- The Gram-positive cell wall is a crucial feature of Gram-positive bacteria, contributing to their structural integrity and interaction with the environment. It is a defining characteristic used to distinguish these bacteria from Gram-negative ones.
- Gram-positive bacteria, such as Staphylococcus aureus, Streptococcus pneumoniae, and Enterococcus faecalis, possess a distinctive cell wall structure that plays a central role in their physiology. The cell wall of Gram-positive bacteria is notably thick, primarily composed of peptidoglycan (PG), which provides rigidity and strength. This peptidoglycan layer is significantly thicker compared to that found in Gram-negative bacteria, forming a robust framework that maintains cell shape and protects against osmotic pressure.
- Embedded within the peptidoglycan layer are various components that contribute to the cell wall’s function and versatility. One of the major components is teichoic acids (TAs). These long, anionic polymers are integrated into the peptidoglycan and the cytoplasmic membrane, playing several roles in bacterial physiology. Teichoic acids are involved in cell wall biosynthesis, regulation of ion transport, and adherence to host tissues. They also influence the bacterium’s interaction with the immune system and its ability to cause disease.
- The Gram-positive cell envelope also includes capsular polysaccharides (CPS), which are covalently attached to the peptidoglycan layer. These polysaccharides contribute to the formation of a protective capsule around the bacterium, enhancing its ability to evade immune detection and contribute to virulence.
- Additionally, the cell wall may contain surface proteins and extracellular polysaccharides that are involved in various functions. Surface proteins often serve as receptors or adhesion factors, facilitating interactions with host cells and contributing to pathogenesis. These proteins can also modulate the passage of nutrients and ions across the membrane, and participate in the enzymatic processes that maintain and remodel the cell wall.
- The complex structure of the Gram-positive cell wall makes it a target for antibiotics. Several antibiotic classes specifically inhibit peptidoglycan synthesis, leading to cell lysis and death. However, the rise of antibiotic resistance has prompted increased research into the cell envelope’s biogenesis and regulation. Understanding these processes is crucial for developing new therapeutic strategies.
Bacterial Cell Wall
- Prokaryotic cells are typically restricted by a rather robust and chemically complex structure found in between cell membranes and the the capsule/slime layer, also known as the cell wall.
- Peptidoglycan is the primary part of the cell’s wall. It is responsible for shaping and strength of the cell.
- Peptidoglycan is a disaccharide and contains two sugar derivatives–N-acetylglucosamine and N-acetylmuramic acid–joined together by short peptide chains.
- N-acetylmuramic acid is the tetrapeptide side chain composed of Damino acids L and – (D-glutamic acid as well as L-alanine) together with mesodiaminopimelic acid (Gram-negative bacteria) or L-lysine (Gram-positive bacteria).
- Side chains of Tetrapeptide are linked via pentaglycine bridges.
- Most Gram-negative cell walls lack interpeptide bridge.
- Cell wall gives shape to cells and guards the bacteria from changes of inosmotic pressure. The pressure inside the cell of bacteria is 5-20 atmospheres.
- Bacterial cells are classified as Gram-positive and Gram-negative, based on the structural differences that exist between Gram-positive and Gram-negative cell walls.
- Cell walls of Gram-positive bacteria are more chemically simple structures than Gram-negative bacteria.
- The cells’ walls Bacillus subtilis, as well as a variety of other Gram-positive bacteria comprise of a single thick 20 to 80-nm uniform layer composed of peptidoglycan (murein) which is located just outside the plasma membrane.
Characteristics of Peptidoglycan
Peptidoglycan is a fundamental component of bacterial cell walls, forming a complex, mesh-like structure known as the sacculus. This structure is crucial for maintaining cell shape and integrity. Here are the key characteristics of peptidoglycan:
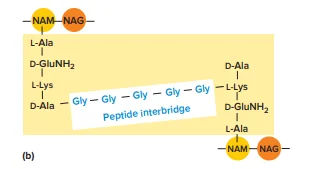
- Composition and Structure: Peptidoglycan is composed of a repeating disaccharide unit and a peptide chain. The disaccharide backbone consists of two sugar derivatives: N-acetylglucosamine (NAG) and N-acetylmuramic acid (NAM). These sugars alternate in the backbone of the peptidoglycan strand.
- Peptide Chains: Attached to the NAM residues are short peptide chains, also known as stem peptides. These peptides typically contain alternating D- and L-amino acids, including D-alanine, D-glutamic acid, and mesodiaminopimelic acid. The peptide chains are linked to the carboxyl group of NAM, forming a robust network.
- Amino Acids: The stem peptides include amino acids not found in proteins, such as D-glutamic acid and mesodiaminopimelic acid. The presence of these unusual amino acids contributes to the structural rigidity and stability of the peptidoglycan layer.
- Cross-Linking: Peptidoglycan strands are interconnected through cross-linking of the peptide chains. There are two types of cross-links:
- Direct Cross-Links: Involves a direct bond between amino acids in adjacent peptide chains. For example, in some bacteria, the carboxyl group of D-alanine at the fourth position in one peptide chain links to diaminopimelic acid at the third position in another chain.
