- The Polymerase Chain Reaction (PCR) is a laboratory technique that is used to amplify (make many copies of) a specific segment of DNA.
- A DNA template is required, from which the target DNA region is defined, so that primers (short single–stranded DNA oligonucleotides) can bind flanking the region
- Denaturation is performed by heating the DNA to separate its two strands.
- Annealing is done when temperature is lowered, so that primers hybridize to complementary sequences on each single‐stranded template.
- Extension is carried out by a thermostable DNA polymerase enzyme which adds nucleotides to the 3′ ends of primers, thereby synthesizing new DNA strands.
- Thermal cycling (repeated cycles of denaturation, annealing, extension) is carried out many times (e.g., 20-40 cycles) to produce exponential amplification of the target DNA segment.
- Taq polymerase (derived from Thermus aquaticus) is often used because of its ability to survive the high denaturation temperatures.
- The small amount of starting DNA is converted into millions/billions of copies, so that enough material becomes available for subsequent study (sequencing, diagnosis, cloning, etc.)
- The PCR process is sensitive which means contamination can lead to amplification of non‐target DNA, giving false positives.
- Specificity is dependent on proper primer design, correct annealing temperature, and integrity of polymerase & reagents.
Types of PCR
Here is the list of Types of PCR based on different categories;
- Standard PCR Types
- Conventional PCR – Basic endpoint PCR amplification
- High-fidelity PCR – Enhanced accuracy with proofreading polymerases
- Hot-start PCR – Prevents non-specific amplification at room temperature
- Fast-cycling PCR – Rapid amplification with shortened cycle times
- Touch down (TD) PCR – Gradually decreasing annealing temperatures
- Real-Time PCR Types
- Real-Time PCR (qPCR) – Quantitative PCR with real-time monitoring
- Digital PCR (dPCR) – Absolute quantification through sample partitioning
- High-Resolution Melt (HRM) PCR – Real-time PCR with melting curve analysis
- RNA-Based PCR Types
- Reverse-Transcriptase PCR (RT-PCR) – RNA to cDNA conversion and amplification
- RT-qPCR – Real-time quantitative reverse transcription PCR
- RNase H-dependent PCR (rhPCR) – RNA-specific amplification method
- Multiplex and Complex PCR Types
- Multiplex PCR – Simultaneous amplification of multiple targets
- Nested PCR – Two-round amplification for enhanced specificity
- Long-range PCR – Amplification of large DNA fragments (>5kb)
- Assembly PCR – Synthetic gene construction method
- Overlap extension PCR – Site-directed mutagenesis and gene fusion
- Genetic Analysis PCR Types
- Allele-specific PCR – Discriminates between allelic variants
- AFLP PCR – Amplified fragment length polymorphism analysis
- VNTR PCR – Variable number tandem repeat analysis
- ISSR PCR – Inter-simple sequence repeat amplification
- Methylation-specific PCR (MSP) – Detects DNA methylation patterns
- SSP-PCR – Single specific primer PCR for HLA typing
- Specialized Application PCR Types
- Colony PCR – Direct amplification from bacterial colonies
- Single cell PCR – Amplification from individual cells
- In situ PCR – Amplification within tissue sections
- Solid phase PCR – Surface-bound amplification
- COLD PCR – Preferential amplification of minority variants
- Advanced PCR Types
- Asymmetric PCR – Unequal primer concentrations for single-strand products
- LATE PCR – Linear after exponential amplification
- Inverse PCR – Amplification of unknown flanking sequences
- TAIL-PCR – Thermal asymmetric interlaced PCR for genome walking
- Ligation-mediated PCR – Unknown sequence amplification with adaptors
- Repetitive Element PCR Types
- Alu PCR – Amplification between Alu repetitive elements
- Repetitive sequence-based PCR – Uses repetitive DNA as primer sites
- Emerging PCR Types
- Nanoparticle-Assisted PCR (nanoPCR) – Enhanced efficiency with nanoparticles
- Miniprimer PCR – Ultra-short primer amplification
- Suicide PCR – Self-limiting amplification system48
1. Conventional PCR
Conventional PCR is the basic type of PCR in which target DNA is amplified by repeated temperature cycles. It is also called end-point PCR or traditional PCR.
In this method, detection of amplified DNA is done only after completion of all cycles. So the result is observed at the end of reaction.
The following are the important points of Conventional PCR–
- Conventional PCR requires DNA template, forward and reverse primers, dNTPs, buffer, Mg²⁺ and thermostable DNA polymerase.
- The reaction is carried out in a thermal cycler. It changes temperature according to the programmed steps.
- The process includes denaturation, annealing and extension. These three steps are repeated for about 25 to 40 cycles.
- In denaturation, the double stranded DNA is separated into single stranded DNA by high temperature.
- In annealing, the primers bind with complementary sequence of the template DNA.
- In extension, Taq DNA polymerase adds nucleotides and forms new DNA strand.
- After amplification, the PCR product or amplicon is analysed by agarose gel electrophoresis.
- In gel electrophoresis, the amplified DNA is separated on agarose gel. It is commonly stained by ethidium bromide and viewed under UV light.
- The specificity of Conventional PCR depends on primer design, annealing temperature, quality of reagents and cycle condition.
- The sensitivity of this method is moderate. It can detect small amount of DNA, but accurate quantification is not possible.
- The main equipments are conventional thermal cycler and electrophoresis apparatus.
- Non-specific amplification and primer-dimer may occur in this method. It may give unwanted bands on gel.
- Plateau effect may occur after many cycles. In this condition, reaction efficiency decreases and amplification does not increase properly.
- There is also risk of contamination during post-PCR handling because tubes are opened for gel analysis.
- Conventional PCR is used in cloning, mutation detection, presence or absence test, and preparation of DNA for sequencing.
2. High-fidelity PCR
High-Fidelity PCR is a type of PCR in which highly accurate DNA polymerase enzyme is used. It reduces the wrong base addition during DNA synthesis.
The following are the important points of High-Fidelity PCR–
- In High-Fidelity PCR, DNA polymerase with low error rate is used. So misincorporation of wrong nucleotide is minimized.
- Many high-fidelity polymerases have 3′→5′ exonuclease activity. This is also called proofreading activity.
- By proofreading activity, the wrongly added nucleotide is removed before further extension of DNA strand.
- Its error rate is much lower than non-proofreading enzyme like Taq polymerase.
- Some high-fidelity enzymes show about 50 to more than 200 times better accuracy than normal Taq polymerase.
- The fidelity of the reaction depends on buffer composition, Mg²⁺ concentration, dNTP concentration, annealing temperature and cycle condition.
- Longer DNA targets are more difficult to amplify correctly. So enzyme selection and number of cycles should be controlled.
- High-Fidelity PCR is useful when the amplified DNA is used for cloning, sequencing and mutation detection.
- It reduces unwanted mutation in the PCR product. So the amplified product becomes more reliable for further work.
- The fidelity of polymerase can be measured by lacI mutation assay, blue-white screening and Next Generation Sequencing (NGS) based method.
- Some commercial high-fidelity polymerases are Q5 polymerase, Phusion polymerase and iProof polymerase.
- iProof polymerase has very low error rate, about 4.4 × 10⁻⁷ in HF buffer. It is around 50-fold lower than Thermus aquaticus Taq polymerase.
- The main limitation is that it is more costly than normal PCR enzyme.
- The reaction may be slower and sometimes needs longer extension time.
- It is more sensitive to reaction components. So small change in reagent concentration may affect the amplification.
- Very low level errors may still occur. It is more possible when cycle number is high or starting template amount is very low.
3. Hot-Start PCR
Hot-Start PCR is a modified type of PCR in which DNA polymerase enzyme remains inactive at low temperature. It becomes active only after high temperature is reached.
The following are the important points of Hot-Start PCR–
- In Hot-Start PCR, the activity of DNA polymerase is suppressed before the first denaturation step.
- The enzyme or other important component is blocked at room temperature. So unwanted amplification does not start during reaction preparation.
- It prevents wrong binding of primers with non-target sequence. It also reduces formation of primer-dimer.
- The polymerase may be blocked by chemical modification. It may also be blocked by antibody or aptamer which bind with the active site of enzyme.
- In some method, modified dNTPs or modified primers are used. In some protocol, polymerase is added only after heating.
- The enzyme becomes active by heat. Generally this activation occurs during first denaturation at about 95°C.
- At high temperature, the blocking agent is removed or denatured. Then DNA polymerase starts synthesis of new DNA strand.
- Hot-Start PCR increases specificity of the reaction. It prevents non-specific amplification before cycling starts.
- It improves yield of correct PCR product or amplicon. Background unwanted bands are reduced.
- The sensitivity of detection may increase because primer-dimer and misprimed products are less formed.
- Reaction setup can be done at room temperature more easily. The reagents remain inactive until heat activation.
- The main limitation is that activation time is needed. The first denaturation step may be longer.
- It is more costly than ordinary PCR because special enzyme, antibody or modified reagent may be required.
- Some enzyme system may show small low temperature activity. This is called leakiness and may still give low level non-specific product.
