Repetitive sequence-based PCR (rep-PCR) is a molecular technique used to produce genetic fingerprint of bacteria and fungi.
It is used for identification and classification of microorganisms at species, subspecies or strain level. It is based on the presence of repetitive DNA sequences in microbial genome. These sequences are non-coding and occur in many copies.
In this technique, specific primers bind to repetitive elements like REP, ERIC and BOX sequences. The DNA region present between these repeated sequences are amplified by PCR. As the distance between these repeated sequences are different in different strains, different sizes of DNA fragments are produced.
After amplification, the DNA fragments are separated by electrophoresis. It gives a specific banding pattern. This banding pattern is known as genomic fingerprint.
The fingerprint is used to compare genetic relatedness among microorganisms. It is also used in hospital infection tracking, environmental contamination study and food safety analysis.
Working Principle of rep-PCR
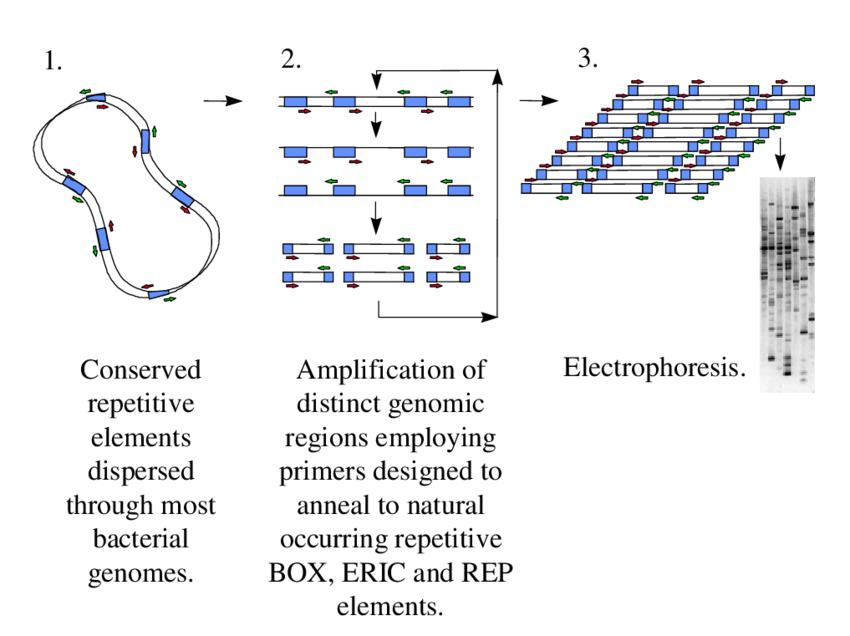
rep-PCR is based on the amplification of DNA regions present between the repetitive sequences of microbial genome. These repetitive sequences are highly conserved, non-coding and present in many copies in the chromosome.
In this technique, specific outward-facing primers bind to the repeated elements like REP, ERIC and BOX sequences. After binding of primers, DNA polymerase extends the primers and amplifies the DNA segment present between two properly oriented repetitive sequences.
The distance and arrangement of these repeated sequences are not same in all bacterial species or strains. So, different sized DNA fragments are produced after amplification. These fragments form a mixture of DNA bands.
After PCR, the amplified fragments are separated by gel electrophoresis or automated system. A specific banding pattern is formed. This pattern is referred to as genomic fingerprint.
This fingerprint helps in identification of microorganisms and also shows genetic relatedness between different bacterial strains.
Sample Requirement for rep-PCR
The sample used for rep-PCR may be purified genomic DNA, crude lysate, whole cells from liquid culture, bacterial colony from agar plate or infected plant tissue.
Purified genomic DNA is mostly preferred for high resolution strain typing. The DNA should be pure and not degraded. The OD 260/280 ratio of DNA should be between 1.8 to 2.0.
For isolated DNA, the concentration is generally maintained between 25 ng/µL to 50 ng/µL. In mixed microbial genomic DNA, total DNA amount should be more than 1 µg and concentration should be more than 10 ng/µL.
The DNA concentration should not be less than 5 ng per reaction. Low DNA amount may produce background amplicons or loss of small and minor bands.