- Indirect Cross-Links: Utilize an interpeptide bridge, a short chain of amino acids connecting peptide chains from different peptidoglycan strands. This type of linkage is common in Gram-positive bacteria.
- Mechanical Properties: The peptidoglycan sacculus is both strong and elastic. It can expand and contract in response to osmotic pressure due to the stiffness of the backbone and the flexible nature of the cross-links. This elasticity is crucial for maintaining cell integrity under varying environmental conditions.
- Permeability: The peptidoglycan layer is porous, allowing proteins with molecular weights up to approximately 50,000 Daltons to pass through. The degree of permeability depends on the sacculus’s state—whether it is stretched or relaxed.
- Variability: There is significant variability in peptidoglycan structures among different bacterial species. For instance, Gram-positive bacteria often have more extensive cross-linking and may substitute certain amino acids, such as using lysine instead of mesodiaminopimelic acid. This variability affects the cell wall’s rigidity and its interaction with antibiotics.
- Role in Antibiotic Targeting: Peptidoglycan is a primary target for antibiotics. Drugs like penicillin inhibit peptidoglycan synthesis, leading to cell lysis. Variations in peptidoglycan structure among bacteria influence their susceptibility to different antibiotics.
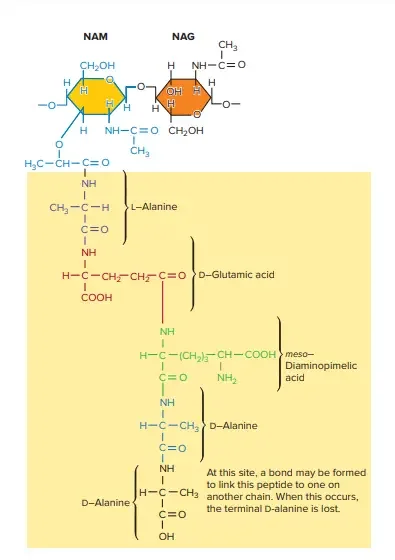
Cell membrane of Gram-Positive Cell Wall
The cell membrane of Gram-positive bacteria is a critical component of the cell envelope, playing essential roles in maintaining cell integrity and mediating interactions with the environment. Here is a detailed overview of its characteristics:
- Lipid Composition: The Gram-positive cell membrane consists of a bilayer membrane that contains a variety of lipids. Major lipids include phosphatidylglycerol and cardiolipin. In certain species, such as those in the genus Bacillus, phosphatidylethanolamine is also prominent. These lipids contribute to membrane stability and function.
- Aminoacylated Phosphatidylglycerol: Some Gram-positive bacteria, including Staphylococcus aureus, produce aminoacylated phosphatidylglycerol, such as lysyl-phosphatidylglycerol. This lipid is synthesized by the enzyme MprF, which transfers lysine from lysyl-tRNA to phosphatidylglycerol. Lysyl-phosphatidylglycerol is crucial for reducing susceptibility to antimicrobial peptides and protecting against various antibiotics, including aminoglycosides and β-lactams.
- Membrane Lipid Dynamics: The composition of membrane lipids, including head groups and fatty acyl chains, can rapidly change in response to environmental conditions like pH, osmotic stress, and temperature. For instance, the presence of branched-chain fatty acids can vary based on growth conditions. These variations impact membrane viscosity, permeability, and protein interactions, thus influencing overall cell function and viability.
- Lipoteichoic Acid (LTA): The cell membrane also contains lipoteichoic acid, which is anchored to the lipid bilayer. LTA plays a role in cell wall synthesis, and its modifications, such as d-alanylation, affect the charge density of the cell envelope. This modification helps in resisting antimicrobial peptides and cationic antibiotics.
- Transmembrane and Lipoproteins: The membrane hosts numerous transmembrane proteins and lipoproteins. These proteins are involved in various functions, including:
- Cell Envelope Synthesis: Assisting in the construction and maintenance of the cell envelope.
- Transport: Facilitating the movement of cell envelope precursors and nutrients.
- Export: Removing toxic compounds from the cell.
- Sensory Functions: Many proteins serve as sensory components in two-component sensing systems, which regulate responses to external stimuli such as cell density and toxic substances.
- Regulation and Resistance: The regulation of lysyl-phosphatidylglycerol and other membrane components is controlled by complex protein systems. For example, the GraRS two-component system and the VraFG ABC-transporter-like system in S. aureus modulate the response to antimicrobial peptides and regulate d-alanylation of teichoic acids. These regulatory mechanisms contribute to the bacterium’s resistance to various antibiotics.
- Surface Protein Display: The membrane supports systems for displaying surface proteins, which are crucial for adhesion and interactions with the environment. These proteins help bacteria adhere to surfaces and engage with host tissues, contributing to pathogenicity.
