Pulsed-field Gel Electrophoresis (PFGE) is a molecular biology technique used for separation of very large DNA molecules in an agarose gel.
It is used as a DNA fingerprinting method for large chromosomal DNA fragments. Normal agarose gel electrophoresis can separate only smaller DNA fragments, but PFGE can separate DNA fragments up to about 10 megabase pairs (Mb).
In PFGE, the electric field is changed in different directions during electrophoresis. Due to this changing field, the long DNA molecules must uncoil and again arrange themselves according to the new direction of electric field.
Large DNA fragments take more time to reorient than small fragments. So they move slowly through the gel. Small DNA fragments reorient quickly and move faster.
Thus different size DNA fragments are separated and form a distinct banding pattern. This banding pattern is used in genomic analysis, physical mapping and detection of small genetic variation.
Principle of Pulsed Field Gel Electrophoresis (PFGE)
Pulsed Field Gel Electrophoresis (PFGE) is based on the separation of very large DNA molecules by alternating electric field.
The large DNA fragments cannot be separated properly by ordinary agarose gel electrophoresis. In continuous electric field, the large DNA molecules moves through the gel in same rate. This movement is called reptation.
In PFGE, the direction of electric field is changed after fixed time interval. Due to this, the DNA molecules changes its direction during migration.
When the electric field is changed, the elongated DNA molecules first uncoil and then orient itself with the new electric field. After this, it starts moving through the gel.
Large DNA molecules takes more time for reorientation. Small DNA molecules takes less time for reorientation. So large fragments moves slowly and small fragments moves faster.
Thus the DNA fragments are separated according to their size. It is used for separation of large DNA fragments up to about 10 megabase pairs (Mb).
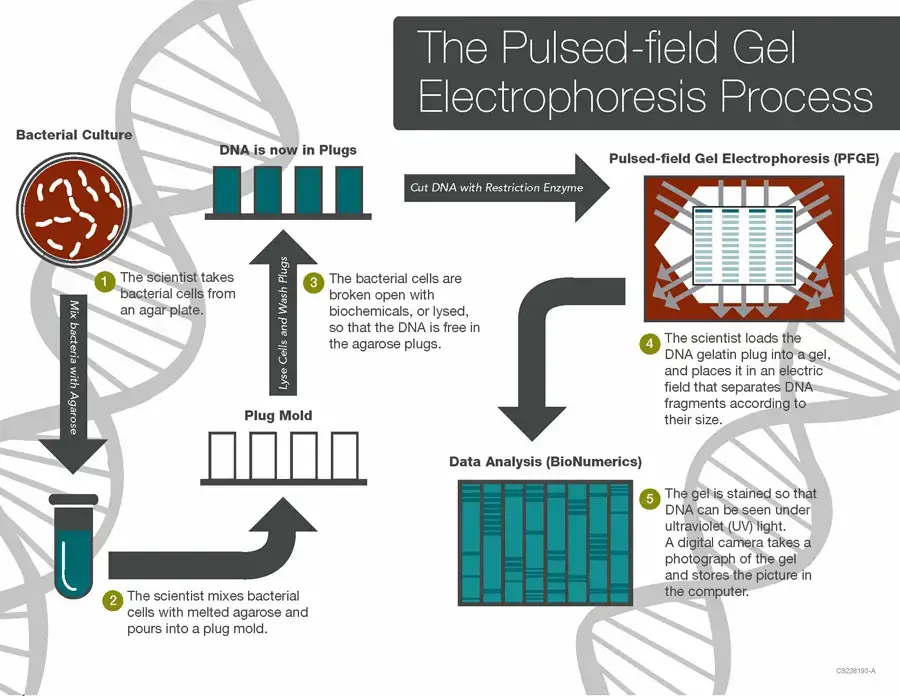
Procedure of Pulsed Field Gel Electrophoresis (PFGE)
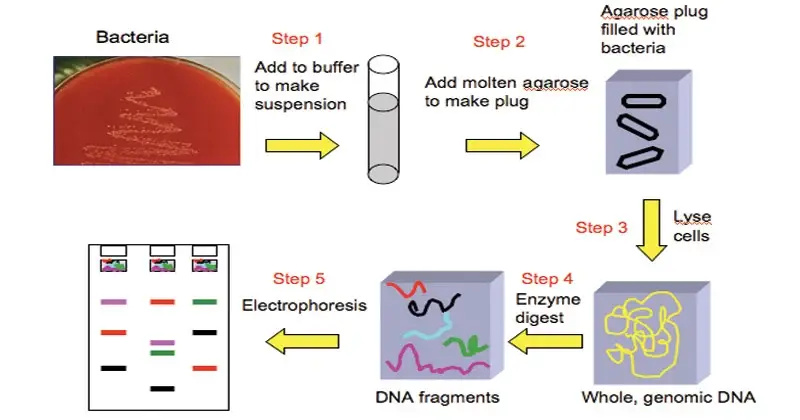
The following are the steps of Pulsed Field Gel Electrophoresis (PFGE)–
- The cells such as bacterial cells are collected from the culture and suspended in a buffer solution. The concentration of cell suspension is adjusted by using spectrophotometer or turbidity meter.
