Pseudomonas Aeruginosa
- Pseudomonas aeruginosa is a fascinating aerobic microorganism, characterized by its motile, Gram-negative rod-shaped bacteria structure. It can be found in various habitats worldwide, from soil and water to living organisms like plants, animals, and humans.
- With an absolute aerobic metabolism, P. aeruginosa shows a positive oxidase reaction, distinguishing it from other bacteria. This unique species serves as the type species for the entire Pseudomonas genus, exhibiting both general properties common to the genus and some relatively uncommon traits.
- One distinguishing feature of P. aeruginosa is its ability to produce pigments that emit fluorescence under UV radiation, classifying it as one of the fluorescent species within the genus. This characteristic gives rise to its species name, “aeruginosa,” derived from the Latin term for “full of copper rust or verdigris (green).”
- Medically, P. aeruginosa has garnered significant attention due to its widespread distribution and resistance against many antibacterial compounds, making it challenging to treat once it infects the host. It is known to be a leading cause of community-acquired infections, including hospital-acquired pneumonia and urinary tract infections, among others.
- Notably, P. aeruginosa is an opportunistic pathogen, commonly found as a part of the normal flora in living beings. However, it can turn pathogenic and induce infections, especially in individuals with compromised immune systems.
- Despite its notoriety as a pathogen, P. aeruginosa also holds industrial significance. It is cultivated for the production of primary and secondary metabolites, making it valuable in various applications. The bacterium’s production of biosurfactants has proven useful in environmental cleanup, restoration, and remediation.
- The challenges associated with treating P. aeruginosa infections lie in its increasing resistance to multiple antibiotics, rendering some strains almost intractable within the host’s body.
- Overall, Pseudomonas aeruginosa remains an intriguing and complex microorganism, with both medical importance and industrial applications, making it a subject of extensive research and study.
Principle
The MacConkey Agar is a specialized culture medium widely used in microbiology for the isolation, identification, and differentiation of gram-negative bacteria, particularly those belonging to the Enterobacteriaceae family. The principle of MacConkey Agar lies in its combination of selective and differential properties, making it an effective tool in microbiological analyses.
The medium’s composition includes pancreatic digest of gelatin and peptones, providing essential nutrients, vitamins, and nitrogenous factors necessary for the growth of microorganisms. Lactose monohydrate serves as the fermentable source of carbohydrates. Additionally, sodium chloride is present to maintain the osmotic balance of the medium.
The selective action of MacConkey Agar is attributed to the inclusion of crystal violet and bile salts. These components act as inhibitors, targeting most gram-positive bacteria and preventing their growth on the agar.
The differentiation of bacteria on MacConkey Agar is based on their ability to ferment lactose. Lactose fermenting strains produce red or pink colonies on the medium. This color change occurs due to the production of acid from lactose fermentation, leading to a decrease in pH. The pH drop causes the neutral red dye present in the agar to change color, resulting in red or pink colonies.
On the other hand, non-lactose fermenting bacteria, including Salmonella, Proteus, Pseudomonas aeruginosa, and Shigella, are unable to utilize lactose. Instead, they utilize peptone, which results in the production of ammonia and a subsequent rise in pH. Consequently, the colonies of non-lactose fermenting bacteria formed on the medium appear colorless or white, and the surrounding agar remains relatively transparent. Sometimes, these colonies can have a golden to brown appearance with dark centers.
MacConkey Agar can be modified by replacing lactose with other sugars, enabling the differentiation of various gram-negative bacteria based on their sugar fermentation capabilities. For example, when sorbitol is used as a replacement sugar, MacConkey Agar becomes useful in isolating and differentiating enteropathogenic E. coli serotypes, such as E. coli O157:H7. Non-sorbitol fermenting E. coli strains will form distinctive white circular colonies on the agar, aiding in their identification.