For difficult samples like Gram-positive bacteria, microbial pellet may be used. More than 10 mL microbial solution with OD 600 more than 1.5 can also be used for extraction and rep-PCR.
Protocol of rep-PCR
The following are the steps of rep-PCR–
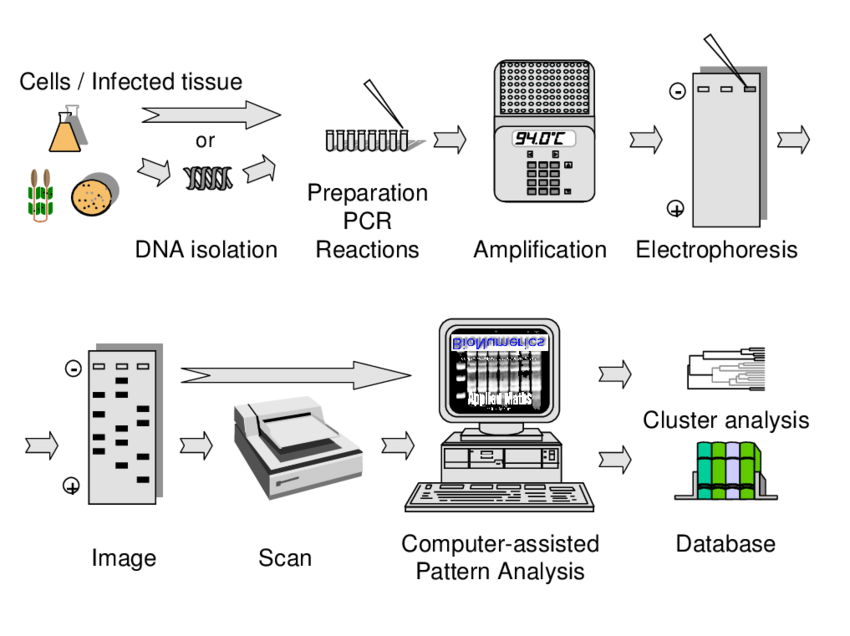
- Genomic DNA is extracted from cultured microbial cells by commercial DNA extraction kit, silica-column method, Chelex-100 method or simple boiling method. The extracted DNA is adjusted to 25 to 50 ng/µL because variation in DNA concentration may affect the final fingerprint pattern.
- The PCR reaction mixture is prepared by adding PCR-grade water, PCR buffer, MgCl₂, dNTPs and Taq DNA polymerase. Specific outward-facing primers are added into the reaction mixture, such as BOXA1R for BOX elements or paired primers for REP and ERIC elements. The standardized template DNA is then added into the master mix.
- The tubes are placed in a thermal cycler for amplification of DNA present between the repetitive elements. Initial denaturation is done at 94°C to 95°C for 2 to 7 minutes. In this step, double stranded DNA is separated into single strands.
- Then 30 to 35 cycles are carried out. Denaturation is done at 94°C to 95°C for 30 to 60 seconds. Annealing is done at 40°C to 65°C for 30 to 60 seconds. Extension is done at 65°C to 72°C for 1.5 to 8 minutes because large DNA fragments may be synthesized.
- Final extension is done at 65°C to 72°C for 15 to 16 minutes. This helps to complete all synthesized DNA strands.
- The amplified PCR products are loaded into agarose gel and separated according to their size by gel electrophoresis. In automated method, the amplified fragments are separated by microfluidic capillary electrophoresis system like DiversiLab platform.
- The agarose gel is stained with DNA binding dye such as ethidium bromide and observed under UV light. A specific banding pattern is formed which is known as genomic fingerprint.
- The banding pattern is digitized by software such as BioNumerics or DiversiLab web platform. Similarity value is calculated by Pearson or Dice coefficient and the strains are grouped into a dendrogram to compare the genetic relatedness.
Uses of rep-PCR
The following are the uses of rep-PCR–
- rep-PCR is used to trace the spread and transmission route of hospital associated infections. It is also used in outbreak investigation.
- It is used in clinical laboratories for routine monitoring of high risk and multidrug resistant pathogens like MRSA and carbapenem-resistant Acinetobacter baumannii.
- It is used in food safety for tracing food borne pathogens like Salmonella and Listeria in food chain. It also helps in microbiological quality control during food manufacturing.