- The standardized cell suspension is mixed with Proteinase K and warm melted agarose. This mixture is poured into small plug moulds and allowed to solidify to form agarose plugs.
- The agarose plug protects the large DNA molecules from mechanical breaking during handling. So the high molecular weight DNA remains intact inside the plug.
- The solidified plugs are placed in lysis buffer containing detergent such as Sarcosyl and Proteinase K. The plugs are incubated in water bath.
- During this step, the cell membrane is broken and cellular proteins are digested. The intact DNA is released inside the agarose plug.
- The plugs are washed several times with warm ultrapure water and TE buffer. This removes cell debris, detergent and remaining Proteinase K.
- The washing step is important because these substances may inhibit the activity of restriction enzymes in the next step.
- A small slice of agarose plug is cut and incubated with rare-cutting restriction enzyme and suitable buffer.
- The restriction enzyme cuts the intact genomic DNA at specific sites. Large DNA fragments are formed.
- Agarose gel is prepared for electrophoresis. The digested plug slices are loaded into the wells of the gel.
- The plug slices are sealed with molten agarose, so that they remain fixed in the wells during electrophoresis.
- The gel is placed in the PFGE chamber containing chilled electrophoresis buffer. The temperature is generally maintained at about 14°C.
- The electrophoresis is run with selected voltage, switch time and total running time. It usually takes about 18 to 21 hours.
- After electrophoresis, the gel is stained with fluorescent dye such as Ethidium bromide or GelRed.
- The gel is observed under UV transilluminator. The separated DNA fragments appear as distinct banding pattern.
Applications of Pulsed Field Gel Electrophoresis (PFGE)
The following are the applications of Pulsed Field Gel Electrophoresis (PFGE)–
- PFGE is used for DNA fingerprinting of bacterial strains, and helps in detection and tracing of food borne disease outbreak caused by E. coli, Salmonella and Listeria.
- It is used for tracking the spread of multidrug resistant organisms in hospitals.
- PFGE is used for estimation of total genome size of bacteria and bacteriophages, and also for preparation of physical and genetic map of whole chromosome.
- It is used to detect large gene rearrangement, deletion and duplication, and useful in diagnosis of genetic diseases like Duchenne muscular dystrophy and Becker muscular dystrophy.
- In cancer study, PFGE is used to detect chromosomal translocation in malignancy such as follicular lymphoma, and also used to measure DNA damage like double strand break caused by radiation or chemical exposure.
- It gives discrete banding pattern, so genetic relation among different strains of same species and their genetic history can be studied.
- PFGE is used for selection and isolation of very large DNA fragments for cloning, and useful in construction and characterization of Yeast Artificial Chromosome (YAC) and Bacterial Artificial Chromosome (BAC) libraries.
- In food and beverage industry, it is used to monitor genetic identity and stability of microbes used in food production, and also used in agricultural products like grapes and hops used in wine and beer making.
- In environmental microbiology, PFGE is used to isolate and detect large megaplasmids, and helps in study of horizontal gene transfer and environmental spread of antibiotic resistance.
- PFGE is used in investigation of bioterrorism agents by microbial fingerprinting.
- It is also used in transgenic research for experimental construction and study of transgenic mice.
Advantages of Pulsed Field Gel Electrophoresis (PFGE)
The following are the advantages of Pulsed Field Gel Electrophoresis (PFGE)–
- PFGE can separate very large DNA fragments up to about 10 megabase pairs (Mb) or more, which is not possible in ordinary agarose gel electrophoresis.
- It is useful for preparation of physical and genetic map of chromosome, measuring genome size and studying genetic relation between different strains.
- Very large restriction fragments can be selected and isolated by PFGE, so it is useful in cloning, gene isolation and functional gene mapping.
- PFGE gives stable macro-restriction pattern in some rapidly changing pathogens, so it is useful for outbreak tracking of such organisms.
- It is simple and less costly method for local outbreak investigation, especially in small hospitals and limited laboratory.
- It can be used in different organisms from viruses to mammals, and useful in tracing food borne disease outbreak, study of DNA damage and diagnosis of genetic diseases and chromosomal translocation in cancers.
Limitations of Pulsed Field Gel Electrophoresis (PFGE)
The following are the limitations of Pulsed Field Gel Electrophoresis (PFGE)–
- PFGE is a time consuming method because separation of large DNA molecules takes long time, and standard diagnostic procedure may need about 2 to 4 days.
- It is a laborious method and requires more resources, because the process is complex, multi-step and needs trained person for proper handling.
- PFGE gives only large restriction fragment pattern, so minute genetic changes cannot be detected properly and true relation between organisms may not always be identified.
- The high molecular weight DNA is very fragile and easily broken, so cells must be embedded and processed inside solid agarose plugs.
- The result of PFGE depends on proper temperature, voltage, pulse time and buffer condition, and small change in these conditions may cause band smearing or distorted bands.