In summary, the principle of MacConkey Agar involves the selective inhibition of most gram-positive bacteria, coupled with the differential ability to distinguish between lactose fermenting and non-lactose fermenting gram-negative bacteria based on colony colors and characteristics. This makes it an indispensable tool in clinical and research laboratories for the identification of specific bacterial species and the detection of pathogenic strains.
Requirement
- MacConkey Agar Plates: MacConkey agar is a specialized culture medium used to isolate, identify, and differentiate gram-negative bacteria, particularly those from the Enterobacteriaceae family. The agar plates are essential for the growth of microorganisms and for observing their colony characteristics.
- Sterile Swabs: Sterile swabs are necessary tools for collecting samples from various sources, such as surfaces, wounds, or biological specimens. They provide a contamination-free way to transfer the sample onto the culture medium without introducing unwanted microorganisms.
- Sterile Saline: Sterile saline, a solution of salt and water, is used to moisten and suspend the collected samples on the swabs before inoculating them onto the agar plates. It helps in the even distribution of the sample and aids in the growth of microorganisms.
- Incubator: An incubator is a controlled environment used to maintain specific temperature and humidity conditions suitable for the growth of microorganisms. After inoculating the samples onto the agar plates, they are placed in the incubator to allow bacterial growth over a specific period.
- Ultraviolet Light: Ultraviolet (UV) light is employed to visualize certain characteristics of bacterial colonies on the MacConkey agar plates. Some bacterial species produce pigments or fluoresce under UV light, aiding in the identification and differentiation of specific microorganisms.
MacConkey Agar Plates Composition
| Ingredients | Amount |
| Peptone (Pancreatic digest of gelatin) | 17 gm |
| Proteose peptone (meat and casein) | 3 gm |
| Lactose monohydrate | 10 gm |
| Bile salts | 1.5 gm |
| Sodium chloride | 5 gm |
| Neutral red | 0.03 gm |
| Crystal Violet | 0.001 gm |
| Agar | 13.5 gm |
Procedure
Preparation of MacConkey Agar Plates
The preparation of MacConkey agar plates involves several steps to create a specialized culture medium used for the isolation and differentiation of gram-negative bacteria, particularly those of the Enterobacteriaceae family. Here’s a step-by-step guide on how to prepare MacConkey agar plates:
- Weighing and Measuring: Measure 49.53 grams of dehydrated MacConkey agar medium using a laboratory balance. Transfer the measured amount into a clean and sterile container.
- Dissolving the Medium: Add 1000 ml (1 liter) of distilled water to the container containing the measured MacConkey agar medium. Stir or swirl the mixture to ensure the medium is evenly dispersed in the water.
- Boiling the Medium: Place the container with the medium and water mixture on a heat source and bring it to a gentle boil. Continue heating until the medium is completely dissolved in the water. The boiling helps to ensure that all components of the medium are well-mixed and evenly distributed.
- Autoclaving and Sterilization: After dissolving the medium, the next step is to sterilize the solution. Transfer the dissolved MacConkey agar medium into autoclave-safe containers, leaving enough headspace to avoid spillage during autoclaving.
Autoclave the containers at 15 lbs pressure (121°C) for 15 minutes. Autoclaving is a crucial step as it effectively eliminates any potential contaminants, ensuring a sterile environment for bacterial growth.
- Cooling the Medium: Once the autoclaving process is completed, remove the containers from the autoclave and allow them to cool to approximately 45°C – 50°C. Cooling the medium is essential to prevent heat-related damage to the agar and to make it safe for handling.
- Mixing the Medium: Before pouring the medium into sterile Petri plates, ensure that it is adequately mixed. Gentle swirling or stirring of the medium will help achieve uniform consistency.
- Pouring the Plates: Using aseptic techniques, pour the cooled and mixed MacConkey agar medium into sterile Petri plates. The plates should be placed on a sterile surface, and the medium should be poured carefully to avoid contamination.
- Solidification and Storage: Allow the poured plates to solidify at room temperature or in a controlled environment. Once solidified, the MacConkey agar plates are ready for use in microbiological experiments and bacterial culture studies.