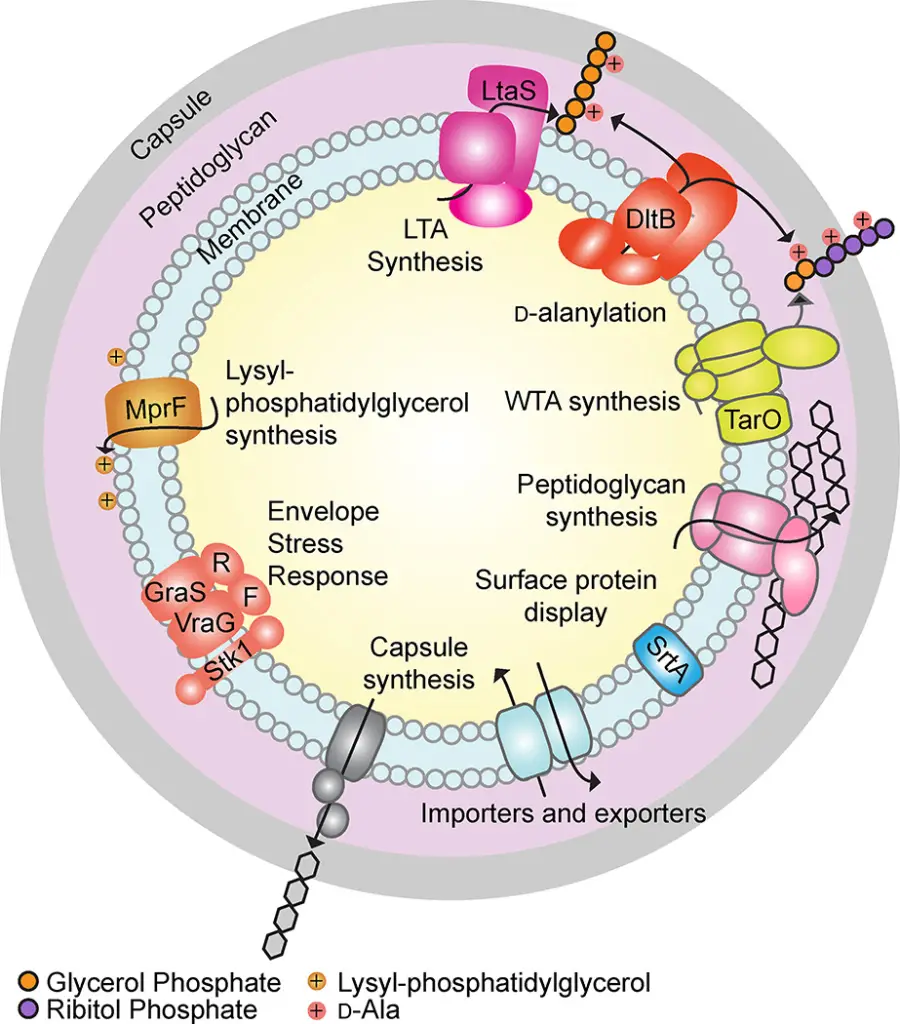
Peptidoglycan
Peptidoglycan (PG) is a crucial component of bacterial cell walls, particularly in Gram-positive bacteria. It provides structural integrity and protects against environmental stress. This macromolecule forms a robust framework around the cell membrane, enabling bacteria to maintain their shape and resist osmotic pressure.
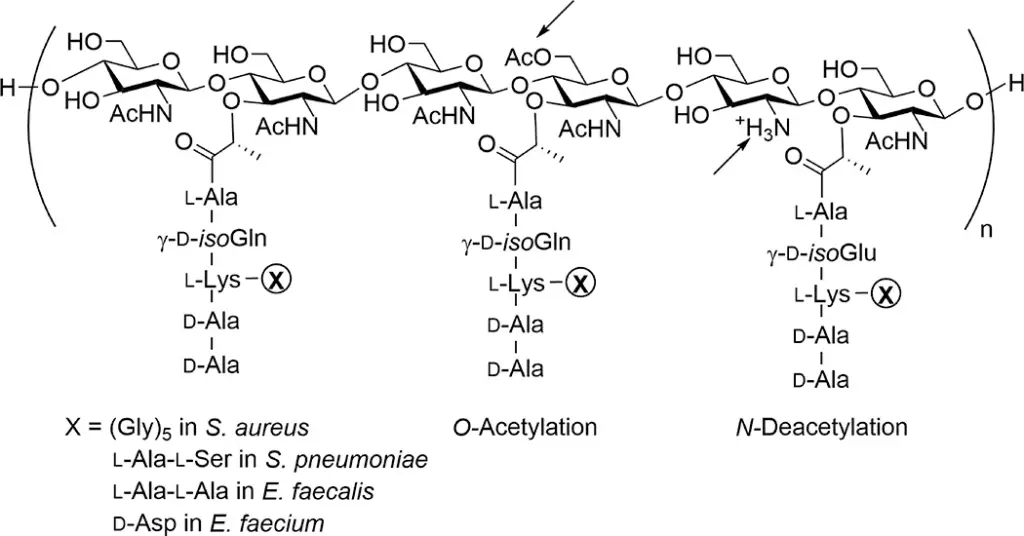
Peptidoglycan Structure
Peptidoglycan consists of repeating disaccharide units, which include N-acetylglucosamine (GlcNAc) and N-acetylmuramic acid (MurNAc). These units are linked through β-1,4-glycosidic bonds, forming linear chains. The disaccharide chains are cross-linked by peptide side chains, contributing to the overall rigidity of the cell wall.
