PCR Machine is an automated laboratory instrument that is used to amplify small segment of DNA or RNA into millions or billions of copies.
It is also known as thermal cycler or DNA amplifier. It contains a thermal block where reaction tubes containing sample, primers, nucleotides and polymerase enzyme are placed.
The machine changes the temperature in pre-programmed cycles. These temperature cycles allow denaturation of DNA, annealing of primers and synthesis of new DNA strands. It complete the amplification process within few hours.
Modern PCR machine also has heated lid which prevents condensation inside the reaction tube. It also has control panel for setting temperature, time and number of cycles according to the experiment.
Principle of PCR
PCR is based on the principle of nucleic acid hybridization and nucleic acid replication. It is a temperature dependent cyclic process where double stranded DNA (dsDNA) is first separated into single stranded DNA (ssDNA) by heat.
In these single stranded DNA templates, primers bind to their complementary sequences. This is called annealing. Then DNA polymerase enzyme adds nucleotides to the 3’ end of primer and forms new double stranded DNA.
The reaction occurs in three steps. These are denaturation, annealing and extension. In denaturation, DNA strands are separated by high temperature. In annealing, primers bind with the target DNA sequence. In extension, Taq DNA polymerase synthesizes new DNA strand at about 72°C.
These three steps are repeated in cyclic manner. So the newly formed DNA again acts as template in next cycle. Thus millions or billions copies of a selected DNA segment are formed in few hours. This is why PCR is also called molecular photocopying.
Parts of a PCR Machine
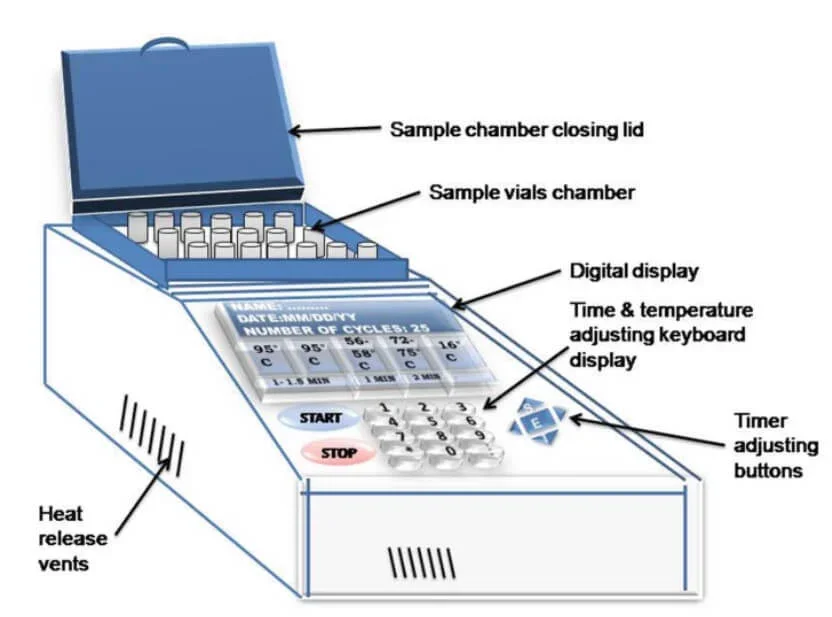
The PCR machine is made up of different parts which are used for holding reaction tubes, changing temperature and controlling the amplification process.
- Thermal block – It is the main part of the PCR machine where PCR reaction tubes are placed. It has small holes for holding tubes containing DNA sample, primers, nucleotides and polymerase enzyme.
- Heated lid – It is present above the thermal block. It presses the reaction tubes and prevent condensation of water inside the tube lid. It also helps in proper contact of tubes with the block.
- Control panel and keypad – It is used to set the temperature, time and number of cycles. The display shows the running condition and program of the machine.
- Air vents – These are openings present on the side, front or bottom part of the machine. They help in air intake and exhaust for cooling the system.
- Peltier elements – These are internal thermoelectric parts which help in rapid heating and cooling of the thermal block. They maintain the cyclic temperature changes during PCR.