Pseudomonas Aeruginosa Inoculation on MacConkey Agar Plates
Inoculation of Pseudomonas aeruginosa on MacConkey agar plates involves a series of steps to culture and identify this gram-negative bacterium. Pseudomonas aeruginosa is a medically important bacterium known for causing various infections in humans. Here’s a step-by-step guide on how to inoculate Pseudomonas aeruginosa on MacConkey agar plates:
- Labeling the MacConkey Agar Plates: Before starting the inoculation process, ensure that the MacConkey agar plates are properly labeled with relevant information such as the date, sample source, and any other necessary identifiers.
- Swabbing the Specimen: Using a sterile swab, collect a sample of the desired specimen suspected to contain Pseudomonas aeruginosa. This specimen could be obtained from various sources, such as wounds, clinical samples, or environmental samples.
- Inactivating the Swab: After swabbing the specimen, it is important to inactivate the swab to prevent any potential growth of bacteria during the transfer process. To achieve this, roll the swab on a sterile saline pad, ensuring that any viable bacteria on the swab are neutralized.
- Streaking the Swab on MacConkey Agar Plate: Next, streak the inactivated swab across the surface of the MacConkey agar plate. Streaking helps to evenly distribute the bacteria across the agar surface and allows for the isolation of individual colonies. Be careful not to press too hard on the agar surface to avoid damaging the medium.
- Incubating the Plate: Once the swab has been streaked on the MacConkey agar plate, place the plate in an incubator set at 37°C. Incubate the plate for 24-48 hours to allow sufficient time for bacterial growth.
- Examination under Ultraviolet Light: After the incubation period, remove the MacConkey agar plate from the incubator and examine it under ultraviolet (UV) light. Pseudomonas aeruginosa is a fluorescent species, and under UV light, it will produce characteristic pigments that fluoresce, aiding in its identification. This fluorescence will help distinguish Pseudomonas aeruginosa from other bacteria present on the plate.
- Interpretation of Results: Based on the colony characteristics, including color and fluorescence, Pseudomonas aeruginosa can be identified. Lactose-fermenting strains of Pseudomonas aeruginosa will produce pink or red colonies on the MacConkey agar plate, while non-lactose fermenting strains will appear colorless or white.
Interpretation
Interpreting Pseudomonas aeruginosa on MacConkey agar plates involves observing the colony characteristics and differentiating them from other gram-negative bacteria, particularly Escherichia coli. MacConkey agar is a selective and differential medium used for the isolation and identification of gram-negative bacteria based on their ability to ferment lactose. Here’s how to interpret Pseudomonas aeruginosa on MacConkey agar plates:
- Colony Color: Pseudomonas aeruginosa colonies on MacConkey agar will typically appear colorless or pale yellow. This is because Pseudomonas aeruginosa is a non-lactose fermenting bacterium, and as a result, it does not produce acid from lactose fermentation. As a non-lactose fermenter, it does not cause the pH of the agar to drop, and therefore, it does not produce the characteristic pink or red color associated with lactose-fermenting bacteria.
- Fluorescence under Ultraviolet Light: One distinctive feature of Pseudomonas aeruginosa is its ability to produce fluorescent pigments. When examined under ultraviolet (UV) light, Pseudomonas aeruginosa colonies may exhibit fluorescence, which sets them apart from many other bacteria on MacConkey agar plates.
- Differentiation from Escherichia coli: Another common gram-negative bacterium found on MacConkey agar is Escherichia coli (E. coli). E. coli is a lactose-fermenting bacterium, and its colonies will appear pink or red due to the production of acid from lactose fermentation. This is a key characteristic used to differentiate E. coli from Pseudomonas aeruginosa, which does not ferment lactose and, therefore, lacks the red or pink coloration.