- Disaccharide Units:
- N-acetylglucosamine (GlcNAc) and N-acetylmuramic acid (MurNAc) form the basic repeating unit.
- The average length of these glycan chains can vary significantly across different bacterial species. For instance, in Staphylococcus aureus, glycan strands average between 6 and 18 disaccharide units, while in Bacillus subtilis, they can extend up to 5000 disaccharide units.
- Peptide Cross-Linking:
- The peptide chains attached to MurNAc residues are typically composed of five amino acids. Common configurations include l-alanine, d-isoglutamine, and a terminal d-Ala-d-Ala.
- Cross-linking occurs between the peptides of adjacent glycan chains, often involving the ε-amino group of lysine or meso-diaminopimelic acid. This cross-linking is essential for the structural integrity of the peptidoglycan layer.
Peptidoglycan Biosynthesis
PG biosynthesis is a multi-step process that involves several key stages:
- Cytoplasmic Phase:
- Assembly of UDP-MurNAc Pentapeptide: Involves the formation of UDP-MurNAc-pentapeptide from UDP-GlcNAc.
- Formation of Lipid I and II: The pentapeptide is transferred to a lipid carrier, forming Lipid I, which is then glycosylated to Lipid II.
- Membrane Translocation:
- Flipping of Lipid II: Lipid II is translocated across the bacterial membrane by a flippase enzyme, MurJ.
- Glycan Chain Polymerization: On the outer side of the membrane, Lipid II is polymerized into long glycan chains.
- Cross-Linking:
- Peptidoglycan Glycosyltransferases (PGTs): These enzymes catalyze the polymerization of glycan strands.
- Transpeptidases: These enzymes cross-link the glycan strands, reinforcing the cell wall structure.
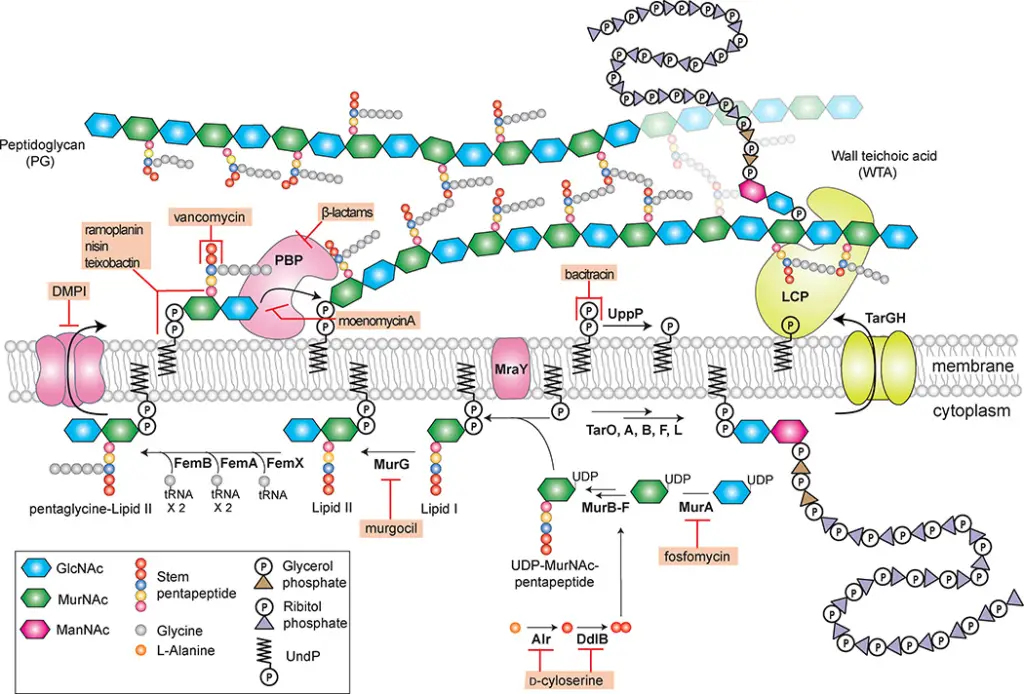
Functions of Peptidoglycan
- Structural Support:
- Peptidoglycan provides structural rigidity, crucial for maintaining cell shape and resisting internal osmotic pressure. In Gram-positive bacteria, the peptidoglycan layer can withstand pressures up to 20 atmospheres.
- Protection:
- The PG layer protects the cell from environmental threats, including antimicrobial agents. Modifications such as N-deacetylation and O-acetylation can enhance resistance to lysozyme and other antimicrobial peptides.