- Indicator lights – These lights show the working condition of the machine. It also gives warning when the thermal block or heated lid is hot.
- Power switch and electrical system – It supplies electricity to the machine. It helps in running the heating, cooling and control system of the PCR machine.
Components/reagents Required for PCR
The following are the components required for PCR reaction.
- DNA template – Template DNA is the sample DNA that contain the target sequence. This selected sequence is amplified in PCR.
- Thermostable DNA polymerase – It is a heat stable enzyme used for synthesis of new DNA strand. Taq DNA polymerase is most commonly used because it remains active after repeated heating.
- Oligonucleotide primers – Primers are short single stranded nucleotide sequence. They are complementary to the target region of DNA. They provide starting point for DNA polymerase.
- Deoxyribonucleotide triphosphates (dNTPs) – These are free nucleotides used for making new DNA strand. It includes dATP, dGTP, dCTP and dTTP.
- Buffer system – Buffer is used to maintain the proper pH of the reaction mixture. It generally contains Tris-HCl and potassium chloride (KCl).
- Magnesium ions (Mg²⁺) – Mg²⁺ is an important cofactor for DNA polymerase enzyme. It is usually added as magnesium chloride (MgCl₂).
- Nuclease-free water – It is pure water used to make the final volume of reaction mixture. It is free from nuclease enzyme and prevent breakdown of DNA.
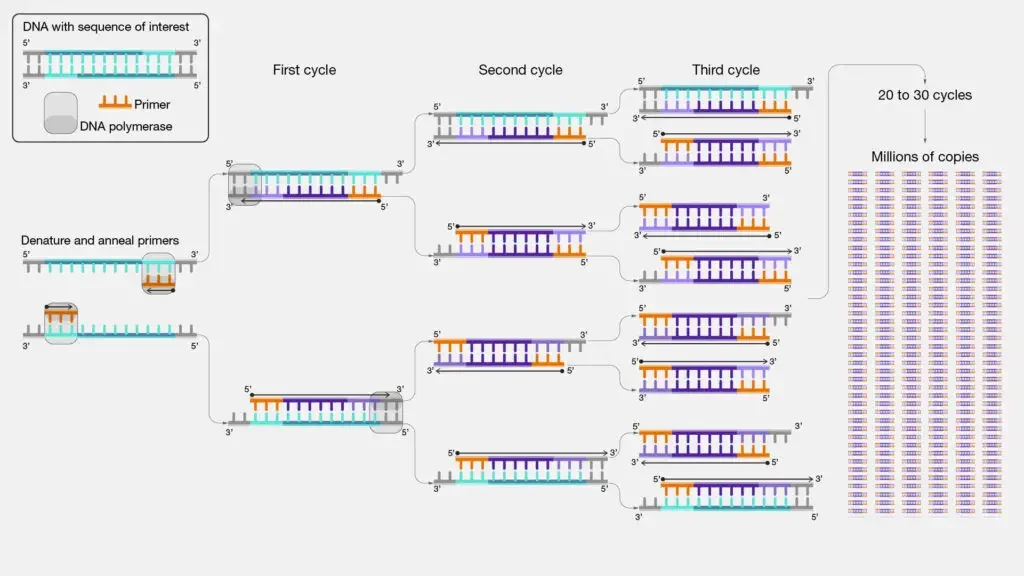
Steps of PCR – Process of PCR (PCR cycling)
PCR cycling is the repeated heating and cooling process of PCR reaction. It is done automatically by PCR machine or thermal cycler.
In this process the temperature is changed in fixed order. The target DNA is copied again and again. Generally 25 to 40 cycles are used in PCR. In every cycle, newly formed DNA also acts as template for next cycle.
The following are the steps of PCR cycling–
- Initial denaturation – In this step, the reaction mixture is first heated at 94-98°C for 2 to 10 minutes. This temperature separates the complex double stranded DNA into single strands.
- Denaturation – In this step, temperature is raised at 94-100°C for 10 to 60 seconds. The hydrogen bonds present between two strands of DNA are broken. As a result, double stranded DNA becomes single stranded DNA.