- In some genetic disease diagnosis, PFGE may give false positive or false negative result because of different haplotype or array structure of patient.
References
- A general method for detecting and sizing large plasmids. (n.d.). PubMed.
- Agilent Technologies. (n.d.). Femto Pulse system.
- Basim, E., & Basim, H. (2001). Pulsed-field gel electrophoresis (PFGE) technique and its use in molecular biology. Turkish Journal of Biology, 25(4), 405-418.
- Bio-Rad. (n.d.). CHEF solutions.
- Bio-Rad. (n.d.). Pulsed field gel electrophoresis.
- Bio-Rad. (2014). Pulsed field gel electrophoresis (Bulletin 6224).
- Blue Cross NC. (2026, January). Genetic testing for muscular dystrophies AHS – M2074.
- Carleton, H. (2019, May 23). 2019: PulseNet laboratories transition to whole genome sequencing. Centers for Disease Control and Prevention Archive.
- Centers for Disease Control and Prevention. (2012, August). Unified pulsed-field gel electrophoresis (PFGE) protocol for Gram positive bacteria.
- Centers for Disease Control and Prevention. (2013, March 24). Standard operating procedure for PulseNet PFGE of Escherichia coli O157:H7, Escherichia coli non-O157 (STEC), Salmonella serotypes, Shigella sonnei and Shigella flexneri.
- Centers for Disease Control and Prevention. (2014, January 23). Modified Pulse-Net procedure for pulsed-field gel electrophoresis of select Gram negative bacilli.
- Centers for Disease Control and Prevention. (2024, March 4). Detecting outbreaks with whole genome sequencing.
- Comparison of pulsed-field gel electrophoresis and whole-genome-sequencing-based typing confirms the accuracy of pulsed-field gel electrophoresis for the investigation of local Pseudomonas aeruginosa outbreaks. (n.d.). PubMed.
- Determination of bacteriophage genome size by pulsed-field gel electrophoresis. (n.d.). ResearchGate.
- Duplication detection in Japanese Duchenne muscular dystrophy patients and identification of carriers with partial gene deletions using pulsed-field gel electrophoresis. (n.d.). PubMed.
- Enhanced detection of the t(14;18) translocation in malignant lymphoma using pulsed-field gel electrophoresis. (n.d.). PubMed.
- Enhanced detection of the t(14;18) translocation in malignant lymphoma using pulsed-field gel electrophoresis. (n.d.). ResearchGate.
- Epidemiology in foodborne outbreaks: The role of PulseNet, whole genome sequencing, and pulsed-field gel electrophoresis. (n.d.). Ron Simon & Associates.
- Exploring PFGE for detecting large plasmids in Campylobacter jejuni and Campylobacter coli isolated from various retail meats. (n.d.). PMC.
- Grand View Research. (2024). Automated sample preparation technology market (2025 – 2033).
- Howes, R. (2026, March 13). Discovery is changing – and automation leads the way. Drug Target Review.
- Napoli, K. A. (2024). Pulsed-field gel electrophoresis. Research Starters, EBSCO Information Services.
- Physical mapping and YAC-cloning connects four genetically distinct 4qter loci (D4S163, D4S139, D4F35S1 and D4F104S1) in the FSHD gene-region. (n.d.). PubMed.
- Physical mapping of large DNA by chromosome fragmentation. (n.d.). PMC – NIH.
- Prinzi, A. (2021, July 28). Whole genome sequencing (WGS) as a tool for hospital surveillance. American Society for Microbiology.
- Pulse field gel electrophoresis. (n.d.). PMC.
- PulseNet international: The missing link between PFGE and WGS. (n.d.). bioRxiv.
- PulseNet international: The missing link between PFGE and WGS. (n.d.). bioRxiv.
- Pulsed field gel electrophoresis and genome size estimates. (n.d.). PubMed.
- Pulsed field gel electrophoresis for detection of gene rearrangements in Duchenne muscular dystrophy. (n.d.). PubMed.
- Pulsed field gel electrophoresis: Past, present, and future. (n.d.). PubMed.
- Pulsed-field gel electrophoresis (PFGE) technique and its use in molecular biology. (n.d.). SciSpace.
- Rotating field gel electrophoresis (ROFE). (n.d.). PubMed.
- Streamlined approach to creating yeast artificial chromosome libraries from specialized cell sources. (n.d.). PNAS.
- The application of pulsed field gel electrophoresis in clinical studies. (n.d.). PMC.
- The comprehensive theory and application of pulsed-field gel electrophoresis in genomic analysis and molecular surveillance. (n.d.).
- The plasmid-host fitness landscape: A new paradigm for predicting the fate of mobile resistance. (n.d.). PMC.
- Wikipedia contributors. (2026, April 26). Pulsed-field gel electrophoresis. In Wikipedia, The Free Encyclopedia.
- Zero Hour Health. (2020, February 19). You may read about “WGS” vs. “PFGE” in the context of outbreaks. Do you need to care?