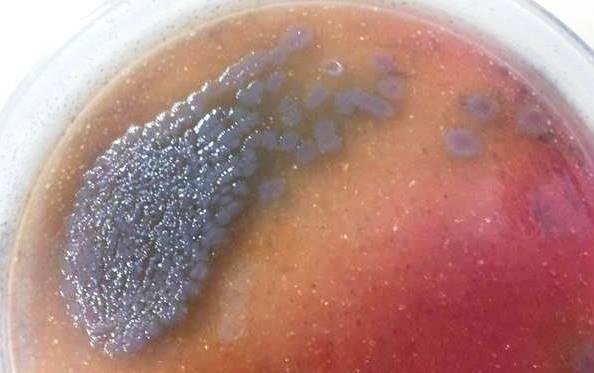
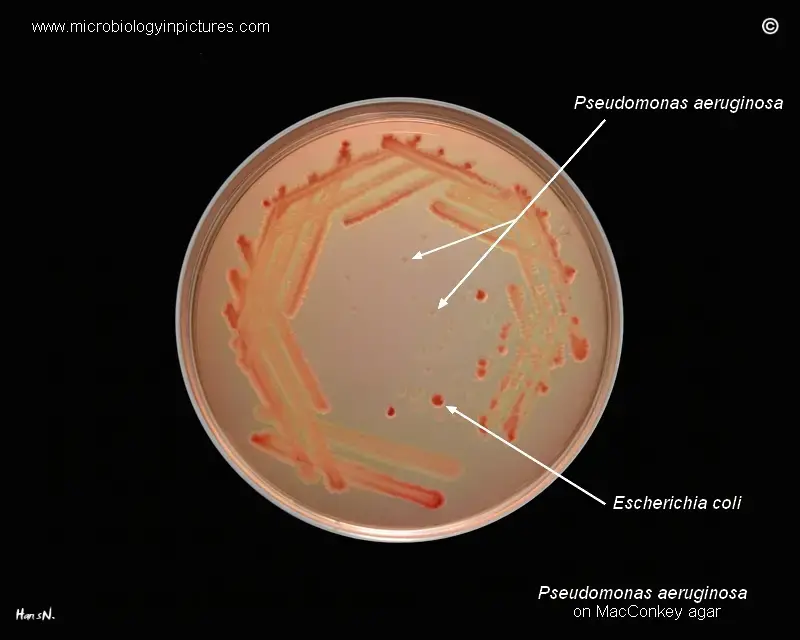
Cultural Characteristics of Pseudomonas Aeruginosa
| Cultural Characteristics | Nutrient Agar Medium (NAM) | Cetrimide Agar medium | MacConkey Agar medium | Blood Agar Medium |
|---|---|---|---|---|
| Shape | Irregular | Circular | Circular | Irregularly edged circular |
| Size | 2-4 mm | 1-3 mm | 2-3 mm | 2-4 mm |
| Elevation | Low Convex | Low Convex | Low Convex | Flat |
| Surface | Smooth (fresh isolation) ; Mucoid (When slime layer is formed) | Smooth (fresh isolation) ; Mucoid (When slime layer is formed) | Smooth (fresh isolation) ; Mucoid (When slime layer is formed) | Smooth (fresh isolation) ; Mucoid (When slime layer is formed) |
| Color | Greenish blue (due to pigment production) | Greenish blue (due to pigment production) | Colorless | Greyish white |
| Structure | Translucent –Opaque | Opaque | Transparent | Translucent –Opaque |
| Hemolysis | —– | —– | —– | β-Hemolysis (in some strains) |
Notes
When working with Pseudomonas aeruginosa on MacConkey agar plates, several important notes should be taken into consideration to ensure accurate results and prevent potential issues:
- Use of Sterile Swabs and Saline: To avoid introducing contaminants onto the agar plates, always use sterile swabs and sterile saline when collecting and transferring specimens. Contamination could lead to false results and hinder the accurate identification of Pseudomonas aeruginosa colonies.
- Incubation Temperature and Time: It is crucial to incubate the MacConkey agar plates at the appropriate temperature of 37°C. This temperature mimics the human body’s conditions, which are favorable for the growth of Pseudomonas aeruginosa. The plates should be incubated for a period of 24-48 hours to allow sufficient time for optimal bacterial growth.