- Annealing – In this step, temperature is lowered at 45-65°C for 20 to 40 seconds. The short primers bind with their complementary sequence on the single stranded DNA template.
- Extension or elongation – In this step, temperature is raised at about 72°C. This temperature is suitable for Taq DNA polymerase enzyme. The enzyme adds nucleotides at the 3’ end of the primer and new DNA strand is formed.
- Cycle repetition – Denaturation, annealing and extension form one complete PCR cycle. These steps are repeated for 25 to 40 times. In each cycle, the amount of target DNA becomes double.
- Final extension – After completion of all cycles, the reaction is kept at 72°C for 2 to 10 minutes. In this step, incomplete DNA strands are completed by DNA polymerase.
- Hold – At last, the temperature is lowered to 4°C. The amplified DNA samples are kept stable until they are taken out from the machine.
Operating Procedure of PCR Machine
The following are the operating procedure of PCR machine–
- Before starting the machine, the thermal block is checked properly. It should be clean and dry. The heat indicator light should be off before touching the block.
- The prepared PCR reaction mixture is taken in PCR tubes or PCR plate. Then the tubes are placed into the holes of the thermal block.
- The tubes should be properly closed. It prevents leakage of reaction mixture and evaporation of sample.
- The tubes are placed evenly in the block. Empty spaces may be balanced with empty tubes when needed.
- The heated lid is closed firmly over the tubes. It presses the tube caps and prevents condensation of water inside the tubes.
- The temperature and time for denaturation, annealing and extension are set by using control panel.
- The number of cycles are also set. Generally 20 to 40 cycles are used in PCR.
- After setting the program, the run is started. The PCR machine automatically changes the temperature according to the selected program.
- In the cycling process, DNA strands are separated by heating. Primers bind during cooling and new DNA strands are synthesized by Taq DNA polymerase.
- After completion of all cycles, the machine may keep the samples at 4°C. This keeps the amplified DNA stable.
- The lid is opened carefully because the heated lid and block may remain hot. The PCR tubes are then removed from the block.
- After removing the samples, the machine is switched off. The block and working area are kept clean for next use.
Types of PCR
The following are the different types of PCR–
- Real-time PCR (qPCR) – It is a type of PCR used to detect and measure the amplified DNA during the reaction. It uses fluorescent dye or probe.
- Reverse Transcriptase PCR (RT-PCR) – It is used for amplification of RNA. In this method, RNA is first converted into complementary DNA (cDNA) by reverse transcriptase enzyme.
- Multiplex PCR – It is a type of PCR in which more than one target DNA sequence is amplified in a single reaction. It uses multiple primer pairs.
- Nested PCR – It is carried out by using two sets of primers in two successive reactions. The second primer binds inside the first amplified product and increase specificity.
- Hot Start PCR – In this method, DNA polymerase remains inactive at low temperature. It becomes active only after high temperature denaturation and prevents non-specific amplification.
- Digital PCR (dPCR) – In this type, one sample is divided into many small reactions. It is used for absolute counting of target DNA molecules.
- Long-range PCR – It is used to amplify large DNA fragments. Special polymerase mixture is used in this type.
- Touchdown PCR – In this method, annealing temperature is kept high in first cycles and then slowly decreased. It helps in specific binding of primers.
- In situ PCR – It is carried out inside fixed cells or tissue section. It is used to detect and locate specific DNA or RNA sequence in cells.
- Single-cell PCR – It is used for amplification of genetic material from single cell. It is useful when very small amount of sample is present.
- Fast-cycling PCR – It is a rapid type of PCR. It uses short time for denaturation, annealing and extension steps.
- Methylation-specific PCR (MSP) – It is used to detect DNA methylation pattern. In this method, DNA is first treated with sodium bisulfite and then amplified.
- Asymmetric PCR – It is used to amplify one strand of double stranded DNA more than the other strand. Unequal amount of primers are used.
- Assembly PCR – It is used to join many short overlapping DNA fragments into one long DNA molecule.