4. Fast-cycling PCR
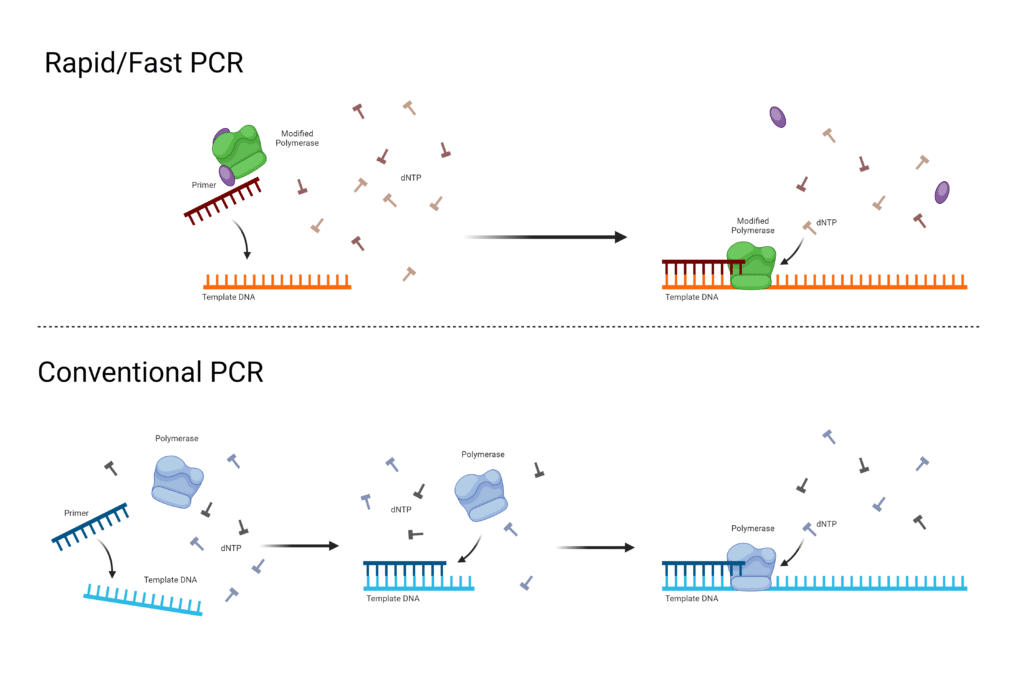
Fast-Cycling PCR is a type of PCR in which the time of denaturation, annealing and extension steps are shortened. It is used to complete the PCR reaction in less time than conventional PCR.
The following are the important points of Fast-Cycling PCR–
- In Fast-Cycling PCR, the holding time of each temperature step is reduced. So the total cycling time becomes short.
- Special DNA polymerase enzyme is used in this method. These enzymes can add nucleotides rapidly during extension step.
- High processivity polymerase is required. It helps in faster synthesis of new DNA strand.
- Fast thermal cycler is used in this method. It has rapid heating and cooling capacity.
- The fast ramp rate of machine reduces the time needed for changing one temperature to another temperature.
- In some protocol, annealing and extension steps are combined together. This is called two-step PCR.
- The time for denaturation, annealing and extension are kept only for minimum effective duration.
- Short amplicons are more suitable for Fast-Cycling PCR. Longer DNA fragments may need more time and special optimization.
- Primer design, buffer composition and polymerase selection are important in this method.
- If condition is properly optimized, specificity and sensitivity of the reaction are maintained.
- Fast-Cycling PCR can complete the reaction within tens of minutes. It is faster than conventional PCR which may take 1 to 2 hours.
- The main limitation is that all enzymes cannot work properly in shortened time.
- Complex DNA template, high GC content and secondary structure may reduce the efficiency of this method.
- Poorly designed primers may give non-specific products. So more optimization is needed in Fast-Cycling PCR.
5. Touch down (TD) PCR
Touchdown PCR (TD-PCR) is a modified type of PCR in which annealing temperature is kept high at first. Then the annealing temperature is slowly decreased in later cycles.
The following are the important points of Touchdown PCR–
- In Touchdown PCR, high annealing temperature is used in early cycles. It increases specific binding of primers with target DNA.
- The starting annealing temperature is usually 5-10°C above the melting temperature (Tm) of primers.
- At this high temperature, only perfectly matched primer-template binding takes place. Non-specific binding is reduced.
- In next cycles, the annealing temperature is decreased step by step. It may be decreased by 1-2°C in each cycle.
- The temperature is reduced until a lower suitable annealing temperature is reached.
- After the touchdown phase, normal PCR cycles are carried out at the final annealing temperature.
- The correct DNA product formed in early cycles acts as template in later cycles.
- The specific product increases more than non-specific product. So unwanted PCR products are reduced.
- Touchdown PCR improves specificity of amplification. It is useful when non-specific bands are formed in regular PCR.
- It can also increase sensitivity and yield when target DNA is present in low amount.
- TD-PCR is useful for difficult templates like high GC content DNA and templates having secondary structure.
- It is also useful when primers are not fully optimized.
- The main limitation is that protocol setting is more complex. Annealing temperature has to be changed in different cycles.
- More optimization may be needed for proper result.
- If the starting annealing temperature is too high, primer binding may not occur properly. So yield may decrease in early cycles.
6. Real-Time PCR (qPCR)
Real-Time PCR is a type of PCR in which amplification of DNA is detected during the reaction. It is also called quantitative PCR (qPCR).
The following are the important points of Real-Time PCR–
- In Real-Time PCR, the amplified DNA is measured after each cycle. It is not detected only at the end of reaction.
- A fluorescent reporter molecule is added in the reaction mixture. The fluorescence increases when the amount of PCR product increases.
- The fluorescence signal is detected by a special Real-Time PCR machine. It contains fluorescence detector with thermal cycler.
- The reaction is done in closed tube. So post-PCR handling is less and contamination risk is reduced.
- Real-Time PCR is used for quantification of starting DNA amount. When RNA is used, it is first converted into cDNA and then quantified.
- Two common detection methods are used in qPCR. These are SYBR Green dye method and TaqMan probe method.
- SYBR Green binds with all double stranded DNA. So fluorescence is produced when amplified DNA is formed.
- TaqMan probe is sequence specific probe. It gives fluorescence only when it binds with its target sequence.
- In qPCR, amplification curve is obtained. In first part, product increases exponentially. Later the reaction reaches plateau phase when reagents become limited.
- Ct or Cq value is the cycle number where fluorescence crosses the threshold level. It is used for calculation of target amount.
- Relative quantification is done by comparing target gene with reference or housekeeping gene.
- Absolute quantification is done by comparing the sample with standard curve prepared from known amount of template.
- Real-Time PCR is used for gene expression study. It is also used for detection and quantification of pathogens.
- It is used for copy number analysis, genetic variant detection and biomarker study.
- The main limitation is that reagents and machine are costly than conventional PCR.
- Primer and probe design should be done carefully. Poor design may give wrong signal.
- Reaction efficiency must be checked properly. Inhibitors may affect fluorescence and DNA polymerase activity.
- Quantification may be incorrect if amplification efficiency is not proper.
7. Digital PCR (dPCR)
Digital PCR (dPCR) is a modified type of PCR in which absolute amount of DNA or RNA is measured. In this method, the reaction mixture is divided into many small reactions.
The following are the important points of Digital PCR–
- In Digital PCR, the sample is partitioned into thousands or many small partitions. These partitions may be droplets, wells or chambers.
- Each partition may contain zero, one or more target nucleic acid molecule.
- Amplification occurs separately in each partition. After amplification, each partition is detected as positive or negative.
- Positive partition means target molecule is present. Negative partition means target molecule is absent.
- This positive and negative result is called binary read-out.
- Poisson statistical model is used in dPCR. It corrects the result when more than one target molecule is present in one partition.
- By this calculation, absolute copy number or concentration of target nucleic acid is measured.
- Digital PCR does not need standard curve like qPCR. The quantification is done from number of positive and negative partitions.
- The reaction uses similar components like qPCR. These are primers, probes and master mix.
- The main difference is partitioning of sample and end-point detection.
- Digital PCR has high sensitivity. It can detect rare mutation and low amount target sequence.
- It is useful when target DNA is present with high amount of background normal DNA.
- It has more tolerance to inhibitors. Partitioning dilutes the inhibitors in different small reactions.
- It gives more precise result because many small reactions are analysed separately.
- The main limitation is that the instrument and consumables are costly.
- Throughput may be lower than some other PCR methods.
- Careful optimization is needed for partitioning, amplification and detection.
8. High-Resolution Melt (HRM) PCR
High-Resolution Melt PCR (HRM PCR) is a PCR based method in which melting nature of double stranded DNA is studied. It is used to detect small changes in DNA sequence.
The following are the important points of HRM PCR–
- In HRM PCR, the target DNA region is first amplified by Real-Time PCR.
- A saturating DNA-binding fluorescent dye is added in the reaction. The dye binds with double stranded DNA and gives fluorescence.
- After amplification, the PCR product is slowly heated. This is called melt phase.
- During heating, double stranded DNA separates into single stranded DNA.
- When DNA melts, the dye is released and fluorescence signal decreases.
- The temperature is increased in very small steps, about 0.1-0.3°C. So the melting curve is recorded very clearly.
- The melting curve shows how the DNA amplicon melts with increasing temperature.
- The curve is then normalized and difference curve may be prepared.
- HRM PCR can distinguish homoduplex DNA, heteroduplex DNA, mutant sequence and wild type sequence.
- The melting temperature (Tm) and curve shape depend on GC content, length of amplicon, base composition and mismatch in DNA.
- Even a single base change may alter the melting curve. So HRM PCR is useful for mutation detection.
- HRM PCR is used for mutation scanning and Single Nucleotide Polymorphism (SNP) genotyping.
- It is also used in methylation analysis, zygosity testing and identification of species or strain.
- This method is cheaper than probe based genotyping for screening of many samples.
- Post-PCR handling is less because analysis is done in closed tube.
- The main limitation is that good quality DNA is required.
- Primers should be properly designed and the amplicon should be small or medium size.
- The instrument must have high temperature precision and suitable fluorescent dye.
- Some small sequence change may still need confirmation by sequencing or other method.