- Observation of Colony Size: Pseudomonas aeruginosa colonies on MacConkey agar plates may appear small and inconspicuous. To ensure a thorough examination, plates should be carefully observed under suitable lighting conditions to detect any potential colonies. It is essential not to overlook small colonies, as they might be indicative of the presence of Pseudomonas aeruginosa.
- Additional Tests for Confirmation: If there is uncertainty regarding the identity of a colony on the MacConkey agar plate, further tests should be performed to confirm if it is indeed Pseudomonas aeruginosa. Biochemical testing, such as the oxidase test, can be carried out to assess specific enzymatic reactions characteristic of Pseudomonas aeruginosa. Microscopy can also provide valuable information about the colony’s morphology and arrangement, assisting in the identification process.
- Proper Aseptic Technique: Maintaining strict aseptic technique throughout the process is crucial to prevent cross-contamination and ensure accurate results. Proper handwashing, using sterile equipment, and working in a clean environment are essential practices when handling specimens and agar plates.
- Record Keeping: Always keep accurate and detailed records of the specimen source, sample collection, incubation time, and any additional tests performed. Proper record keeping is essential for traceability, quality control, and reference purposes.
FAQ
What is Pseudomonas aeruginosa, and why is it important to study it on MacConkey agar?
Pseudomonas aeruginosa is a gram-negative bacterium known for causing various infections in humans. MacConkey agar is used to isolate and differentiate gram-negative bacteria based on their lactose fermentation abilities, making it a valuable medium for studying Pseudomonas aeruginosa and identifying its unique characteristics.
How do Pseudomonas aeruginosa colonies appear on MacConkey agar?
Pseudomonas aeruginosa colonies on MacConkey agar will be colorless or pale yellow since it is a non-lactose fermenter. Additionally, these colonies may exhibit fluorescence under ultraviolet light due to the production of fluorescent pigments.
What temperature and duration are suitable for incubating MacConkey agar plates to detect Pseudomonas aeruginosa?
MacConkey agar plates should be incubated at 37°C for 24-48 hours to allow optimal growth of Pseudomonas aeruginosa.
Are Pseudomonas aeruginosa colonies easy to spot on MacConkey agar plates?
Pseudomonas aeruginosa colonies can be small and inconspicuous, making them challenging to detect. Close and careful examination of the plates is necessary to identify these colonies accurately.
Can other gram-negative bacteria grow on MacConkey agar?
Yes, MacConkey agar supports the growth of various gram-negative bacteria. Notably, Escherichia coli will produce pink or red colonies due to lactose fermentation, allowing differentiation from Pseudomonas aeruginosa.
What happens if I’m unsure whether a colony is Pseudomonas aeruginosa on MacConkey agar?
If there is uncertainty regarding the identity of a colony, additional tests, such as biochemical testing or microscopy, can be performed to confirm whether it is Pseudomonas aeruginosa.
How should I handle the swabs and saline while working with Pseudomonas aeruginosa on MacConkey agar?
Always use sterile swabs and sterile saline to prevent contamination of the plates. Proper aseptic technique is essential to obtain accurate results.
Can Pseudomonas aeruginosa cause infections in humans?
Yes, Pseudomonas aeruginosa is an opportunistic pathogen known to cause a wide range of infections in humans, particularly in immunocompromised individuals or those with underlying health conditions.
What are some of the diseases caused by Pseudomonas aeruginosa?
Pseudomonas aeruginosa can cause various infections, including hospital-acquired pneumonia, urinary tract infections, skin and soft tissue infections, bloodstream infections, and infections in burn patients.
How can understanding Pseudomonas aeruginosa on MacConkey agar help in healthcare settings?
Studying Pseudomonas aeruginosa on MacConkey agar aids in the identification of this bacterium and its antimicrobial susceptibility, allowing healthcare providers to implement appropriate treatment strategies and infection control measures.