- High-fidelity PCR – It uses proofreading DNA polymerase enzyme. It reduces wrong nucleotide addition during DNA synthesis.
- RAPD (Rapid Amplified Polymorphic DNA) – It uses short random primers for amplification of random regions of genomic DNA. It gives different banding pattern.
- RACE (Rapid Amplification of cDNA Ends) – It is used to amplify unknown ends of mRNA after conversion into cDNA. It is used to study 3’ or 5’ end sequence.
- Differential Display PCR – It is used to compare mRNA expression in two different cells or tissues. It helps to identify different expressed genes.
What Is Taq Polymerase?
Taq polymerase is a heat stable DNA polymerase enzyme. It is used in PCR for synthesis of new DNA strand.
It is obtained from a thermophilic bacterium named Thermus aquaticus. This bacterium lives in hot spring and can tolerate high temperature.
Taq polymerase adds nucleotides at the 3’ end of primer. It uses the single stranded DNA as template and forms complementary DNA strand.
The optimum temperature for Taq polymerase is about 72°C. At this temperature, extension of DNA takes place.
The important property of Taq polymerase is thermostability. It can withstand the high temperature used in denaturation step of PCR. So it does not get destroyed after repeated heating and cooling cycle.
Due to this property, Taq polymerase is the most commonly used enzyme in PCR. It allows automatic amplification of DNA without adding new enzyme in each cycle.
Applications of PCR
The following are the important applications of PCR–
- PCR is used for diagnosis of bacterial, fungal and viral diseases. It is used to detect HIV, SARS-CoV-2, Hepatitis virus and other infectious agents.
- It is used to measure viral load in patient sample. The amount of viral DNA or RNA can be detected by using PCR.
- PCR is used for diagnosis of genetic disorders. It can detect mutation or abnormal gene sequence.
- It is used in forensic science for DNA fingerprinting. It helps to identify criminal, victim and missing person.
- PCR is used in paternity testing. The DNA pattern of child and parents are compared.
- It is used in molecular archaeology. Ancient DNA from old bones, fossils or preserved sample can be amplified.
- PCR is used in genetic research. It helps in study of gene, genotype, rare allele and Single Nucleotide Polymorphism (SNP).
- It is used to study gene expression. mRNA is first converted into cDNA and then amplified.
- PCR is used in molecular cloning. The selected DNA fragment is amplified and inserted into vector.
- It is used in site directed mutagenesis. Specific mutation can be introduced into a gene.
- PCR is used for preparation of DNA sample for sequencing. It is also used in Next Generation Sequencing (NGS) library preparation.
- It is used in prenatal diagnosis. Fetal DNA can be tested for chromosomal abnormality and genetic disease.
- PCR is used in preimplantation genetic diagnosis during in vitro fertilization (IVF). It helps to test embryo before implantation.
- It is used in environmental microbiology. Pathogens present in water, soil and environment can be detected.
- PCR is used in food safety testing. It detects infectious pathogen and contamination in food sample.
Advantages of PCR
The following are the advantages of PCR–
- PCR is highly sensitive technique. It can amplify target DNA from very small amount of sample.
- It can also amplify DNA from degraded or very less starting material. So small sample is enough for PCR.
- PCR is highly specific method. It uses specific primers which bind only with the selected target sequence.
- It produces millions to billions copies of target DNA within few hours. So it is a rapid technique.
- PCR is simple and easy technique. It does not need long culture process for many organisms.
- It is relatively inexpensive method. So it is commonly used in many laboratories.
- PCR is useful in rapid clinical diagnosis. It can detect bacteria, virus and other pathogens in short time.
- It is useful for diagnosis of diseases which are difficult to grow in culture.
- PCR can detect antimicrobial resistance genes. This helps in selection of proper treatment.
- It can be used for many samples at a time. It is suitable for high-throughput analysis.
- PCR can amplify more than one target in same reaction by multiplex PCR.
- It is useful in research, diagnosis, forensic science and genetic analysis.