9. Reverse-Transcriptase PCR (RT-PCR)
Reverse-Transcriptase PCR (RT-PCR) is a type of PCR in which RNA is first converted into complementary DNA (cDNA). Then this cDNA is amplified by PCR.
The following are the important points of RT-PCR–
- RT-PCR is used when RNA is the starting material.
- In this method, reverse transcriptase enzyme converts RNA into cDNA.
- The formed cDNA then acts as template for PCR amplification.
- RT-PCR needs reverse transcriptase enzyme for reverse transcription step and DNA polymerase enzyme for amplification step.
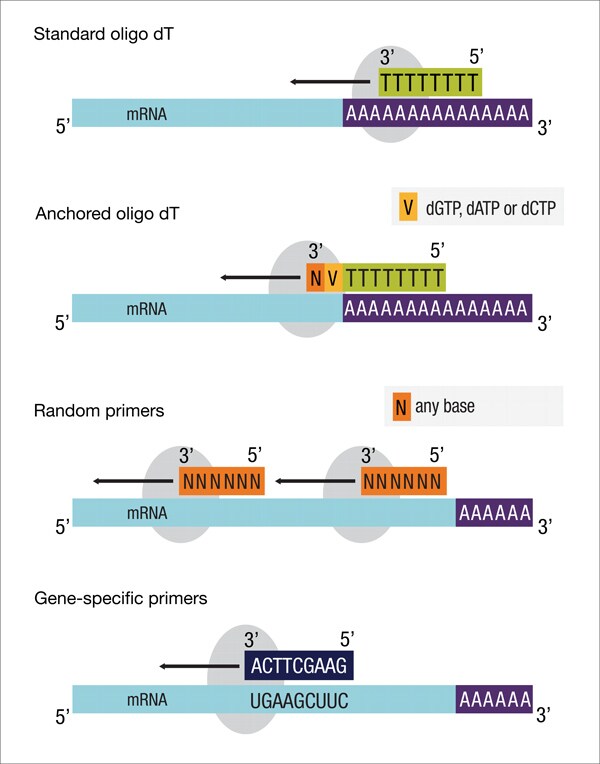
- Different types of primers may be used. These include target specific primers, oligo-dT primers and random primers.
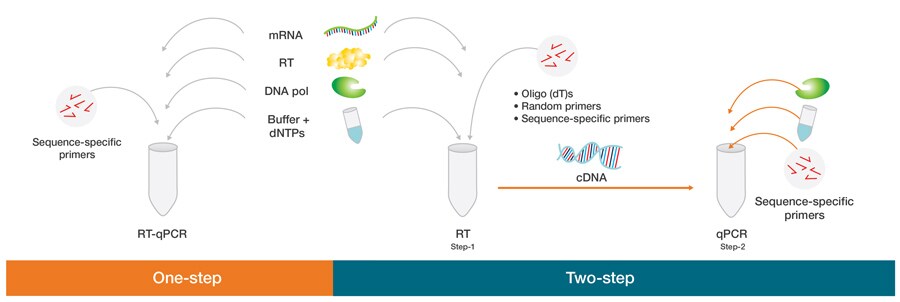
- One-step RT-PCR is a method where reverse transcription and PCR amplification are done in same tube.
- Two-step RT-PCR is a method where cDNA is first prepared from RNA. Then a part of this cDNA is used for PCR reaction.
- The amplified product may be detected by gel electrophoresis in end-point method.
- When RT-PCR is combined with Real-Time PCR, the product is detected during the reaction by fluorescent dye or probe.
- RT-PCR is highly sensitive method. It can detect low amount of RNA transcript from small sample.
- It is used for gene expression study. It is also used for detection of viral RNA.
- The specificity depends on primer design, enzyme quality, purity of RNA and temperature condition of reverse transcription step.
- RNA is easily degraded. So sample should be handled carefully.
- Genomic DNA contamination may give background signal.
- Reverse transcription step may show variation. So quantification may become incorrect if reaction efficiency is not proper.
10. RT-qPCR – Real-time quantitative reverse transcription PCR
RT-qPCR is a combined molecular method in which RNA is first converted into complementary DNA (cDNA). Then this cDNA is amplified and measured in real time by PCR.
The following are the important points of RT-qPCR–
- RT-qPCR is also called Reverse Transcription Quantitative Real-Time PCR.
- In this method, starting material is RNA. It may be total RNA or messenger RNA (mRNA).
- The RNA is first converted into cDNA by reverse transcriptase enzyme.
- The formed cDNA acts as template for quantitative Real-Time PCR.
- Fluorescent dye or probe is added in the reaction mixture. It gives signal when the amplified product is formed.
- The fluorescence is measured after each cycle. So quantification is done during amplification, not only at the end.
- One-step RT-qPCR is a method where reverse transcription and qPCR are done in same tube.
- In one-step method, handling is less. So contamination risk is also less.
- Two-step RT-qPCR is a method where cDNA is first prepared separately. Then a part of this cDNA is used for qPCR.
- In two-step method, the same cDNA can be used for many target genes. It also gives more flexibility in primer selection and optimization.
- RT-qPCR is highly sensitive method. It can detect low amount of RNA transcript.
- It can detect small changes in gene expression and viral RNA amount.
- It has wide dynamic range. So different amount of RNA can be quantified.
- Normalization is required in RT-qPCR. Reference or housekeeping genes are used for this purpose.
- Normalization controls variation in RNA amount, RNA quality and efficiency of reverse transcription.
- The main limitation is that RNA may degrade easily. So sample should be handled carefully.
- Both reverse transcriptase and DNA polymerase efficiency should be checked.
- Inhibitors may affect enzyme activity and give incorrect result.
- The reagents and instrument are more costly than simple PCR.
- RT-qPCR is used for gene expression study, viral RNA quantification, biomarker study, genetic testing and validation of microarray result.
11. RNase H-dependent PCR (rhPCR) – RNA-specific amplification method
RNase H-dependent PCR (rhPCR) is a modified type of PCR in which special blocked primers are used. The primer becomes active only after cleavage by RNase H2 enzyme.
The following are the important points of rhPCR–
- In rhPCR, the primers are blocked at their 3′ end. So DNA polymerase cannot extend the primer directly.
- These special primers are called rhPrimers. They contain one RNA base near the 3′ blocking group.
- The 3′ blocking group prevents extension by DNA polymerase before proper primer binding.
- A thermostable RNase H2 enzyme is added in the reaction mixture. It is often obtained from Pyrococcus abyssi.
- When the rhPrimer binds correctly with the template DNA, an RNA:DNA heteroduplex is formed.
- RNase H2 recognizes this RNA:DNA heteroduplex. It cuts at the 5′ side of the RNA base.
- After cleavage, the blocking group is removed. Then DNA polymerase can extend the primer and new DNA strand is formed.
- If the primer does not bind properly, the block remains present. So wrong amplification is reduced.
- rhPCR strongly reduces primer-dimer formation and mis-priming.
- This method increases specificity of the PCR reaction. Background products and artifacts are less formed.
- RNase H2 should be heat stable because it has to tolerate high temperature of PCR cycles.
- Its low activity at room temperature gives hot-start like effect. So unwanted amplification during reaction setup is reduced.
- Mismatch near the RNA base reduces cleavage by RNase H2. This helps in discrimination of SNPs and variant sequence.
- rhPCR is used for SNP genotyping and detection of rare alleles.
- It is useful in multiplex PCR because many primer pairs can be used with less primer interaction.
- It is also used in environmental DNA species discrimination and sequencing of immune receptor or TCR regions.
- The main limitation is that rhPrimer design should be proper. The position of RNA base and blocker stability are important.
- Enzyme concentration and buffer composition need optimization.
- Some leakage may occur if blocking group is removed non-specifically.
- The cost is higher than simple PCR because special primers and enzyme are required.
12. Multiplex PCR – Simultaneous amplification of multiple targets
Multiplex PCR is a type of PCR in which many target DNA sequences are amplified in one reaction tube. It uses more than one primer pair in the same reaction.
The following are the important points of Multiplex PCR–
- In Multiplex PCR, several target DNA sequences are amplified at the same time.
- Multiple primer pairs are used in one reaction mixture.
- Primer design is very important in this method. All primers should work at nearly same annealing temperature.
- Primer pairs should not bind with each other. It prevents primer-dimer and unwanted products.
- The size of different amplicons should be different enough. Then the products can be separated by gel electrophoresis.
- In fluorescent method, different probes or fluorescent labels may be used. It helps to identify different amplified products.
- Multiplex PCR is used for detection of many pathogens in one sample.
- It is used in SNP genotyping, mutation detection and polymorphism analysis.
- It is also used in forensic profiling and GMO detection.
- This method saves sample material. Small amount of DNA or RNA can be tested for many targets.
- It reduces reagent use and time because many reactions are done together.
- Multiplex PCR increases throughput. Many targets or many samples can be tested faster.
- Specificity and sensitivity may decrease if the reaction is not optimized properly.
- Competition may occur among different primer pairs. Some targets may amplify more and some may amplify less.
- Unequal amplification efficiency may reduce the yield of some products.
- Optimization is strongly required in Multiplex PCR.
- Primer concentration, annealing temperature, buffer and polymerase mixture should be adjusted properly.
- Each primer pair should be first checked separately and then checked in multiplex reaction.
- The initial setup takes more time and testing. But later it saves time, reagent and sample when many targets are tested routinely.
13. Nested PCR – Two-round amplification for enhanced specificity
Nested PCR is a modified type of PCR in which two rounds of amplification are done. It uses two sets of primers for increasing specificity and sensitivity.