Tailoring Modifications
- N-deacetylation:
- The removal of acetyl groups from GlcNAc and MurNAc residues can protect bacteria from lysozyme, an enzyme that cleaves glycosidic bonds in peptidoglycan.
- O-acetylation:
- The addition of acetyl groups to MurNAc residues contributes to resistance against lysozyme and may affect the overall properties of the cell wall.
Antibiotic Targeting
Given the essential role of peptidoglycan in bacterial cell wall integrity, its biosynthetic pathway is a target for various antibiotics:
- β-Lactams:
- These antibiotics, including penicillin, inhibit peptidoglycan cross-linking by binding to penicillin-binding proteins (PBPs). This prevents proper cell wall synthesis and leads to bacterial cell lysis.
- Vancomycin:
- Vancomycin interferes with the synthesis of peptidoglycan by binding to the d-Ala-d-Ala termini of the peptide side chains, inhibiting cross-linking.
Teichoic Acids
Teichoic acids (TAs) are key components of the cell envelopes of Gram-positive bacteria. They are classified into two main types: wall teichoic acids (WTAs) and lipoteichoic acids (LTAs). Each type plays distinct roles in bacterial physiology and structure.
1. Wall Teichoic Acids (WTAs)
Structure and Composition
Wall teichoic acids are anchored to the peptidoglycan (PG) layer of the bacterial cell wall. They consist of a disaccharide linkage unit connected at one end to PG through a phosphodiester bond and at the other end to a main chain polymer. The main chain of WTAs varies among species but universally includes phosphodiester linkages, which contribute anionic charges to the cell wall.
- In Staphylococcus aureus and Bacillus subtilis, the main chain of WTAs is composed of glycerol-phosphate or ribitol-phosphate repeats.
- In Streptococcus pneumoniae, the main chain includes 2-acetamido-4-amino-2,4,6-trideoxygalactose, glucose, ribitol-phosphate, and two GalNAc moieties, decorated with phosphorylcholine, a rare modification.
- In Enterococcus faecalis, the repeating unit contains d-glucose, d-galactose, 2-acetamido-2-deoxy-d-galactose, 2-acetamido-2-deoxy-d-glucose, and ribitol-phosphate.
- In Enterococcus faecium, the WTA polymer is simpler, with repeating units of 2-acetamido-2-deoxy-D-galactose and glycerol-phosphate.
Biosynthesis
The biosynthesis of WTAs involves several steps:
- Precursor Formation: PhosphoGlcNAC is transferred to an undecaprenyl phosphate (Und-P) lipid carrier, forming a polymeric precursor.
- Polymerization: The precursor is extended by various intracellular enzymes.
- Transport and Ligation: The polymer is transported to the cell surface and attached to PG via a two-component ABC transporter. The exact enzymatic functions of some steps remain under investigation.
In S. aureus, WTAs are not essential for survival in vitro if specific early genes in the pathway are deleted. However, later genes are conditionally essential due to their role in preventing the accumulation of toxic intermediates that inhibit PG synthesis.
Modifications
WTAs and LTAs can be modified with d-alanine esters, which neutralize negative charges and affect cell envelope properties. The d-alanylation involves the dlt operon proteins, which add d-alanine residues to ribitol or glycerol groups. These modifications are crucial for bacterial resistance to antimicrobial peptides and other functions.
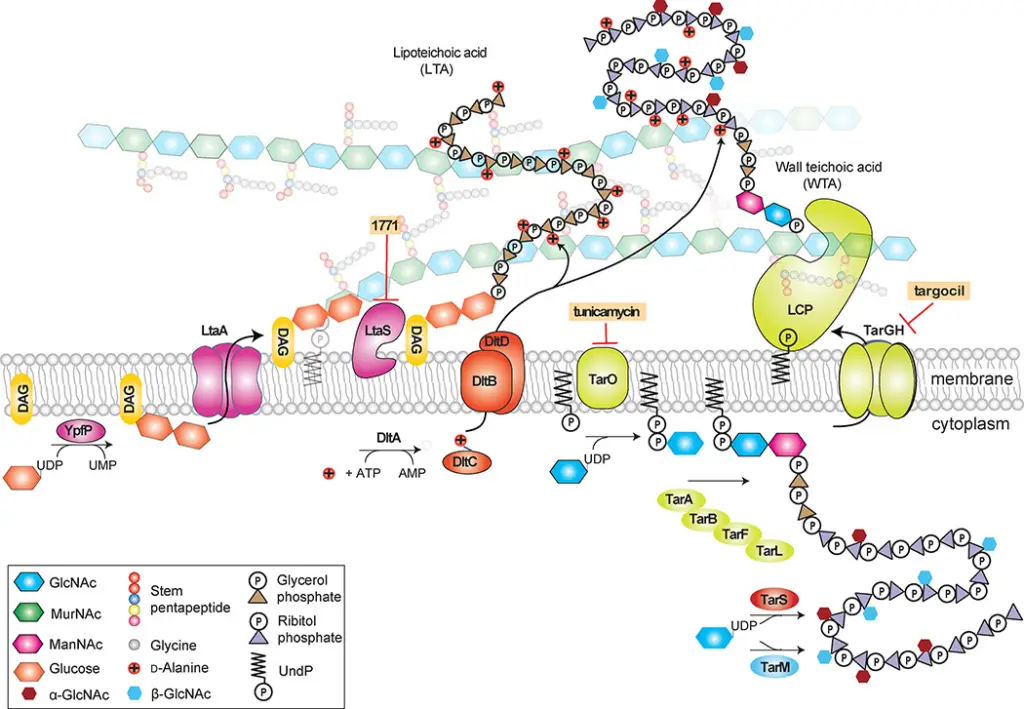
2. Lipoteichoic Acids (LTAs)
Structure and Composition
Lipoteichoic acids are anchored to the cytoplasmic membrane via a glycolipid. The most common structure includes a polyglycerol-phosphate chain attached to a glycolipid anchor. This glycolipid anchor is often diacylglycerol with glucose moieties, although variations exist:
- In S. aureus and B. subtilis, the anchor typically contains two glucose residues.