- It is used to identify pseudo-outbreaks in laboratory. It helps to check whether the positive pathogen result is due to actual infection or cross-contamination in clinical or food laboratory.
- It is used in environmental monitoring and water quality management. It is also used in microbial source tracking to identify the source of fecal pollution.
- It is used for microbial identification and classification. It helps to study microbial diversity and genetic relatedness of different bacterial and fungal isolates.
- It is used in agricultural and veterinary diagnosis. It helps in tracking veterinary infections and characterization of plant pathogens.
- It is used in biodefense and microbial forensic science because it can identify microorganisms at strain level.
Advantages of rep-PCR
The following are the advantages of rep-PCR–
- rep-PCR is a rapid and simple technique. It gives result within 12 to 24 hours, which is faster than pulsed-field gel electrophoresis (PFGE).
- It is an inexpensive genotyping method. The startup cost, labour cost and equipment cost are lower than many other molecular typing methods.
- It can be performed with different types of samples. Purified genomic DNA, crude cell lysate, whole cells from liquid culture, colonies from plates and infected plant tissues can be used.
- It has high discriminatory power. It can differentiate microorganisms at species, subspecies and strain level.
- It can distinguish closely related isolates during short term outbreak. In some cases, it gives better discrimination than 16S/18S rRNA restriction analysis and MLST.
- It has broad application. It can be used for many Gram-negative bacteria, Gram-positive bacteria and some fungal species.
- It gives stable and reproducible genomic fingerprint when the protocol is standardized. Automated systems like DiversiLab can increase the reproducibility.
- It is suitable for high-throughput screening. Large number of samples or microbial populations can be analyzed in short time.
Limitations of rep-PCR
The following are the limitations of rep-PCR–
- rep-PCR may have lower discriminatory power in some cases. It may not separate very closely related epidemic strains or isolates with very small genomic difference.
- It is slightly less discriminatory than some gold standard typing methods like PFGE and whole genome sequencing (WGS).
- Traditional manual rep-PCR may show low reproducibility between different laboratories. So, comparison of results become difficult without standardized or automated system.
- The interpretation of DNA banding pattern in agarose gel may be subjective. It may take more time and human error can occur during visual analysis.
- The result of rep-PCR is highly affected by technical variables. Small change in primer-template ratio, enzyme concentration, MgCl₂ concentration and PCR condition may change the fingerprint pattern.
- Different thermocycler machines may give different genomic fingerprints. This occurs due to difference in ramp speed and temperature uniformity of heating block.
- Different source or concentration of Taq DNA polymerase may affect amplification. It may produce artificial bands or weak amplification.
- High MgCl₂ concentration may cause non-specific binding. Low MgCl₂ concentration may fail to amplify larger DNA fragments.
- In environmental samples, interpretation of rep-PCR profile is complex because wild bacterial populations have high genetic diversity. So, simple clustering may not always give correct source identification.
References
- CD Genomics. (n.d.). Rep-PCR profiling. https://www.cd-genomics.com/microbioseq/rep-pcr-profiling.html.
- Creative Biolabs-Live Biotherapeutics. (n.d.). Rep-PCR for microbial molecular identification. https://live-biotherapeutic.creative-biolabs.com/rep-pcr-in-microbial-molecular-identification.htm.
- European and Mediterranean Plant Protection Organization. (2010). Contributed resources detail: Rep-PCR tests for identification of bacteria. https://www.ippc.int/en/about/core-activities/capacity-development/guides-and-training-materials/contributed-resource-detail/rep-pcr-tests-identification-bacteria/.
- PubMed. (n.d.). [REP and ERIC repetitive DNA sequences in bacteria–diagnostic significance]. https://pubmed.ncbi.nlm.nih.gov/10803021/.
- ResearchGate. (n.d.). Overview of automated rep-PCR and the DiversiLab system. (A) rep-PCR… https://www.researchgate.net/figure/Overview-of-automated-rep-PCR-and-the-DiversiLab-system-A-rep-PCR-primers-bind-to-many_fig1_8349667.
- PMC. (n.d.). Impact of small repeat sequences on bacterial genome evolution. https://pmc.ncbi.nlm.nih.gov/articles/PMC3184786/.