The following are the important points of Nested PCR–
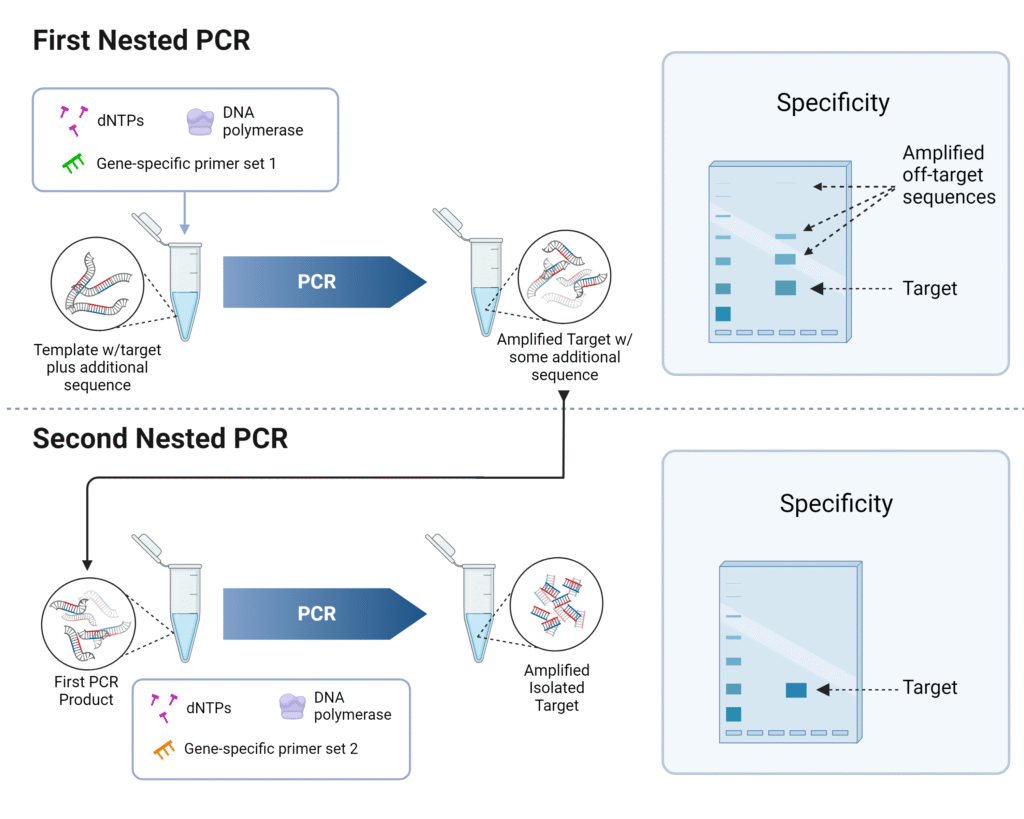
- In Nested PCR, first PCR is carried out with outer primers.
- The outer primers bind outside the target region. It amplifies a larger DNA fragment.
- The product of first PCR is then used as template for second PCR.
- In second PCR, inner or nested primers are used.
- These inner primers bind inside the first amplified PCR product.
- Only the correct first round product contains binding sites for both inner primers.
- Non-specific product formed in first PCR usually do not contain binding sites for inner primers.
- So unwanted amplification is reduced in second round.
- Nested PCR increases specificity of amplification.
- It also increases sensitivity because target DNA is amplified two times.
- Low copy number target sequence can be detected by this method.
- More total cycles are used in Nested PCR because two PCR rounds are performed.
- It needs proper knowledge of target sequence. Outer and inner primers should be designed correctly.
- Nested PCR is used for detection of rare pathogens.
- It is used for detection of low copy number genes.
- It is useful when sample contains high amount of background non-target DNA.
- It may be used for degraded DNA, such as formalin-fixed tissue sample.
- The main limitation is that it takes more time than simple PCR.
- It needs more reagents and extra primers.
- Contamination risk is higher because first PCR product is handled before second PCR.
- Optimization of reaction condition is more difficult in Nested PCR.
14. Long-range PCR – Amplification of large DNA fragments (>5kb)
Long-Range PCR is a special type of PCR in which large DNA fragments are amplified. It is used for amplification of DNA regions longer than standard PCR, usually more than 5 kb.
The following are the important points of Long-Range PCR–
- In Long-Range PCR, long DNA amplicons are produced.
- It can amplify large fragments like 6.6 kb, 13 kb, 20 kb or more.
- Standard PCR often fails to amplify such long DNA fragments.
- Special DNA polymerase or mixture of polymerases are used in this method.
- The enzyme should have high processivity. So it can synthesize long DNA strand continuously.
- The enzyme should also have high fidelity or proofreading activity. It reduces error during long DNA synthesis.
- Buffer composition is important in Long-Range PCR.
- Some additives or enhancers are added in reaction mixture. These reduce problem of secondary structure, GC-rich region and difficult template.
- Thermal cycling condition is changed from ordinary PCR.
- Longer extension time is used because long DNA strand need more time for synthesis.
- Sometimes lower extension temperature like 68°C is used instead of 72°C.
- Short denaturation time is used to reduce damage of long template DNA.
- Template DNA should be good quality and intact. Fragmented or damaged DNA cannot give proper long amplification.
- Primer design is more important in this method. Primers should be specific and their Tm should be well matched.
- Primers should not form secondary structure and should not bind with repeat or high GC region.
- Long-Range PCR is used for sequencing of large genomic region.
- It is used for study of structural variation.
- It is useful for cloning of long genes including introns.
- It is also used for detection of large deletion or insertion.
- It may be used for making long probes or long DNA templates.
- The main limitation is that yield may be low.
- Reaction failure is more common than ordinary PCR.
- Optimization takes more time.
- Cost of reagent and instrument may be higher.
15. Assembly PCR – Synthetic gene construction method
Assembly PCR is a type of PCR in which many short oligonucleotides are joined to form a long DNA molecule. It does not require complete natural DNA template.
The following are the important points of Assembly PCR–
- In Assembly PCR, many short single stranded oligonucleotides are used.
- These oligonucleotides have overlapping ends.
- The overlapping ends help the adjacent oligonucleotides to bind with each other in correct order.
- The overlap region is usually about 20 bp or near this size.
- During first cycles, the overlapping oligos anneal with each other.
- DNA polymerase extends the annealed oligos and forms partial DNA fragments.
- These partial fragments again anneal and extend in next cycles.
- In this way, longer DNA strands are gradually formed.
- After assembly cycles, outer primers are used.
- The outer primers bind at the two ends of the full desired sequence.
- Then normal PCR amplification is done.
- The full-length assembled DNA product is amplified more than short incomplete fragments.
- Assembly PCR is used to prepare synthetic genes.
- It can make DNA sequence of hundreds to thousands of base pairs.
- It is useful when natural DNA template is not available.
- The fidelity of Assembly PCR depends on quality of oligonucleotides, DNA polymerase, overlap design and error correction step.
- Wrong base, deletion or mismatch may occur during synthesis and assembly.
- Oligonucleotide concentration, annealing temperature and cycle number should be optimized.
- Low annealing temperature may cause wrong binding of oligos.
- Too many oligos may also cause mis-annealing and incomplete assembly.
- The main limitation is that long constructs are difficult to prepare.
- Secondary structure and incomplete overlap may reduce proper assembly.
- Cost increases when the size of DNA construct becomes large.
- It is more laborious than simple PCR using existing DNA template.
16. Overlap extension PCR – Site-directed mutagenesis and gene fusion
Overlap Extension PCR is a type of PCR in which two or more DNA fragments are joined by their overlapping ends. It is also called Splicing by Overlap Extension PCR (SOE-PCR).
The following are the important points of Overlap Extension PCR–
- In Overlap Extension PCR, separate DNA fragments are first amplified.
- The primers are designed in such a way that they add overlapping ends to the adjacent DNA fragments.
- These overlapping ends are complementary to each other.
- The fragments are then denatured and allowed to anneal through their overlap region.
- DNA polymerase extends from the annealed overlap region.
- As a result, fused hybrid DNA fragment is formed.
- After this, outer primers are used for amplification of full fused product.
- The overlapping overhangs are usually present at the 5′ end of primers.
- The overhangs should be long enough for proper annealing. It may be several nucleotides or dozens of nucleotides.
- High-fidelity DNA polymerase is preferred in this method.
- It reduces unwanted mutation during fusion and extension of DNA fragments.
- When many fragments are joined, the method is done in steps.
- First, individual fragments are amplified.
- Then overlap extension is done without outer primers.
- At last, final amplification is done with external primers.
- Overlap Extension PCR is used for construction of chimeric gene.
- It is used for insertion of desired mutation or tag into a gene.
- It is used for promoter and terminator fusion.
- It is also used for protein fusion and synthetic gene assembly.
- The main advantage is that restriction enzyme site is not required.
- It can produce scarless fusion product.
- It gives flexibility in designing fused DNA constructs.
- Large construct can also be prepared by this method when proper polymerase and condition are used.
- The main limitation is that overhang design should be precise.
- Concentration of fragments and primers should be balanced.
- Incomplete fusion or mis-annealing may occur.
- Annealing temperature and cycle number should be optimized properly.