- In Clostridium difficile, the glycolipid anchor may include more glucose residues.
- In L. monocytogenes, additional sugar moieties such as galactose are present.
Biosynthesis
The synthesis of LTAs begins in the cytoplasm:
- Glycolipid Anchor Formation: The glycolipid anchor, such as Glc2DAG, is synthesized and flipped across the membrane by LtaA.
- Polymerization: Glycerol-phosphate units are transferred to the glycolipid anchor by LtaS. In some organisms, a primase enzyme (e.g., LtaP) adds initial units to the glycolipid anchor.
Differences in LTA synthesis exist among species. For instance, S. aureus and L. monocytogenes employ different sets of enzymes for polymer chain formation.
Functional Roles and Modifications
Teichoic acids are integral to bacterial cell function and structure:
- Cell Morphology: WTAs help maintain rod-shaped morphology in B. subtilis. Disruption in their synthesis leads to abnormal cell shapes and growth defects.
- Cell Division: Both LTAs and WTAs influence cell division and shape. For example, S. aureus mutants with disrupted LTA synthesis show severe cell division defects.
- Immune Evasion: Modifications such as d-alanylation help bacteria evade host immune responses. These modifications are essential for resistance to antimicrobial peptides and other stresses.
What is Capsular Polysaccharides?
Capsular polysaccharides (CPS) are complex glycopolymers found on the surface of many bacterial cells. These structures are anchored to the peptidoglycan layer of the bacterial cell wall, extending outward. CPS play a critical role in the interaction between bacteria and their hosts, influencing both pathogenicity and immune system evasion.
1. Structural Diversity of Capsular Polysaccharides
Capsular polysaccharides exhibit significant structural variability among different bacterial strains. This variability is primarily due to differences in the repeating oligosaccharide units that make up the CPS chains. For instance, in Streptococcus pneumoniae, a large number of serotypes have been identified—93 to be precise. Each serotype has a distinct CPS composition. The differences arise from variations in the gene sequences responsible for CPS biosynthesis.
The CPS of S. pneumoniae can change through natural recombinational events at the biosynthetic locus. This allows the bacteria to switch serotypes, potentially enhancing their ability to evade the host immune system. This switching is linked to variations in the genes responsible for adding or modifying sugar moieties in the CPS structure.
For example, the CPS of S. pneumoniae serotype 2 consists of a backbone made up of glucose-rhamnose units, with side chains of glucose and glucuronic acid. Recent studies have revealed that some serotypes contain two different repeating units within their CPS.
Similarly, Staphylococcus aureus has multiple CPS serotypes, with serotypes 5 and 8 being particularly prevalent in human infections.
2. Role of CPS in Host Immunity
Capsular polysaccharides are crucial for bacterial virulence. They contribute to the pathogen’s ability to avoid detection and destruction by the host’s immune system. CPS can mask the bacterial surface, reducing the effectiveness of phage infection and phagocytosis by immune cells.
For instance, CPS can obstruct the binding of opsonic C3 fragments to complement receptors on immune cells, thus diminishing opsonization and subsequent phagocytosis. This masking effect has been observed in various bacteria, including Enterococcus faecalis. Additionally, in Group B Streptococcus, CPS terminal sialic acid groups interact with Siglecs on human leukocytes, mimicking human cell surface glycans and reducing the innate immune response.
3. CPS and Vaccine Development
The immunomodulatory properties of CPS have made them a target for vaccine development. Historically, vaccines using polyvalent pneumococcal polysaccharides have proven effective. More recent advances involve conjugating CPS to carrier proteins, which enhances vaccine efficacy.