Limitations of PCR
The following are the limitations of PCR–
- PCR needs prior knowledge of target DNA sequence. Without knowing the sequence, specific primers cannot be designed.
- PCR is highly sensitive technique. So even very small amount of contaminating DNA or RNA can be amplified.
- Contamination may give false positive result. So the sample, reagents and working area should be handled carefully.
- Taq DNA polymerase has no proofreading activity. So wrong nucleotide may be added during DNA synthesis.
- These errors may produce mutation in PCR product. It may be single base change, insertion or deletion.
- PCR is less efficient for amplification of large DNA fragments. As the target size increases, amplification becomes difficult.
- Non-specific amplification may occur in PCR. Sometimes primers bind with other similar sequence instead of target sequence.
- Primer-dimer may be formed when primers bind with each other. It uses primers and nucleotides and reduces the proper amplification.
- Incomplete templates may form chimeric PCR products. These products can interfere with correct result.
- The result of PCR may become confusing when non-specific bands or unwanted products are formed.
Precautions using PCR Machine
The following are the precautions of PCR–
- Separate working area should be used for pre-PCR and post-PCR work. It prevents contamination of sample.
- PCR preparation should be done in laminar flow hood when possible. UV light may be used for decontamination of the working area.
- Clean gloves, mask, hair cover and lab coat should be used during PCR preparation. Gloves should be changed regularly.
- Sterile and filtered pipette tips should be used. It prevents aerosol contamination during pipetting.
- PCR tubes, tips and other disposable materials should be used directly from sterile packet. They should not be touched unnecessarily.
- Reagents should be aliquoted in small amount. It prevents repeated freeze and thawing of main stock.
- Nuclease-free water should be used only for PCR work. It should not be mixed with other laboratory use.
- Tubes containing PCR reagents should be centrifuged for few seconds before opening. It brings the liquid to the bottom and reduces aerosol formation.
- Before starting the run, the thermal block should be checked. It should be clean and free from any spill.
- Only proper PCR tubes or PCR plates should be used in the machine. Other tubes may not fit properly.
- All tubes and plates should be tightly closed. It prevents evaporation and leakage during high temperature step.
- Tubes should be placed evenly in the thermal block. It helps the heated lid to sit properly on the tubes.
- Heated lid should not be forced or over tightened. It may damage the lid mechanism.
- Air vents and power cable of PCR machine should be checked before use. Ventilation should not be blocked.
- After completion of run, hot lid should not be touched directly. It may cause burn injury.
- Samples should be cooled before removing. Forceps may be used for taking out hot tubes.
- Any spill of reaction mixture should be cleaned immediately. Waste should be discarded in proper biohazard container.
- After use, the PCR machine should be switched off. The block and working area should be kept clean for next use.
References
- Anton Paar. (2026). ISO 17025 calibration.
- Applied Biological Materials Inc. (n.d.). Polymerase chain reaction (PCR) – Tips for successful PCR and troubleshooting PCR-qPCR.
- Bamerni, P., König, J., Mistry, L.-A., Schmitt, K., & Wöllenstein, J. (2025). Design and validation of a high-speed miniaturized thermocycler with Peltier elements for efficient PCR thermal cycling. Sensors, 25(22), 7046.
- Belgrader, P., Young, S., Yuan, B., Primeau, M., Christel, L. A., Pourahmadi, F., & Northrup, M. A. (2001). A battery-powered notebook thermal cycler for rapid multiplex real-time PCR analysis. Analytical Chemistry, 73(2), 286-289.
- Bennett, K. (2025). 17025 accreditation: Calibration requirements white paper. Transcat Inc.
- Bento Bioworks Ltd. (2026). PCR thermocycler / heat block module – Bento Lab user manual.
- Bio-Rad Laboratories, Inc. (2026). Introduction to qPCR system.
- Bio-Rad Laboratories, Inc. (2026). Introduction to qPCR system.
- Bio-Rad Laboratories, Inc. (2026). PCR troubleshooting.