17. Allele-specific PCR
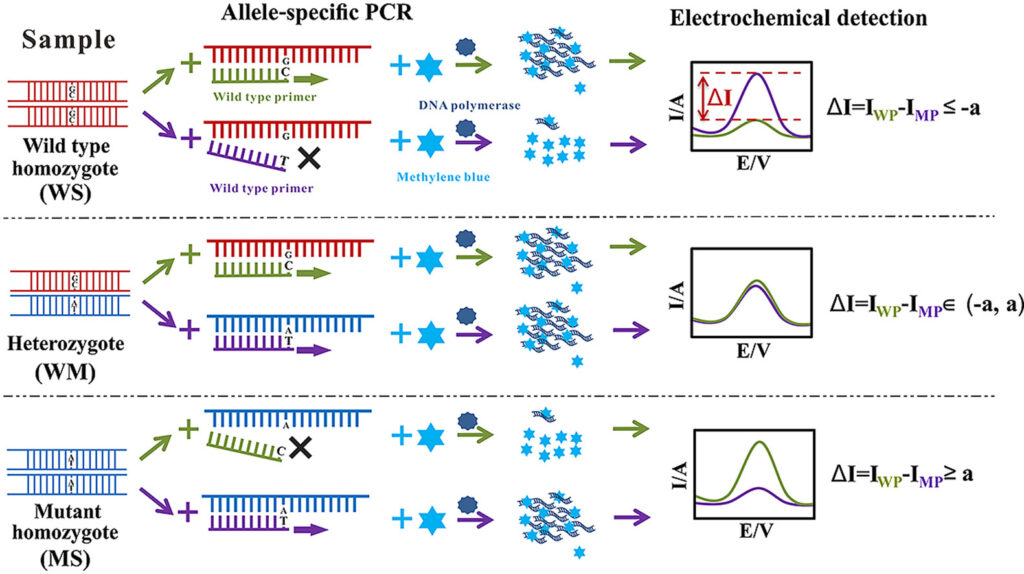
Allele-Specific PCR (AS-PCR) is a type of PCR in which specific allele or variant is detected by using allele specific primers. Amplification occurs only when the target allele is present.
The following are the important points of Allele-Specific PCR–
- In AS-PCR, the primer is designed in such a way that its 3′ end base matches with the selected SNP or mutation. If perfect matching is present, DNA polymerase extends the primer and amplification occurs.
- If the 3′ end of primer does not match with the template DNA, extension does not occur properly. So the non-matching allele is not amplified.
- Two allele specific primers may be used in this method. One primer is for wild type allele and another primer is for mutant allele, with one common primer.
- The reaction may be done in separate tubes or in multiplex form. The amplified product shows which allele is present in the sample.
- Specificity may be increased by adding another mismatch near the 3′ end of primer. This makes wrong primer-template binding unstable and reduces non-specific amplification.
- The amplified allele specific product is detected by gel electrophoresis. It may also be detected by fluorescence, probe or melting curve method in real-time system.
- Allele-Specific PCR has moderate to high sensitivity. It can detect variant allele even in low amount when reaction condition is optimized.
- The main limitation is that primer design should be very precise. Weak mismatch discrimination may give false positive or false negative result.
- Annealing temperature, primer concentration and buffer condition should be optimized properly. Otherwise non-specific amplification may occur.
- AS-PCR is used for SNP genotyping, point mutation detection, pharmacogenetics, diagnosis of genetic diseases and population genetics study.
18. AFLP PCR
AFLP PCR is a molecular fingerprinting method in which genomic DNA is cut by restriction enzymes, adaptor is ligated and selected fragments are amplified by PCR. It is also called Amplified Fragment Length Polymorphism PCR.
The following are the important points of AFLP PCR–
- In AFLP PCR, genomic DNA is first digested with restriction enzymes. Usually two enzymes are used, one rare cutter and one frequent cutter, which produce many DNA fragments of different size.
- Adaptors are then ligated to the ends of restriction fragments. These adaptors are known sequence of double stranded oligonucleotides and provide binding site for PCR primers.
- Selective amplification is done by using primers which bind with adaptor sequence, restriction site sequence and extra selective nucleotides at the 3′ end. These extra bases reduce the number of fragments to be amplified.
- The amplified fragments are separated by gel electrophoresis or capillary electrophoresis. After separation, band or peak pattern is obtained which acts as genetic fingerprint.
- The band pattern is scored as present or absent. Present band is written as 1 and absent band is written as 0, and this helps to compare individuals, strains or populations.
- AFLP PCR can detect many polymorphisms without much prior sequence information. So it is useful for organisms whose full genome sequence is not known.
- This method is sensitive and reproducible when good quality DNA, proper restriction enzymes and selected primers are used.
- The main limitation is that band scoring may be confusing in some cases. Some weak bands or overlapping fragments may be difficult to score.
- AFLP markers are mostly dominant markers. So it may not clearly separate heterozygous and homozygous condition.
- Reproducibility may vary between different gels, instruments or laboratories. Large fragment analysis may need high-throughput system.
- AFLP PCR is used in population genetics, strain fingerprinting, variety identification, phylogenetic study, genetic mapping and detection of genetic variation.
19. VNTR PCR
VNTR PCR is a PCR based technique in which Variable Number Tandem Repeat (VNTR) region of DNA is amplified. It is used to detect difference in number of repeat sequences.
The following are the important points of VNTR PCR–
- In VNTR PCR, the target region contains tandem repeat sequences. These repeats are present one after another in the DNA.
- The repeat unit of VNTR is usually longer than microsatellite. It may be about 10-100 base pairs.
- The number of repeat units may vary in different individuals or strains. So the size of amplified DNA fragment also becomes different.
- Primers are designed on the flanking region of VNTR. These primers amplify the repeat region with the nearby flanking sequence.
- The amplified product is separated by gel electrophoresis or capillary electrophoresis. The size of the band or peak shows the number of repeats.
- VNTR PCR gives length polymorphism. This polymorphism is used as genetic marker for comparison of individuals or strains.
- In forensic science, many VNTR loci are tested together. The combined pattern gives individual specific DNA fingerprint.
- The sensitivity and specificity depends on primer design, quality of DNA and PCR condition. The flanking region should be conserved for proper primer binding.
- Very large VNTR alleles may produce large PCR products. These products may be difficult to amplify and separate clearly.
- Stutter bands may be formed due to polymerase slippage during amplification. This may create difficulty in result interpretation.
- Good resolution system is required for correct sizing. High quality gel or capillary system gives better result.
- VNTR PCR is used in molecular typing of microorganisms, genetic diversity study, parentage analysis, pedigree analysis, human identification and epidemiological tracing.
20. ISSR PCR
ISSR PCR is a molecular marker technique in which simple sequence repeat (SSR) based primers are used. It amplifies the DNA regions present between two nearby microsatellite repeat loci.
The following are the important points of ISSR PCR–
- ISSR PCR is also called Inter-Simple Sequence Repeat PCR. In this method, primers bind with microsatellite repeats and amplify the region present between adjacent repeats.
- The primers are usually 16-25 base pairs long. They contain repeat motifs like di-nucleotide, tri-nucleotide or tetra-nucleotide repeats.
- Some primers are anchored at 3′ end or 5′ end with one or few non-repeat bases. This increases binding specificity of the primer.
- In one reaction, many inter-SSR regions may be amplified. The amplified fragments give many bands of different sizes.
- The products are separated by gel electrophoresis or capillary electrophoresis. The band pattern acts as multilocus fingerprint.
- ISSR markers show high polymorphism. This is because the inter-repeat regions vary in length and sequence among individuals or strains.
- The method is reproducible when good quality DNA, proper primer and optimized annealing temperature are used.
- ISSR PCR is cost effective method. It does not need full genome sequence information for primer design.
- Same ISSR primers may be used in related species. So it is useful in organisms where sequence information is less.
- The main limitation is that ISSR markers are mostly dominant markers. So heterozygous and homozygous condition may not be separated clearly.
- Large bands may amplify poorly. Faint band or smeared band may make scoring difficult.
- PCR condition should be optimized properly. Annealing temperature, primer concentration and cycle number affect the result.
- ISSR PCR is used in genetic diversity study, population structure analysis, phylogenetic relationship, cultivar identification, germplasm characterization, taxonomy and conservation genetics.
21. Methylation-specific PCR (MSP)
Methylation-Specific PCR (MSP) is a type of PCR in which methylation status of DNA is detected. It shows whether the target DNA region is methylated or unmethylated.
The following are the important points of MSP–
- In MSP, the DNA is first treated with bisulfite. This treatment changes unmethylated cytosine into uracil, which is read as thymine during PCR, but methylated cytosine (5-methylcytosine) remains unchanged.
- Two sets of primers are used in this method. One primer set is specific for methylated DNA sequence and another primer set is specific for unmethylated DNA sequence after bisulfite treatment.
- The amplification is done separately with each primer set. Only the template which perfectly matches with the primer is amplified.
- The PCR product is detected by gel electrophoresis. The presence or absence of band shows the methylation status of specific CpG sites or CpG islands.
- MSP is highly sensitive method. It can detect small amount of methylated DNA even when large amount of unmethylated DNA is present.
- Primer design is very important in MSP. Several CpG sites should be present in primer binding region, mainly near the 3′ end, for better discrimination.
- Proper bisulfite conversion is essential. Incomplete conversion of unmethylated cytosine may give false positive result as methylated DNA.
- The main limitation is that many MSP methods are qualitative. It mainly shows presence or absence, not exact amount.
- Mixed methylation pattern may give confusing result. It may be difficult to interpret in some sample.
- DNA may be degraded during bisulfite treatment. So yield of PCR product may be reduced.
- Primer designing in dense CpG rich region may be difficult.
- MSP is used for study of gene methylation, cancer epigenetics, gene silencing, biomarker detection and methylation analysis of CpG islands.
22. SSP-PCR
SSP-PCR is a type of PCR in which sequence specific primers are used. It amplifies only the exact matching allele or sequence variant.
The following are the important points of SSP-PCR–
- SSP-PCR is also called Sequence Specific Primer PCR. In this method, primers are designed for one particular allele or sequence, so amplification occurs only when that allele is present.
