Phenylalanine Deaminase Test (PDA test) is a biochemical test used to detect the enzyme phenylalanine deaminase produced by some bacteria.
It is used to identify and differentiate gram-negative enteric bacteria, mainly the Proteeae group from other Enterobacteriaceae.
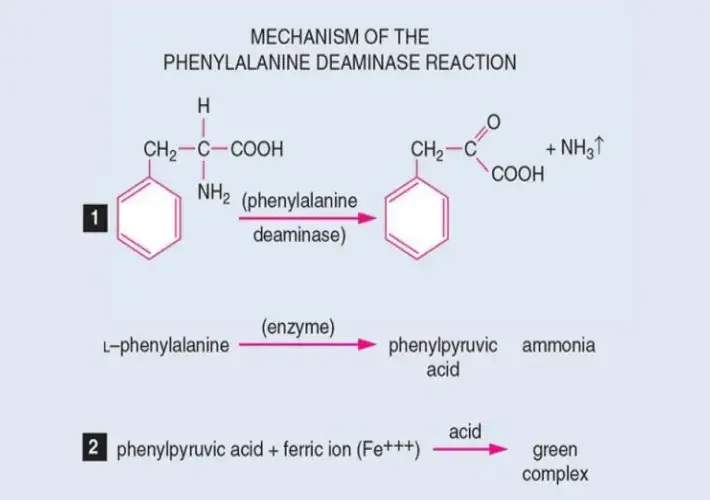
In this test, the organism is grown on a medium containing phenylalanine. If the enzyme is present, phenylalanine is deaminated and phenylpyruvic acid and ammonia is formed.
After incubation, 10% ferric chloride reagent is added. Phenylpyruvic acid reacts with ferric chloride and a light to dark green colour is produced immediately, this is a positive test.
If phenylpyruvic acid is not formed, no green colour is seen. The colour remains yellow or amber of the reagent, this is a negative test.
Objectives of Phenylalanine Deaminase Test
- To determine the ability of an organism to produce the enzyme phenylalanine deaminase.
- To test whether the bacterium can oxidatively deaminate phenylalanine and form phenylpyruvic acid and ammonia.
- To differentiate Proteeae tribe (Proteus, Providencia and Morganella) from other members of Enterobacteriaceae.
- To learn the procedure of PDA test and to interpret the result after adding 10% ferric chloride reagent.
- To identify unknown microbial species in clinical microbiology by using this biochemical test result.
Principle of Phenylalanine Deaminase Test
Principle of Phenylalanine Deaminase Test is based on the ability of some bacteria to produce the enzyme phenylalanine deaminase and utilize phenylalanine as a source of carbon and energy.
In this test, the organism is grown on a medium containing phenylalanine. The enzyme oxidatively removes the amine group from phenylalanine in presence of oxygen and phenylpyruvic acid, ammonia and water are formed.
Phenylpyruvic acid is colourless, so a developer is required. After incubation, 10% ferric chloride is added and it acts as a chelating agent.
If phenylpyruvic acid is present, it reacts with ferric ions and a light to deep green coloured complex is produced which indicates positive test. If the enzyme is absent, phenylpyruvic acid is not formed and no green colour appears, it remains yellow or straw colour of the reagent.
Requirements of Phenylalanine Deaminase Test
- Phenylalanine agar slants (DL-phenylalanine containing medium).
- Phenylalanine deaminase tablets or Urea-PDA disks (rapid method) with sterile distilled water or sterile saline.
- 10% Ferric chloride (FeCl3) solution (developer reagent).
- Hydrochloric acid (HCl) (0.1N or 1N) (used in some methods for acidification).
- Incubator (35–37°C) with aerobic condition.
- Inoculating loop or sterile swab.
- Sterile test tubes.
- Dropper or Pasteur pipette.
- Bunsen burner or incinerator.
- PPE and cleaning items (gloves, alcohol wipes, forceps).
- Positive control strain- Proteus mirabilis / Proteus vulgaris / Proteus hauseri.
- Negative control strain- Escherichia coli.
Procedure of Phenylalanine Deaminase Test
- Take a pure culture of the test organism (18 to 24 hours culture).
- With a sterile inoculating loop, pick a heavy inoculum from the culture.
- Inoculate a phenylalanine agar slant by streaking the surface in zigzag or fishtail pattern. Only the slant surface is streaked, butt is not stabbed.
- Keep the cap loosened for air exchange, because oxygen is required for the reaction.
- Incubate the tube aerobically at 33–37°C (generally 35°C) for 18 to 24 hours. If very heavy inoculum is used, 4 to 6 hours incubation may also give result.
- After incubation, add 4 to 5 drops of 10% ferric chloride solution directly on the slant.