- bioRxiv. (n.d.). Donut PCR: A rapid, portable, multiplexed, and quantitative DNA detection platform with single-nucleotide specificity.
- Chai. (2018). OpenPCR design.
- Eppendorf SE. (2020, October 26). Peltiers in PCR matter!.
- Esco Lifesciences. (n.d.). PCR thermal cycler maintenance tips and procedures.
- Farrar, J. S., & Wittwer, C. T. (2015). Extreme PCR: Efficient and specific DNA amplification in 15-60 seconds. Clinical Chemistry, 61(1), 145-153.
- FBI.gov. (n.d.). CODIS archive.
- Iridian Spectral Technologies Ltd. (n.d.). What optical filters are used in qPCR instruments?.
- Karki, P. (2026, April 11). PCR machine (thermal cycler): Principle, parts, types, uses, examples. Microbe Notes.
- Khan Academy. (n.d.). Polymerase chain reaction (PCR).
- Khehra, N., Padda, I. S., & Zubair, M. (2025, July 7). Polymerase chain reaction (PCR). StatPearls Publishing.
- Krishnapur, P. (2023, August 23). Types of PCR – 7 methods, principles & applications. Assay Genie.
- LabX. (n.d.). PCR and thermal cycler technology: Fundamentals, applications, and emerging trends in DNA amplification.
- Lawrence Livermore National Laboratory. (n.d.). Rapid PCR instrumentation technologies.
- Let’s Talk Academy. (2025, March 16). Three steps of PCR cycle – Denaturation, annealing, and extension.
- Merck KGaA. (2026). Polymerase chain reaction. MilliporeSigma.
- MyBioSource. (n.d.). Types of PCR: Conventional, real-time, reverse transcriptase, digital, and multiplex PCR.
- National Human Genome Research Institute. (2020, August 17). Polymerase chain reaction (PCR) fact sheet.
- NCBI. (n.d.). Extreme PCR meets high-speed melting: A step closer to….
- NCBI. (n.d.). Forensic DNA profiling: Autosomal short tandem repeat as a prominent marker in crime investigation.
- NCBI. (n.d.). PCR in forensic science: A critical review.
- NCBI. (n.d.). Polymerase chain reaction: Basic protocol plus troubleshooting and optimization strategies.
- NCBI. (n.d.). Rapid PCR powered by microfluidics: A quick review under the background of COVID-19 pandemic.
- New England Biolabs. (n.d.). PCR optimization (E0555).
- New England Biolabs. (n.d.). PCR troubleshooting guide.
- Photonics Spectra. (n.d.). Optical filters help PCR tests quickly diagnose COVID-19.
- Plant and Soil Sciences eLibrary. (2026). Three temperature cycles | Polymerase chain reaction (PCR).
- Promega Corporation. (2026). STR amplification | STR analysis | STR kits.
- Purtell, Z. (2024, January 4). PCR thermal cycler maintenance and care tips. Labcare Service Ltd.
- QIAGEN. (2026). Types of PCR.
- ResearchGate. (n.d.). Temperature control for PCR thermocyclers based on Peltier-effect thermoelectric.
- Shrivastava, P., Kaitholia, K., Dixit, S., & Kumawat, R. K. (2025, December 2). Enhancing forensic DNA profiling efficiency through alternative STR-based multiplex systems. ISHI News.
- SIMCO. (2026, February 23). ISO/IEC 17025 calibration audit guide: Requirements and common pitfalls.
- Takara Bio Inc. (2026). Troubleshooting your PCR.
- Unknown Author. (n.d.). The architectural and biochemical synthesis of polymerase chain reaction instrumentation: A comprehensive engineering and clinical analysis.
- Wikipedia. (2025, February 19). Thermal cycler.
- Wikipedia. (2026, March 26). Variants of PCR.
- Wittwer, C. T. (2019, October 22). Extreme molecular diagnostics [Video lecture]. ARUP Laboratories.
- Wittwer, C. T., & Farrar, S. J. (2015, April 30). Extreme PCR (U.S. Patent Application Publication No. US20150118715A1). U.S. Patent and Trademark Office.