- The 3′ end of primer is very important in this method. If mismatch is present at this end, DNA polymerase cannot extend the primer properly and non-target allele is not amplified.
- SSP-PCR is commonly used in HLA typing. Many allele specific primer pairs are used and the presence or absence of band shows which HLA allele is present in the sample.
- In some cases, SSP-PCR is also referred as Single Specific Primer-PCR. It is used when only one end of DNA sequence is known and unknown adjacent region is amplified by genome walking.
- The reaction is carried out under stringent PCR condition. This condition reduces non-specific priming and allows amplification only from properly matched primer-template combination.
- The amplified product is generally detected by gel electrophoresis. Presence of band gives positive result and absence of band gives negative result for that allele.
- SSP-PCR has high sensitivity and specificity when primers are properly designed. So it is useful for allele discrimination and genetic typing.
- The main limitation is that many separate reactions or many primer pairs may be needed for testing several alleles.
- False positive or false negative result may occur when primer binding is not proper or reaction condition is not optimized.
- Primer design and validation takes more time and cost than simple PCR.
23. Colony PCR
Colony PCR is a screening PCR method in which bacterial or yeast colony is used directly as template source. It is used to check recombinant clones without plasmid purification.
The following are the important points of Colony PCR–
- In Colony PCR, a single colony is picked after transformation and plating. The colony is picked by sterile toothpick or pipette tip and transferred into PCR master mix.
- The bacterial cell is lysed during initial heating or denaturation step. Due to lysis, plasmid DNA or genomic DNA comes out and becomes available for amplification.
- Different types of primers can be used in this method. Insert specific primers, vector flanking primers or orientation specific primers are used to check presence, size and orientation of insert.
- The amplification is carried out by normal PCR cycles. Sometimes longer initial heating or adjusted annealing temperature is used because cell debris and impurities may affect the reaction.
- The PCR products are analysed by agarose gel electrophoresis. Colonies having correct insert give band of expected size.
- Colony PCR saves time and cost. It avoids plasmid miniprep and restriction digestion for checking many colonies.
- It is useful for rapid screening of positive recombinant colony after cloning experiment.
- The sensitivity and specificity is good when reaction is properly optimized.
- The main limitation is that false positive or false negative result may occur.
- Too much colony material may inhibit PCR because excess cell debris is present in reaction.
- Poor primer design may give non-specific band or no amplification. So primer design and colony amount should be controlled properly.
24. Single cell PCR
Single Cell PCR is a PCR technique in which genetic material from one single cell is amplified. It is used to study DNA or RNA of that one cell.
The following are the important points of Single Cell PCR–
- In Single Cell PCR, the starting template amount is very small. The template may be present in picogram level, so careful handling and sensitive reagent are required.
- The single cell is first isolated by micro-dissection, micromanipulation, fluorescence activated cell sorting (FACS) or other single cell capture method.
- The isolated cell is lysed to release its DNA or RNA. If RNA is used, it is first converted into complementary DNA (cDNA) by reverse transcription.
- Whole Genome Amplification (WGA) or Whole Transcriptome Amplification (WTA) may be done when many genes or many loci are studied from one cell.
- The amplification is carried out by PCR with optimized cycle number, primer concentration, polymerase type and reaction volume, so that allele loss and unequal amplification are reduced.
- Single-cell RT-qPCR is used when gene expression from one cell is measured. It detects RNA level and gives quantitative result.
- Single Cell PCR has very high sensitivity. It can detect rare transcript, low copy gene and mutation present in single cell.
- This method is useful because bulk cell PCR may hide the variation of one cell. Single cell study shows cell to cell difference.
- The main limitation is amplification bias. Some allele or sequence may amplify more than other.
- Allelic dropout may occur in this method. One allele may fail to amplify and result may become incomplete.
- Contamination risk is very high because the starting material is extremely small.
- Reproducibility and accuracy depend on very careful optimization of lysis, amplification and handling condition.
25. In situ PCR
In situ PCR is a molecular method in which specific DNA or RNA sequence is amplified inside intact cells or tissue section. It keeps the position of target nucleic acid in the cell or tissue.
The following are the important points of In situ PCR–
- In In situ PCR, amplification is done within fixed cells or tissue sections. So the anatomical location of the target DNA or RNA is preserved.
- The tissue is first fixed and permeabilized. Proteolytic digestion may be done, so that primers and DNA polymerase can enter inside the cells.
- The target sequence is amplified inside the cell or tissue section by PCR. This shows which cell or tissue region contain the target nucleic acid.
- The amplified product may be detected by labelled nucleotides during PCR. This is called direct In situ PCR.
- In indirect In situ PCR, the amplified product is detected later by hybridization with labelled probe.
- Detection may also be done by histochemical or immunohistochemical method.
- In situ RT-PCR is used when RNA target is detected. In this method, RNA is first converted into cDNA and then amplified inside the tissue section.
- In situ PCR has high sensitivity. It is useful for detection of low copy genes when fixation, permeabilization and amplification are properly controlled.
- In formalin-fixed paraffin-embedded tissue, nucleic acid may be damaged. So amplification may be poor and background signal may increase.
- PCR product may diffuse out from cells or tissue. This may give wrong localization of signal.
- Non-specific labelled nucleotide incorporation may occur in fragmented DNA. So negative control is needed.
- Quantification is difficult in In situ PCR. It mainly shows location of target sequence.
- The method is more tedious than ordinary PCR. Optimization of fixation, reagent entry and amplification is laborious.
- In situ PCR is used in diagnostic pathology, detection of viral infection in tissue, mapping of gene expression, developmental biology and detection of gene rearrangement or translocation.
26. Solid phase PCR
Solid-Phase PCR (SP-PCR) is a method in which one or both primers are attached to a solid surface. In this method, DNA amplification occurs with at least one primer fixed on the surface.
The following are the important points of Solid-Phase PCR–
- In Solid-Phase PCR, the primer is immobilized on solid support like glass, beads, microarray or chip surface. The primer is generally attached by its 5′ end, and the 3′ end remains free for extension by DNA polymerase.
- The attached primer must be stable during repeated heating and cooling cycles. So covalent linkage or strong binding chemistry is used for fixing the primer on the solid surface.
- During reaction, template DNA and other free primer anneal with the immobilized primer. Then DNA polymerase extends the primer and forms surface-bound amplified DNA product.
- The efficiency of SP-PCR is usually lower than conventional PCR. This is because movement of template, steric hindrance and reduced availability of immobilized primer affect the reaction.
- Specificity may be improved in this method. Since one primer is fixed on the surface, unwanted primer interaction and primer-dimer formation are reduced.
- The steps of denaturation, annealing and extension are adjusted according to the solid support. The surface and attached oligonucleotide should not be damaged during thermal cycling.
- Solid-Phase PCR is used in microarray based target detection, SNP detection, diagnostic platform and preparation of immobilized amplicons for sequencing.
- It is also useful where detection is needed directly on a surface. Washing and signal capture becomes easier in such system.
- The main limitation is low yield and slow amplification. More cycle number may be needed in comparison to normal PCR.
- The template may not easily reach the surface bound primer. So amplification may be reduced.
- Surface chemistry, primer density, primer orientation and cycling condition should be optimized properly.
27. COLD PCR
COLD PCR is a modified type of PCR in which low amount mutant or variant allele is enriched more than normal wild type allele. It is also called Co-amplification at Lower Denaturation Temperature PCR.
The following are the important points of COLD PCR–
- COLD PCR is based on the use of lower denaturation temperature. This temperature is called critical denaturation temperature (Tc), which is lower than the melting temperature (Tm) of the target DNA.
- At Tc, the mismatched heteroduplex DNA denatures more easily. But the perfectly matched wild type homoduplex DNA remains mostly double stranded.
- Due to selective denaturation, mutant or mismatched DNA becomes available for primer binding and amplification. So variant allele is amplified more in next cycles.
- COLD PCR helps to detect low level mutation in a mixture of normal and mutant DNA. The mutant allele may become more visible after amplification.
- There are two main formats of COLD PCR. These are Fast COLD PCR and Full COLD PCR.
- In Full COLD PCR, intermediate annealing and hybridization step is included. This helps in formation of heteroduplex DNA before selective denaturation.
- In Fast COLD PCR, some steps are skipped to make the reaction faster. But mutation enrichment may be less in some cases.
- Different types of mutation may be enriched by this method. It may include base substitution, small insertion and deletion when condition is optimized.
- The amplified products of COLD PCR are usually analysed by Sanger sequencing, pyrosequencing, High-Resolution Melting (HRM) or other mutation detection method.
- COLD PCR increases sensitivity of mutation detection. It may give about 10 to 100 fold enrichment of low abundance mutant allele.
- Proper control of Tc is very important. Even small change of about 0.3°C may reduce enrichment or allow wild type DNA to denature.
- The amplicon size should be kept short for good discrimination. Usually short target region gives better result.
- The main limitation is that optimization is needed for each target sequence. Some mutation may not change Tm enough for good enrichment.
- Polymerase error or poor amplification efficiency may affect the result.
- COLD PCR is used for detection of rare mutation, cancer mutation analysis, somatic mutation study and identification of low amount variant allele in clinical sample.
28. Asymmetric PCR
Asymmetric PCR is a type of PCR in which one strand of DNA is amplified more than the other strand. It is used to produce single stranded DNA (ssDNA).
The following are the important points of Asymmetric PCR–
- In Asymmetric PCR, two primers are used but in unequal concentration. One primer is added in high amount and it is called excess primer, while the other primer is added in low amount and it is called limiting primer.