- Gently rotate or agitate the tube so that reagent contacts with the growth and some surface colonies are dislodged.
- Observe the colour change within 1 to 5 minutes. Light to dark green colour indicates positive test. If it remains yellow or amber colour of the reagent, it indicates negative test.

Quality control strains
Positive Control Organisms (Produce a green color):
- Proteus mirabilis (e.g., ATCC 12453)
- Proteus vulgaris (e.g., ATCC 13315, ATCC 8427, ATCC 6380)
- Providencia alcalifaciens (e.g., ATCC 9886)
Negative Control Organisms (Remain yellow with no color change):
- Escherichia coli (e.g., ATCC 25922)
- Enterobacter aerogenes / Klebsiella aerogenes (e.g., ATCC 13048)
Result of Phenylalanine Deaminase Test
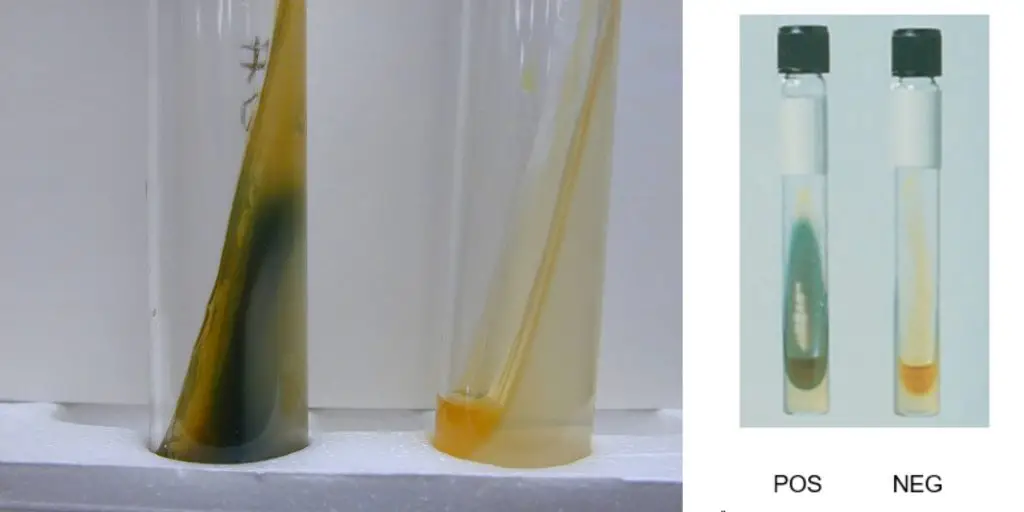
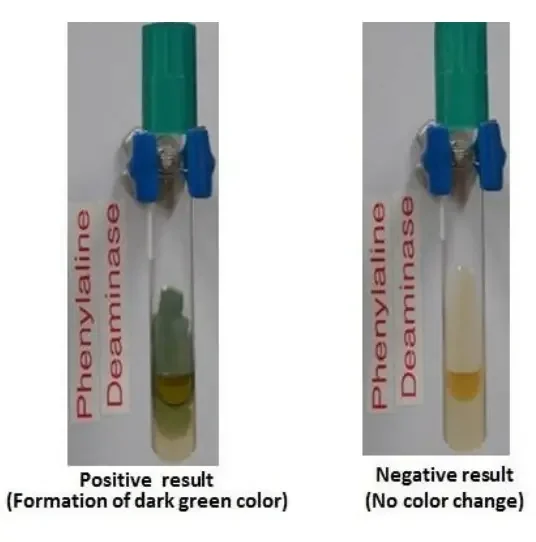
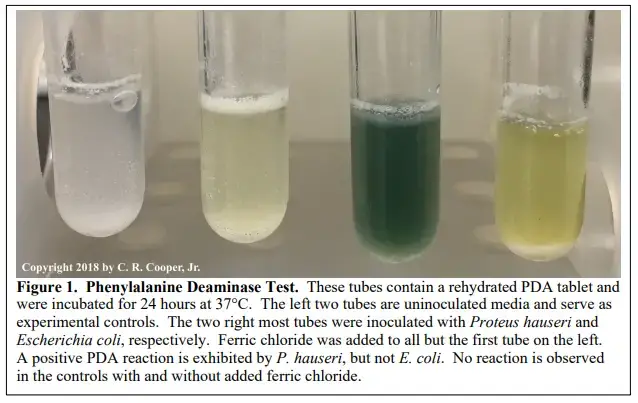
Positive result
Light to dark green colour develops on the surface of the medium within 1 to 5 minutes after adding 10% ferric chloride reagent.
It indicates phenylpyruvic acid is formed and the organism produces phenylalanine deaminase enzyme.