Pneumococcal vaccines today include variations such as PPSV23, which covers 23 serotypes, and conjugate vaccines like PCV7 and PCV13, which cover 7 and 13 serotypes, respectively. PCV13 is recommended for infants and adults over 50 due to its T-cell dependent immune response, which is more suitable for younger and older populations.
However, the use of these vaccines raises concerns about serotype replacement. Studies have shown that while vaccines reduce the incidence of vaccine-covered serotypes, they may lead to an increase in non-vaccine serotypes. This issue highlights the need for continuous monitoring and development of vaccines to address evolving bacterial populations.
4. Challenges and Future Directions
Capsular conjugate vaccines for S. aureus serotypes 5 and 8 have been explored but have not yet proven successful in clinical trials. One potential reason for reduced vaccine efficacy is the presence of natural non-opsonic antibodies to PNAG, another polysaccharide in S. aureus.
Gram-Positive Cell Wall Structure – Overview
The cell’s Gram-positive wall can be described as thick (15-80 nm) and more homogeneous than the thin (2 millimeters) Gram-negative wall. The Gram-positive cell wall has a an extensive amount of peptidoglycan in multiple layers, which makes up around 40-80% of the its dry mass. The cell wall of the Gram-positive cells is composed mostly of teichoic and Teichuronic acids. These two substances make up as much as 50 percent from the total dry mass of the wall, and 10 percent in the weight dry of the entire cell.
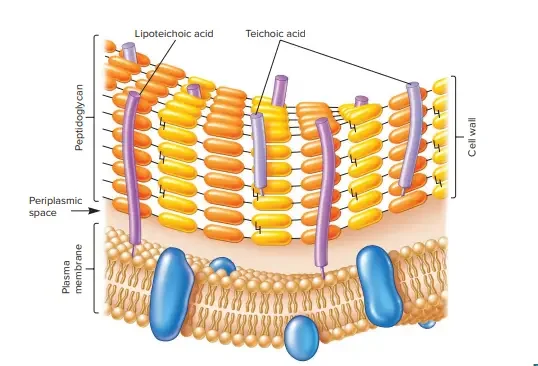
The periplasmic area is located between the plasma membrane as well as the cell wall . It is so small that it is rarely accessible to electron microscopy. The periplasm is home to a small amount of proteins. This is likely because the peptidoglycan sacculus porous, which means that a lot of proteins that move over the plasma membrane travel across the sacculus.
The secreted proteins of some are known as exoenzymes. Exoenzymes are often used to break down polymers, such as polysaccharides and proteins that are otherwise too big for transportation over the plasma membrane. the products of degradation, which are monomer-based building blocks, are then utilized by cells. The proteins that remain in the periplasmic area are typically linked with the plasma membrane. Some proteins that are secreted do not traverse the peptidoglycan sacculus but some of them are attached by it. These proteins play a role in cell interactions with the environment.
A. Teichoic acids
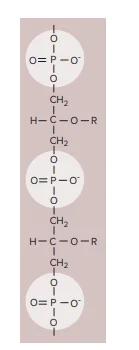
Teichoic acids are important structural components found predominantly in the cell walls of Gram-positive bacteria. These polymers are either polyribitol phosphate or polyglycerol phosphate. They consist of repeating units of ribitol or glycerol linked by phosphate groups. Teichoic acids may also feature sugar or amino acid substitutions, either as side chains or within the polymer chain itself.
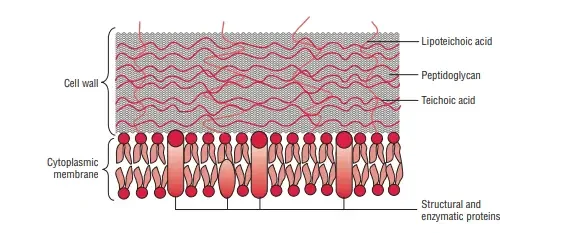
There are two primary types of teichoic acids: wall teichoic acids (WTA) and lipoteichoic acids (LTA). Each plays a distinct role in bacterial cell wall structure and function.
- Wall Teichoic Acids (WTA):
- Structure and Bonding: WTAs are covalently linked to the peptidoglycan layer of the bacterial cell wall. They are attached through the hydroxyl group of N-acetylmuramic acid residues in the peptidoglycan.
- Function: WTAs extend across the surface of the peptidoglycan. They are positively charged, contributing to the overall negative charge of the bacterial cell wall. This negative charge is critical for interactions with the environment and for maintaining cell wall integrity.
- Lipoteichoic Acids (LTA):
- Structure and Bonding: LTAs are covalently attached to the plasma membrane lipids. This linkage involves the glycolipid components of the bacterial membrane.
- Function: LTAs extend from the membrane into the peptidoglycan layer, influencing cell wall stability and function. They also play a role in bacterial adhesion and in modulating the immune response.
Teichoic acids are not present in all bacteria. Their presence is primarily a feature of Gram-positive bacteria, which belong to the phyla Firmicutes and Actinobacteria. These bacteria typically have thick cell walls rich in peptidoglycan, and teichoic acids contribute to the structural complexity of these walls.