- PMC. (n.d.). Enterobacterial repetitive intergenic consensus sequence repeats in Yersiniae: Genomic organization and functional properties. https://pmc.ncbi.nlm.nih.gov/articles/PMC1291288/.
- ASM Journals. (1997). Factors affecting reliability and reproducibility of … https://journals.asm.org/doi/pdf/10.1128/jcm.35.2.339-346.1997.
- Bilung, L. M., Pui, C. F., Su’ut, L., & Apun, K. (2018). Evaluation of BOX-PCR and ERIC-PCR as molecular typing tools for pathogenic Leptospira. Disease Markers, 2018, 1351634. https://pmc.ncbi.nlm.nih.gov/articles/PMC6092967/.
- Khan Academy. (n.d.). Polymerase chain reaction (PCR) (article). https://www.khanacademy.org/science/ap-biology/gene-expression-and-regulation/biotechnology/a/polymerase-chain-reaction-pcr.
- The Plant Pathology Journal. (n.d.). Genetic diversity and population structure of phyllosphere-associated Xanthomonas euvesicatoria bacteria in Physalis pubescens based on BOX-PCR and ERIC-PCR in China. https://www.ppjonline.org/journal/view.php?number=2490.
- PMC. (n.d.). Comparison of an automated repetitive sequence-based PCR microbial typing system to pulsed-field gel electrophoresis for analysis of outbreaks of methicillin-resistant Staphylococcus aureus. https://pmc.ncbi.nlm.nih.gov/articles/PMC1287783/.
- Healy, M., Huong, J., Bittner, T., Lising, M., Frye, S., Raza, S., Schrock, R., Manry, J., Renwick, A., Nieto, R., Woods, C., Versalovic, J., & Lupski, J. R. (2005). Microbial DNA typing by automated repetitive-sequence-based PCR. Journal of Clinical Microbiology, 43(1), 199–207. https://pmc.ncbi.nlm.nih.gov/articles/PMC540112/.
- European and Mediterranean Plant Protection Organization. (2010). PM 7/100(1): RepPCR tests for identification of bacteria. EPPO Bulletin, 40, 365–368. https://gd.eppo.int/download/standard/224/pm7-100-1-en.pdf.
- PMC. (n.d.). Rapid detection of Pseudomonas aeruginosa and Acinetobacter baumannii harboring blaVIM-2, blaIMP-1 and blaOXA-23 genes by using loop-mediated isothermal amplification methods. https://pmc.ncbi.nlm.nih.gov/articles/PMC4697338/.
- PMC. (n.d.). Validation of use of whole-cell repetitive extragenic palindromic sequence-based PCR (REP-PCR) for typing strains belonging to the Acinetobacter calcoaceticus-Acinetobacter baumannii complex and application of the method to the investigation of a hospital outbreak. https://pmc.ncbi.nlm.nih.gov/articles/PMC228981/.
- MilliporeSigma. (n.d.). Standard PCR protocol. https://www.sigmaaldrich.com/US/en/technical-documents/protocol/genomics/pcr/standard-pcr.
- PMC. (n.d.). Repetitive-sequence-based PCR using the DiversiLab system for identification of Aspergillus species. https://pmc.ncbi.nlm.nih.gov/articles/PMC2395099/.
- YouTube. (n.d.). Basic molecular biology: PCR and real-time PCR – Principle of PCR. https://www.youtube.com/watch?v=db0HzFTJtCs.
- PMC. (n.d.). Clinical evaluation of the DiversiLab microbial typing system using repetitive-sequence-based PCR for characterization of Staphylococcus aureus strains. https://pmc.ncbi.nlm.nih.gov/articles/PMC1081226/.
- Applied Maths. (n.d.). Brochure BioNumerics.indd. https://www.bockytech.com.tw/PDF-File/bn_brochure.pdf.
- Coleman, E. W. (2009). Performing comparisons in BioNumerics. PulseNet International. https://www.pulsenetinternational.org/assets/PulseNet/uploads/bionumerics/bion_comparisons.pdf.
- ResearchGate. (n.d.). Common applications with BioNumerics. https://www.researchgate.net/profile/Mohamed-Mourad-Lafifi/post/How-do-you-create-composite-fingerprints-from-RiboPrinter-patterns-using-Bionumerics-software/attachment/5a72fb954cde266d5887f457/AS%3A589188734922766%401517484949839/download/bionumerics.pdf.