- In early cycles, both strands of DNA are amplified like normal PCR. This gives exponential amplification until the limiting primer is used up.
- After the limiting primer is exhausted, only the strand made from excess primer is amplified. So the reaction becomes linear and single stranded DNA is formed.
- More cycles are needed in this method because linear amplification is slower than exponential amplification.
- Primer design is important in Asymmetric PCR. The melting temperature (Tm) of primers should be suitable, mainly for the limiting primer.
- Poor primer binding or mismatch reduces the yield of single stranded DNA.
- Primer ratio, cycle number, annealing temperature, template concentration and DNA polymerase should be optimized properly.
- LATE-PCR is an improved form of Asymmetric PCR. It is called Linear-After-The-Exponential PCR.
- In LATE-PCR, limiting primer has higher Tm than the excess primer. So amplification continues better even when limiting primer concentration becomes low.
- Asymmetric PCR is used for preparation of single stranded DNA probes, DNA sequencing, aptamer library preparation (SELEX), allele discrimination and microarray hybridization.
- The main limitation is that yield is lower than normal symmetric PCR.
- Reproducibility depends on exact primer ratio and reaction condition.
- More cycle number may produce non-specific products.
- The formed single stranded DNA may need purification before further use.
29. LATE PCR
LATE PCR is a modified form of Asymmetric PCR in which unequal concentration of primers are used. It is designed to produce single stranded DNA with better efficiency and specificity.
The following are the important points of LATE PCR–
- In LATE PCR, one primer is used in low concentration and it is called limiting primer. Another primer is used in high concentration and it is called excess primer.
- The limiting primer is designed with higher or equal melting temperature than excess primer. This concentration adjusted melting temperature helps the limiting primer to work properly even when its amount becomes low.
- In early cycles, both primers are present in the reaction. So both strands of DNA are amplified and double stranded DNA product is formed like normal PCR.
- After the limiting primer is depleted, the reaction changes into linear amplification. In this phase, mainly the strand primed by excess primer is produced.
- As a result, single stranded DNA (ssDNA) accumulates in the reaction mixture. This product can be used for probe binding or further analysis.
- Primer design is very important in LATE PCR. The Tm values should be adjusted according to primer concentration and the primer binding site should not form mispriming or secondary structure.
- LATE PCR gives more predictable amplification pattern than traditional Asymmetric PCR. The exponential phase occurs first and then the linear phase occurs.
- This method is useful in real-time detection. The single stranded product can bind with probe or molecular beacon with low background signal.
- LATE PCR is used for detection and quantification of low amount DNA or RNA target.
- It is also used in single-cell genetic diagnosis, multiplex assay and viral detection.
- The main limitation is that more optimization is required. Primer concentration and Tm adjustment should be done carefully.
- More cycles may be required to get enough single stranded DNA product.
- Probe design should match with the strand produced in linear phase.
- Sample quality and template integrity strongly affect the result of LATE PCR.
30. Inverse PCR
Inverse PCR (iPCR) is a modified type of PCR in which unknown DNA region present outside a known sequence is amplified. It is used when internal sequence is known but flanking sequence is not known.
The following are the important points of Inverse PCR–
- In Inverse PCR, the DNA sample is first cut by restriction enzyme. The enzyme cuts around the known region and produces DNA fragments containing known sequence with unknown flanking regions.
- The restriction fragments are then ligated under suitable condition. Due to self ligation, the linear DNA fragment becomes circular DNA molecule and the unknown flanking ends come close to each other.
- The primers are designed from the known internal sequence. These primers are oriented outward, so extension moves away from the known region and passes through the ligated junction.
- The circularized DNA acts as template for amplification. Because of circular arrangement, the outward facing primers can amplify the unknown flanking DNA region.
- The PCR product contains known core sequence with adjoining unknown flanking sequence. This product can be sequenced for identification of surrounding DNA region.
- Inverse PCR is used for identification of retroviral or transposon integration site. It is also used for mapping unknown chromosomal DNA present near a known sequence.
- It is useful for cloning of promoter or regulatory region present upstream of a gene. It is also used in genome walking and site directed mutagenesis of circular DNA like plasmid.
- The main limitation is that proper size of restriction fragment is required. Very large fragment may not circularize properly and may amplify poorly.
- Restriction enzyme selection is important in this method. The enzyme should not cut inside the known core region.
- Ligation and template preparation should be done carefully. Poor circularization gives weak or no amplification.
31. TAIL-PCR
TAIL-PCR is a PCR method in which unknown DNA sequence present near a known sequence is amplified. It is used for recovery of flanking genomic DNA.
The following are the important points of TAIL-PCR–
- In TAIL-PCR, nested specific primers are used with arbitrary degenerate (AD) primers. The specific primer binds with known DNA region and AD primer binds at different possible sites in unknown flanking DNA.
- The reaction is done in primary, secondary and tertiary PCR rounds. In these rounds, high stringency and low stringency annealing cycles are used, so that specific primer binding and AD primer binding can occur in controlled way.
- AD primers have lower melting temperature (Tm) than specific primers. So in low stringency cycle they can bind at many genomic sites, but in high stringency cycle specific primers work more strongly.
- In secondary and tertiary rounds, inner nested specific primers are used. These primers bind closer to the known region and increase amplification of correct product containing known region with unknown flanking DNA.
- Non-specific products may be formed in early cycles because AD primers can bind at many sites. But nested rounds increase specificity and suppress many unwanted products.
- TAIL-PCR is used for isolation of flanking region near T-DNA or transposon insertion site. It is also used for recovery of promoter or regulatory region present near a known gene.
- It is useful in genome walking. Unknown genomic DNA adjacent to a known sequence can be obtained without knowing the whole sequence.
- hiTAIL-PCR is an improved form of TAIL-PCR. It uses optimized primer design and thermal cycling, and gives larger flanking amplicons with higher success rate.
- The main limitation is non-specific amplification. This occurs because AD primers bind at many different genomic sites.
- More than one PCR round is required. So contamination risk becomes high.
- The size of unknown flanking region may affect amplification. Very long unknown region may not be amplified properly.
- Proper nested primer design is required. A known DNA sequence must be available before starting TAIL-PCR.
32. Ligation-mediated PCR
Ligation-mediated PCR (LM-PCR) is a technique in which DNA fragment having one known end is amplified by ligating an adapter to the unknown end. Then one primer is used from known region and another primer is used from adapter sequence.
The following are the important points of Ligation-mediated PCR–
- In LM-PCR, the DNA is first digested by restriction enzyme or sheared into fragments. The ends are prepared as blunt end or sticky end and then a known sequence adapter or linker is ligated to the fragment.
- The adapter provides binding site for adapter specific primer. Another primer is designed from the known internal DNA region and both primers are used for amplification of adjacent unknown sequence.
- Nested primers may be used in further rounds of PCR. It increases specificity and reduces background amplification from unwanted fragments.
- LM-PCR has high sensitivity. It can detect low amount DNA fragments when only one end of the sequence is known.
- Linker or adapter should be designed properly. It should ligate efficiently and should not self-ligate in high amount.
- The main limitation is that it has many steps like digestion, ligation and purification. So sample loss and contamination risk may occur.
- Some fragments may not ligate properly or may be too long for amplification. The unknown region recovered by this method is often limited in size.
- LM-PCR is used in genome walking, mapping of transposon or viral integration site, detection of promoter or regulatory region, DNA methylation study and in vivo footprinting.
33. Alu PCR
Alu PCR is a DNA fingerprinting method in which genomic regions between Alu repetitive elements are amplified. It is mainly used for detection of polymorphism and length variation.
The following are the important points of Alu PCR–
- In Alu PCR, primers are designed from Alu sequences. These primers amplify the DNA region present between two Alu repeats and produce many fragments of different sizes.
- The amplified fragments are separated by gel electrophoresis or capillary electrophoresis. The band or peak pattern shows variation between samples.
- Genetic polymorphism is detected because Alu insertion, deletion, mutation near Alu element or difference in length between two Alu repeats changes the size of PCR product.
- Alu PCR is sensitive for detection of human DNA. This is because Alu elements are highly abundant in human genome and more than 1 million copies are present.
- Alu PCR may also be used in quantitative or semi-quantitative method. Alu-based qPCR can detect small amount of human DNA in mixed sample.
- The main limitation is that the fragment pattern may be complex and difficult to interpret.
- Non-specific amplification and background band may occur if primer design and reaction condition are not proper.
- Degraded DNA may give poor amplification, mainly for large inter-Alu fragments.
- Alu PCR is used in DNA fingerprinting, human DNA detection, polymorphism analysis, forensic study and comparison of genetic variation.
34. Repetitive sequence-based PCR
Repetitive Sequence-Based PCR (rep-PCR) is a molecular typing method in which repetitive DNA elements present in genome are amplified. It produces strain or species specific fingerprint pattern.
The following are the important points of rep-PCR–
- In rep-PCR, primers are designed against repetitive DNA elements. These elements include REP (Repetitive Extragenic Palindromic), ERIC (Enterobacterial Repetitive Intergenic Consensus) and BOX sequences.
- These repetitive sequences are scattered in bacterial genome. So primers can bind at many places and amplify the DNA regions present between these repeat elements.
- Many PCR products of different lengths are formed in this reaction. The size of products depend on distance between two repetitive elements present in the genome.
- The amplified products are separated by gel electrophoresis or capillary electrophoresis. The band or peak pattern obtained is called fingerprint pattern.
- The fingerprint pattern of different strains or isolates are compared. Similarity and difference in band pattern helps in discrimination of strains.
- rep-PCR is useful for screening of many isolates. Usually one PCR or few PCR reactions with different primer sets are enough for typing.