Proteus, Providencia and Morganella species usually gives positive test.
Negative result
No green colour is produced after adding 10% ferric chloride reagent.
The medium remains yellow or amber colour of the reagent.
It indicates phenylpyruvic acid is not formed and the organism does not produce phenylalanine deaminase enzyme.
Escherichia coli and most other Enterobacteriaceae usually gives negative test.



List of organisms for Phenylalanine Deaminase Test
Positive result (Green colour)
- Proteus mirabilis.
- Proteus vulgaris.
- Proteus penneri.
- Proteus hauseri.
- Proteus myxofaciens.
- Providencia alcalifaciens.
- Providencia stuartii.
- Providencia rettgeri.
- Providencia rustigianii.
- Providencia heimbachae.
- Morganella morganii.
- Enterobacter agglomerans (Pantoea agglomerans).
- Few other non-fermenting gram-negative bacilli and some Enterobacter strains.
Negative result (No green colour, remains yellow)
- Escherichia coli.
- Klebsiella pneumoniae.
- Klebsiella aerogenes.
- Yersinia pestis.
- Yersinia enterocolitica.
- Yersinia pseudotuberculosis.
- Salmonella species.
- Shigella species.
- Citrobacter species.
- Most Enterobacter species.
- Most other members of Enterobacteriaceae family.
Uses of Phenylalanine Deaminase Test
- It is used to detect phenylalanine deaminase enzyme production by a microorganism.
- It is used to identify bacteria that oxidatively deaminates phenylalanine and forms phenylpyruvic acid.
- It is used to differentiate gram-negative enteric bacilli of Enterobacteriaceae based on metabolic reaction.
- It is used to separate Proteeae tribe (Proteus, Providencia and Morganella) from most other Enterobacteriaceae.
- It is used as a biochemical screening test in clinical microbiology for identification of unknown isolates from patient samples.
- It is used to determine the ability of bacteria to utilize phenylalanine as a carbon and energy source.
Advantages of Phenylalanine Deaminase Test
- It is a highly specific test for detection of phenylalanine deamination in gram-negative bacilli.
- It is used to reliably identify and separate Proteeae tribe (Proteus, Providencia and Morganella) from other Enterobacteriaceae.
- It prevents misidentification, because Proteeae are non-lactose fermenters and can be confused with Salmonella and Shigella in primary screening, PDA test helps to differentiate them.
- It gives clear visual result. Light to dark green colour is produced immediately after adding ferric chloride reagent.
- It is a simple and rapid biochemical test and interpretation is easy.
- It can be adapted into rapid methods using tablets or combined assays (urea-phenylalanine) and result can be obtained within 2 to 6 hours in some rapid formats.
Limitations of Phenylalanine Deaminase Test
- It is not a confirmatory test. It cannot identify an unknown organism alone and other biochemical or molecular tests are also required.
- The green colour reaction is unstable. The colour starts fading quickly after formation.
- Result must be read within 1 to 5 minutes after adding 10% ferric chloride reagent, otherwise false negative result can occur.
- Heavy inoculum is required. If light inoculum is taken, phenylpyruvic acid may not be produced in sufficient amount and delayed or false negative result may be seen.
- Ferric chloride reagent can also react in tryptophan deaminase (TDA) test and gives brown or reddish-brown colour. This colour can be misinterpreted if not observed properly.
Precautions of Phenylalanine Deaminase Test
- A fresh pure culture (18–24 hours) should be used and heavy inoculum should be taken. Light inoculum may give delayed or false negative result.
- After adding 10% ferric chloride reagent, the result should be read within 1 to 5 minutes. Green colour fades quickly and late reading can cause false negative.
- Ferric chloride reagent should be stored properly in dark bottle and kept away from heat and light, otherwise reagent may deteriorate.
- After adding ferric chloride drops, the tube should be gently rolled or agitated so that reagent contacts the growth properly.
- If rapid urea-PDA disks or tablets are used, the disks should be checked for discoloration before use.
- Brown or reddish-brown colour should not be taken as positive PDA. Brown colour indicates tryptophan deaminase (TDA) activity, PDA positive must be green.
- PDA test should not be used as a standalone test for identification. It should be used along with other biochemical or molecular tests.
References
- Acharya, S., Kushwaha, P., Desai, S., & Chaubey, G. (2025). Proteus mirabilis entails a switch from commensal to pathogen – Genomic insights from a blaTEM-1B-harboring novel isolate from India [Preprint].
- Admin. (2015, February 8). Enterobacteriaceae. In Basicmedical Key / Clinical Gate.
- BD Biosciences. Phenylalanine agar ferric chloride reagent. In Difco & BBL Manual (2nd ed.).