Functions of Teichoic acids
Teichoic acids play a variety of crucial roles in the physiology and interactions of Gram-positive bacteria. Their functions can be broadly categorized into structural, protective, and antigenic roles. Here’s a detailed explanation of their key functions:
- Structural Functions:
- Cell Wall Stability: Teichoic acids contribute to the structural integrity of the bacterial cell wall. Wall teichoic acids (WTA) are integrated into the peptidoglycan layer, extending across its surface. This integration helps maintain cell wall rigidity and supports cell wall stability.
- Membrane Anchoring: Lipoteichoic acids (LTA) are covalently linked to the plasma membrane. They anchor the cell wall to the underlying membrane, stabilizing the bacterial envelope.
- Protective Functions:
- External Permeability Barrier: Teichoic acids form an external barrier for Gram-positive bacteria. This barrier protects the bacterial cell from harmful substances such as antibiotics, toxins, and host defense molecules.
- Magnesium Ion Binding: Teichoic acids can bind magnesium ions. This binding may play a role in supplying essential magnesium ions to the cell, which is critical for various cellular processes.
- Antigenic Functions:
- Surface Antigens: Teichoic acids serve as major surface antigens in Gram-positive bacteria. For instance, in Streptococcus pneumoniae, they carry Forssman antigens, which are used in serological identification.
- Serological Classification: They are utilized in the serological classification of bacteria, aiding in the identification and differentiation of bacterial species based on their antigenic properties.
- Functional Support:
- Substrate for Autolytic Enzymes: Teichoic acids act as substrates for various autolytic enzymes. These enzymes contribute to the remodeling and turnover of the cell wall.
- Cell Division and Attachment: In Streptococcus pyogenes, LTAs are associated with M proteins. These proteins, together with LTAs, form microfibrils that facilitate bacterial attachment to host cells, which is crucial for the initiation of infection.
- Ion Uptake and Pathogen Interaction:
- Ion Uptake: Teichoic acids are involved in the uptake of ions, including magnesium. This function is essential for maintaining cellular homeostasis.
- Pathogen Binding: They assist in the binding of pathogenic organisms to host tissues, which is a key step in the infection process.
B. Teichuronic acid
Teichuronic acid is a structural component found in the cell walls of certain Gram-positive bacteria. It serves as an alternative to teichoic acids under conditions where phosphate availability is limited. Here’s an overview of its structure and function:
- Structure:
- Repeat Units: Teichuronic acid consists of repeating units of sugar acids. The primary sugar acids involved are D-glucuronic acid and N-acetylmannuronic acid.
- Polysaccharide Subunits: These sugar acids are linked together to form a polymeric structure. This structure incorporates both neutral sugars and acidic sugars.
- Formation and Function:
- Phosphate Limitation: Teichuronic acid is synthesized when the availability of phosphate is insufficient for the production of teichoic acids. Therefore, it acts as a substitute under these conditions.
- Cell Wall Composition: Besides teichuronic acid, the Gram-positive cell wall includes various neutral sugars such as mannose, arabinose, rhamnose, and glucosamine. It also contains acidic sugars like glucuronic acid and mannuronic acid. These sugars are integral components of the polysaccharides in the cell wall.
- Role and Significance:
- Structural Role: Teichuronic acid contributes to the structural integrity of the bacterial cell wall, similar to teichoic acids. It helps maintain cell wall stability and provides support to the bacterial envelope.
- Adaptation to Phosphate Availability: The ability to switch from teichoic acid to teichuronic acid synthesis allows bacteria to adapt to varying environmental conditions, particularly phosphate scarcity.
References
- Rajagopal M, Walker S. Envelope Structures of Gram-Positive Bacteria. Curr Top Microbiol Immunol. 2017;404:1-44. doi: 10.1007/82_2015_5021. PMID: 26919863; PMCID: PMC5002265.
- Chapot-Chartier, MP., Kulakauskas, S. Cell wall structure and function in lactic acid bacteria. Microb Cell Fact 13, S9 (2014). https://doi.org/10.1186/1475-2859-13-S1-S9
- https://en.wikipedia.org/wiki/Gram-positive_bacteria
- http://www.textbookofbacteriology.net/structure_9.html
- https://bio.libretexts.org/Bookshelves/Microbiology/Book%3A_Microbiology_(Kaiser)/Unit_1%3A_Introduction_to_Microbiology_and_Prokaryotic_Cell_Anatomy/2%3A_The_Prokaryotic_Cell_-_Bacteria/2.3%3A_The_Peptidoglycan_Cell_Wall/2.3A%3A_The_Gram-Positive_Cell_Wall#:~:text=The%20Gram%2Dpositive%20cell%20wall%20consists%20of%20many%20interconnected%20layers,interwoven%20through%20the%20peptidoglycan%20layers.
- https://open.oregonstate.education/generalmicrobiology/chapter/bacteria-cell-walls/
- https://www.slideshare.net/AshfaqAhmad52/bacterial-cell-wall-67925149