- Specificity and reproducibility depends on primer selection, PCR condition, annealing temperature, extension time and quality of DNA.
- rep-PCR can detect variation among closely related strains. But small mutation like SNP may not be detected if repetitive element position is not changed.
- It is less costly than full genome sequencing or PFGE (Pulsed-Field Gel Electrophoresis). It needs simple PCR machine and gel electrophoresis system.
- The main limitation is that band pattern may be complex. Gel quality, band scoring and laboratory condition may affect interpretation.
- Some species may have less repetitive elements. In such case, usefulness of rep-PCR becomes low.
35. Nanoparticle-Assisted PCR (nanoPCR)
Nanoparticle-Assisted PCR (nanoPCR) is a modified PCR method in which nanoparticles are added into the reaction mixture. It is used to improve sensitivity, specificity and speed of amplification.
The following are the important points of nanoPCR–
- In nanoPCR, different nanoparticles may be used. Gold nanoparticles (AuNPs) are commonly used because they have good thermal conductivity and improve heating and cooling during PCR.
- Nanoparticles interact with PCR components like primers, DNA polymerase and single stranded DNA. This interaction helps in specific amplification and reduces non-specific primer binding.
- The concentration, size and type of nanoparticles should be optimized. Metal, oxide or carbon based nanoparticles may be used, but excess amount may inhibit the PCR reaction.
- nanoPCR increases sensitivity of detection. In some viral detection studies, it can detect about 10 to 1000 fold lower copy number than conventional PCR.
- Specificity is also improved when nanoparticle concentration is proper. Non-specific bands and background amplification are reduced.
- nanoPCR may reduce reaction time. Annealing and extension time can be shortened in some optimized protocols.
- It is useful for samples having low template amount, degraded DNA or some inhibitory substances. So it is used in clinical, field and veterinary diagnosis.
- The main limitation is that nanoparticle concentration should be carefully controlled. High concentration or wrong size of nanoparticle may inhibit enzyme activity.
- The effect of nanoparticles on DNA polymerase stability and fidelity should be checked. Otherwise wrong result may occur.
- Reproducibility may vary between laboratories because nanoparticle type, size and reaction condition can change the result.
36. Miniprimer PCR
Miniprimer PCR is a type of PCR in which very short primers are used. These short primers are called miniprimers and are usually about 9-10 nucleotides long.
The following are the important points of Miniprimer PCR–
- In Miniprimer PCR, short primers are used for amplification of more divergent DNA sequences. These primers can bind in regions where normal long primers may not work properly.
- Special engineered DNA polymerase is used in this method. Example is S-Tbr polymerase, which can extend from short miniprimers with proper specificity and yield.
- Annealing temperature is kept lower than normal PCR. This is because short primers have lower melting temperature and need suitable condition for binding with template DNA.
- Miniprimer PCR is used in environmental samples like soil and microbial mats for 16S rRNA gene amplification. It can detect novel and highly divergent microbial sequences which may be missed by standard PCR.
- The specificity and reproducibility depends on primer design, annealing temperature and enzyme choice. Short primers may bind at many non-target sites, so optimization is important.
- The sensitivity is moderate to high. It can detect rare or low abundance divergent sequences, but yield may be lower than standard PCR in some cases.
- The main limitation is non-specific annealing. Since primers are short, they may bind broadly and produce unwanted products.
- It may show amplification bias and poor discrimination between very similar sequences. Some target regions with secondary structure or low complexity may not be suitable.
- Miniprimer PCR is used in microbial diversity study, environmental DNA survey, community profiling, genotyping and fingerprinting of bacteria or plant pathogens.
37. Suicide PCR
Suicide PCR is a PCR method in which primer combination is used only one time in laboratory. It is used mainly for ancient DNA or paleogenetic study to reduce contamination.
The following are the important points of Suicide PCR–
- In Suicide PCR, primers are not used before in that laboratory for any positive control or previous amplification. The target genomic region also should not be previously amplified in that lab.
- This method prevents carry-over contamination from old PCR products. So false positive result from previous amplicons is reduced.
- Suicide PCR is useful when the sample contains very low amount and degraded DNA. It is mainly used in ancient DNA, paleogenetics and ancient pathogen detection.
- Strict laboratory practice is required in this method. Reagents, pipettes, surfaces and work area should be very clean, and the unused primer rule should be followed properly.
- The amplification condition is like normal PCR, but the amplified product is usually confirmed by sequencing. This helps to check whether the product is authentic.
- No positive control is used in Suicide PCR because it may create contaminating amplicon. For this reason, false negative result may increase if the reaction fails.
- The specificity is high when the laboratory is clean and target DNA is really present in the sample.
- The main limitation is that very degraded DNA may not amplify. DNA damage may prevent primer binding and extension.
- It is difficult to prove reaction failure because positive control is avoided. So documentation and strict discipline are important.
- Suicide PCR is used in ancient DNA study, paleogenetics, ancient microbial detection and low copy number target detection.
References
- AAT Bioquest. (n.d.). Standard vs. Touchdown PCR.
- abm Inc. (n.d.). Polymerase Chain Reaction (PCR) – Variations to the System.
- Bento Bio. (n.d.). Touchdown and Stepdown PCR.
- Biocompare. (n.d.). qPCR or dPCR: Choosing the Right Solution for Your Research Needs.
- BioGenex. (n.d.). in-situ PCR.
- Bio-Resource. (n.d.). Allele Specific PCR (AS-PCR) / Amplification Refractory Mutation System (ARMS) – Genotyping PCR [Video]. YouTube.
- BioTnA. (n.d.). In Situ PCR.
- BOC Sciences. (n.d.). Differences Between Various PCR Techniques.
- Chauhan, T. (2019, March 13). What is touchdown (TD)-PCR? Genetic Education.
- Chauhan, T. (2019, March 29). Inverse PCR: Principle, Procedure, Protocol and Applications. Genetic Education.
- Comparable Sensitivity and Specificity of Nested PCR and Single-Stage PCR Using a Thermally Activated DNA Polymerase. (n.d.). ResearchGate.
- Comparison of Conventional, Nested, and Real-Time PCR Assays for Rapid and Accurate Detection of Vibrio vulnificus. (n.d.). PubMed Central (PMC).
- Comparison of the Specificity and Sensitivity of PCR, Nested PCR, and Real-Time PCR for the Diagnosis of Histomoniasis. (n.d.). ResearchGate.
- Eppendorf US. (2018, July 18). In-Situ PCR: PCR Directly Inside a Cell?
- GMP Plastic. (2025, February 27). How do to Colony PCR.
- Gonzalez, S. (2024, February 28). The Different Types of PCR Methods. Pipette.com.
- Inverse Polymerase Chain Reaction (PCR). (n.d.). PubMed.
- Inverse Polymerase Chain Reaction (PCR). (n.d.). ResearchGate.
- Khan Academy. (n.d.). Polymerase chain reaction (PCR) (article).
- Khehra, N., Padda, I. S., & Zubair, M. (2025, July 7). Polymerase Chain Reaction (PCR). In StatPearls. StatPearls Publishing.
- Krishnapur, P. (2023, August 23). Types of PCR – 7 Methods, Principles & Applications. Assay Genie.
- Lowe-Power, T. (2021). Colony PCR | protocols. Lowe-Power Lab.
- LubioScience. (2025, May 22). Bench tip: how to do colony PCR properly.
- McDonald, C., Taylor, D., & Linacre, A. (2024). PCR in Forensic Science: A Critical Review. Genes, 15(4), 438.
- MilliporeSigma. (n.d.). Colony PCR.
- MilliporeSigma. (n.d.). Hot Start PCR.
- MilliporeSigma. (n.d.). Standard PCR Protocol.
- MyBioSource. (n.d.). Types of PCR: Conventional, Real-Time, Reverse Transcriptase, Digital, and Multiplex PCR.
- Patsnap Synapse. (2025, May 7). PCR Machines Explained: Conventional vs. Real-Time vs. Digital PCR.
- Principles of digital PCR and its applications in current obstetrical and gynecological diseases. (n.d.). PubMed Central (PMC).
- QIAGEN. (n.d.). dPCR vs qPCR vs end-point PCR.
- QIAGEN. (n.d.). Types of PCR.
- Rajalakshmi, S. (2017). DIFFERENT TYPES OF PCR TECHNIQUES AND ITS APPLICATIONS. International Journal of Pharmaceutical, Chemical and Biological Sciences, 7(3), 285-292.
- Research Techniques Made Simple: Polymerase Chain Reaction. (n.d.). PubMed Central (PMC).
- Sapkota, A. (2025, September 2). 37 Types of PCR with Definition, Principle, and Uses. Microbe Notes.
- Sensitivity and reproducibility of standardized-competitive RT-PCR for transcript quantification and its comparison with real time RT-PCR. (n.d.). PubMed Central (PMC).
- Shoba. (2023, March 6). Touchdown PCR: Easy Intro Plus 5 Expert Tips. Bitesize Bio.
- Takara Bio. (2023, September 28). What is touchdown PCR (TD-PCR) and when would I need to use it?
- The Architectural Evolution and Methodological Diversification of Polymerase Chain Reaction Technologies: A Comprehensive Analysis of Principles, Technical Variants, and Diagnostic Utility. (n.d.).
- Thermo Fisher Scientific. (n.d.). Real-Time vs Digital vs Traditional PCR.
- Types of PCR. (n.d.). Slideshare.
- Wikipedia contributors. (2023, December 11). Inverse polymerase chain reaction. In Wikipedia, The Free Encyclopedia.
- Wikipedia contributors. (2026, May 4). Polymerase chain reaction. In Wikipedia, The Free Encyclopedia.