- Pasanen, T., Koskela, S., Mero, S., Tarkka, E., Tissari, P., & Kirveskari, J. (2014). Rapid molecular characterization of Acinetobacter baumannii clones with rep-PCR and evaluation of carbapenemase genes by new multiplex PCR in Hospital District of Helsinki and Uusimaa. PLoS ONE, 9(1), e85854. https://journals.plos.org/plosone/article/file?id=10.1371/journal.pone.0085854&type=printable.
- PMC. (n.d.). Antimicrobial resistance patterns and molecular characterization of Acinetobacter baumannii isolates in non-COVID-19 patients admitted to intensive care units during the pandemic: A retrospective study at a tertiary hospital in Tehran, Iran. https://pmc.ncbi.nlm.nih.gov/articles/PMC12619513/.
- PubMed. (n.d.). Citywide clonal outbreak of multiresistant Acinetobacter baumannii and Pseudomonas aeruginosa in Brooklyn, NY: The preantibiotic era has returned. https://pubmed.ncbi.nlm.nih.gov/12090889/.
- Osong Public Health and Research Perspectives. (n.d.). A comparison of subtyping methods for differentiating Salmonella enterica serovar Enteritidis isolates obtained from food and human sources. https://ophrp.org/journal/view.php?number=165.
- ASM Journals. (n.d.). Comparison of pulsed-gel electrophoresis and a commercial repetitive-element PCR method for assessment of methicillin-resistant Staphylococcus aureus clustering in different health care facilities. https://journals.asm.org/doi/10.1128/jcm.03466-13.
- Liang, H., Yu, Z., Wang, B., Ndayisenga, F., Liu, R., Zhang, H., & Wu, G. (2021). Synergistic application of molecular markers and community-based microbial source tracking methods for identification of fecal pollution in river water during dry and wet seasons. Frontiers in Microbiology, 12, 660368. https://www.frontiersin.org/journals/microbiology/articles/10.3389/fmicb.2021.660368/full.
- Southern California Coastal Water Research Project. (n.d.). Microbial source tracking. https://www.sccwrp.org/about/research-areas/microbial-water-quality/microbial-source-tracking/.
- Journal of Water and Health. (n.d.). Microbial source tracking of fecal contamination in Laguna Lake … https://iwaponline.com/jwh/article/19/5/762/83842/Microbial-source-tracking-of-fecal-contamination.
- PMC. (n.d.). Performance, design, and analysis in microbial source tracking studies. https://pmc.ncbi.nlm.nih.gov/articles/PMC1855604/.
- PMC. (n.d.). Recent trends and developments of PCR-based methods for the detection of food-borne Salmonella bacteria and Norovirus. https://pmc.ncbi.nlm.nih.gov/articles/PMC9579247/.
- Hygiena. (n.d.). Comprehensive guide to real-time PCR for food safety testing. https://www.hygiena.com/learning-center/technology-guide/comprehensive-guide-to-real-time-pcr-for-food-safety-testing.
- USDA Food Safety and Inspection Service. (n.d.). Outbreak investigations: Response. https://www.fsis.usda.gov/food-safety/foodborne-illness-and-disease/outbreaks/outbreak-investigations-response.
- Marshall, K. E., Nguyen, T. A., Ablan, M., Nichols, M. C., Robyn, M. P., Sundararaman, P., Whitlock, L., Wise, M. E., & Jhung, M. A. (2020). Investigations of possible multistate outbreaks of Salmonella, Shiga toxin–producing Escherichia coli, and Listeria monocytogenes infections — United States, 2016. MMWR Surveillance Summaries, 69(6), 1–14. https://www.cdc.gov/mmwr/volumes/69/ss/ss6906a1.htm.
- Integrated Food Safety Centers of Excellence. (n.d.). Foodborne outbreak investigation case study: Salmonella associated with salami sticks [Video]. YouTube. https://www.youtube.com/watch?v=SZCQAbKU0VI.
- PMC. (n.d.). Case report of Salmonella cross-contamination in a food laboratory. https://pmc.ncbi.nlm.nih.gov/articles/PMC4787000/.