- Benek, A., Canlı, K., & Altuner, E. M. (2026). Determination of biological activity and biochemical composition of Polytrichastrum formosum (Hedw.) G.L.Sm..
- Chemistry Stack Exchange. (2021). Why is it advised to use neutral ferric chloride solution while performing confirmatory test for acetate ions?.
- Condalab. Phenylalanine agar (Cat. 1040) [Technical data sheet].
- Cooper, C. R., Jr. (2018). BIOL 3702 lab exercise – Phenylalanine deamination test. Youngstown State University.
- Darland, G., Ewing, W. H., & Davis, B. R. (1975). The biochemical characteristics of Yersinia enterocolitica and Yersinia pseudotuberculosis. U.S. Department of Health, Education, and Welfare, Public Health Service, Center for Disease Control.
- Ehrenkranz, N. J., Alfonso, B. C., Eckert, D. G., & Moskowitz, L. B. (1989). Proteeae species bacteriuria accompanying Proteeae species groin skin carriage in geriatric outpatients. Journal of Clinical Microbiology.
- Hadi, A. M. KLEBSIELLA Proteus, Morganella and Providencia. Al-Mustaqbal University.
- Hardy Diagnostics. (2020). Ferric chloride reagent, 10% [Instructions for Use].
- Hardy Diagnostics. (2020). Phenylalanine agar [Instructions for Use].
- Hardy Diagnostics. (2020). Phenylalanine deaminase tablets [Instructions for Use].
- HiMedia Laboratories. (2026). Phenyl alanine agar M281 [Technical Data].
- Jabur, S. G. (2025). Molecular investigation of Proteus vulgaris virulence genes from asymptomatic bacteriuria in pregnant women.
- Kbah, A. Q. A. (2026). Antibiotics sensitivity of bacteria types isolated from burns and wounds in Babylon Governorate. American Journal of Bioscience and Clinical Integrity.
- Kimemia, B. B., Musila, L. A., Langat, S. K., & Eyase, F. L. (2025). Characterization of Proteus mirabilis isolated from ticks collected in Isiolo and Kilifi Counties, Kenya.
- Microbiology in Pictures. (2015). Tests used in the identification of Yersinia enterocolitica bacteria.
- Middleton, J. A., Martin, B. C., Kidd, D., & Jenkins, S. N. (2025). Black soldier fly (Hermetia illucens) larvae residue (frass) influences bacterial more than fungal communities in field-grown crops and pastures.
- O’Hara, C. M., Brenner, F. W., & Miller, J. M. (2000). Classification, identification, and clinical significance of Proteus, Providencia, and Morganella. Clinical Microbiology Reviews, 13(4), 534–546. https://doi.org/10.1128/CMR.13.4.534-546.2000.
- O’Hara, C. M., Westbrook, G. L., & Miller, J. M. (1997). Evaluation of Vitek GNI+ and Becton Dickinson Microbiology Systems Crystal E/NF identification systems for identification of members of the family Enterobacteriaceae and other gram-negative, glucose-fermenting and non-glucose-fermenting bacilli. Journal of Clinical Microbiology.
- Root, R. K., Waldvogel, F., Corey, L., & Stamm, W. E. (Eds.). (1998). Clinical infectious diseases: A practical approach. Oxford University Press.
- Singh, S., Dubal, Z., Kumar, G. R., & Shivaraman, G. K. (2025). Genomic constellation of foodborne Proteus mirabilis isolates harboring AMR, virulence genes and comparative WGS analysis. Current Microbiology.
- Smith, M., & Selby, S. (2021). 3.11: Phenylalanine agar. In Microbiology for Allied Health Students: Lab Manual. Biology LibreTexts.
- Spach, D. H., & Liles, W. C. (1998). Antimicrobial therapy for bacterial diseases. In R. K. Root, F. Waldvogel, L. Corey, & W. E. Stamm (Eds.), Clinical infectious diseases: A practical approach. Oxford University Press.
- Thermo Fisher Scientific (Remel). (2010). Phenylalanine agar [Instructions for Use].
- Valencia-Toxqui, G., S, S., & Ramsey, J. (2025). Isolation and characterization of biofilm-disrupting proteus phage Premi.
- Virtual Microbiology Lab Simulator Software (VUMIE). (2022). Ferric chloride (10%) reagent.
- Virtual Microbiology Lab Simulator Software (VUMIE). (2022). Phenylalanine deaminase test.
- Yin, Z., Chen, X., Xiao, J., & Peng, L. (2026). Insights into novel diagnostic assay development, antimicrobial resistance, and pathogenicity in Proteus mirabilis through pan-genome analysis. Applied and Environmental Microbiology.