- New England Biolabs. (n.d.). Understanding variability in DNA amplification reactions. https://www.neb.com/en/tools-and-resources/feature-articles/understanding-variability-in-dna-amplification-reactions.
- Mills, M. G., Bruce, E., Huang, M. L., Crothers, J. W., Hyrien, O., Oura, C. A. L., Blake, L., Jordan, A. B., Hester, S., Wehmas, L., Mari, B., Barby, P., Lacoux, C., Fassy, J., … & Jerome, K. R. (2022). An international, interlaboratory ring trial confirms the feasibility of an extraction-less “direct” RT-qPCR method for reliable detection of SARS-CoV-2 RNA in clinical samples. PLoS ONE, 17(1), e0261853. https://pmc.ncbi.nlm.nih.gov/articles/PMC8758094/.
- Mills, M. G., Bruce, E., Huang, M. L., Crothers, J. W., Hyrien, O., Oura, C. A. L., Blake, L., Jordan, A. B., Hester, S., Wehmas, L., Mari, B., Barby, P., Lacoux, C., Fassy, J., … & Jerome, K. R. (2022). An international, interlaboratory ring trial confirms the feasibility of an extraction-less “direct” RT-qPCR method for reliable detection of SARS-CoV-2 RNA in clinical samples. PLoS ONE, 17(1), e0261853. https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0261853.
- Mills, M. G., Bruce, E., Huang, M. L., Crothers, J. W., Hyrien, O., Oura, C. A. L., Blake, L., Jordan, A. B., Hester, S., Wehmas, L., Mari, B., Barby, P., Lacoux, C., Fassy, J., … & Jerome, K. R. (2022). An international, interlaboratory ring trial confirms the feasibility of an extraction-less “direct” RT-qPCR method for reliable detection of SARS-CoV-2 RNA in clinical samples. PLoS ONE, 17(1), e0261853. https://pubmed.ncbi.nlm.nih.gov/33880478/.
- Mills, M. G., Bruce, E., Huang, M. L., Crothers, J. W., Hyrien, O., Oura, C. A. L., Blake, L., Jordan, A. B., Hester, S., Wehmas, L., Mari, B., Barby, P., Lacoux, C., Fassy, J., … & Jerome, K. R. (2021). An international, interlaboratory ring trial confirms the feasibility of an open-source, extraction-less “direct” RT-qPCR method for reliable detection of SARS-CoV-2 RNA in clinical samples. medRxiv. https://www.medrxiv.org/content/10.1101/2021.04.10.21254091v2.
- Mills, M. G., Bruce, E., Huang, M. L., Crothers, J. W., Hyrien, O., Oura, C. A. L., Blake, L., Jordan, A. B., Hester, S., Wehmas, L., Mari, B., Barby, P., Lacoux, C., Fassy, J., … & Jerome, K. R. (2022). An international, interlaboratory ring trial confirms the feasibility of an extraction-less “direct” RT-qPCR method for reliable detection of SARS-CoV-2 RNA in clinical samples. PLoS ONE, 17(1), e0261853. https://repositorio.udd.cl/entities/publication/75188968-2e01-46e4-9e79-d097dc97c46d.
- ResearchGate. (n.d.). Comparison of multilocus sequence typing (MLST), pulsed-field gel electrophoresis (PFGE), and amplified fragment length polymorphism (AFLP) for genetic typing of Staphylococcus aureus. https://www.researchgate.net/publication/6459778_Comparison_of_multilocus_sequence_typing_MLST_pulsed-field_gel_electrophoresis_PFGE_and_amplified_fragment_length_polymorphism_AFLP_for_genetic_typing_of_Staphylococcus_aureus.
- PMC. (n.d.). A comparison of molecular typing methods applied to Enterobacter cloacae complex: hsp60 sequencing, rep-PCR, and MLST. https://pmc.ncbi.nlm.nih.gov/articles/PMC5394936/.
- PMC. (n.d.). Comparison of pulsed-field gel electrophoresis & repetitive sequence-based PCR methods for molecular epidemiological studies of Escherichia coli clinical isolates. https://pmc.ncbi.nlm.nih.gov/articles/PMC4311324/.