What is an Anticodon?
Anticodons play a crucial role in protein production by ensuring that the correct amino acids are incorporated in the appropriate positions. These sequences of nucleotides are found within transfer RNA (tRNA) molecules. tRNAs act as carriers, bringing the specific amino acids that match the instructions encoded in the messenger RNA (mRNA) to the ribosome, where protein synthesis takes place.
The process of protein production involves linking amino acids together to form a polypeptide chain. The correct placement of amino acids is essential because each amino acid possesses unique properties. Placing the wrong amino acid in a particular position can result in a nonfunctional or even harmful protein for the cell.
The anticodon is a trinucleotide sequence located on one arm of the tRNA molecule. It is specifically complementary to a corresponding codon, a sequence of three nucleotides, found on the mRNA. The pairing of the anticodon with the codon allows the tRNA to accurately read the mRNA instructions and bring the appropriate amino acid to the ribosome for incorporation into the growing protein chain.
In the process of translation, as the ribosome complex proceeds along the mRNA, tRNAs carrying amino acids enter and bind to the exposed mRNA. Only tRNAs with the correct anticodons can successfully form these bonds, ensuring that only the intended amino acids are added to the protein chain. This quality control mechanism helps maintain the fidelity and accuracy of protein synthesis.
The anticodon sequence is located on the anticodon loop of the tRNA molecule. It plays a vital role in the translation process, ensuring the precise alignment between the mRNA codons and the corresponding amino acids. The attachment of the amino acid to the 3′ adenosine end of the tRNA forms an aminoacyl tRNA, ready to deliver the correct amino acid to the ribosome for protein synthesis.
Overall, anticodons serve as essential molecular tools that facilitate the accurate translation of genetic information into functional proteins. By correctly pairing with mRNA codons, they ensure that the correct amino acids are incorporated into the growing polypeptide chain, enabling the cell to produce proteins with specific structures and functions.
Definition of Anticodon
An anticodon is a sequence of three nucleotides located on one arm of a transfer RNA (tRNA) molecule. It is specifically complementary to a corresponding codon, a sequence of three nucleotides found on the messenger RNA (mRNA). The anticodon plays a crucial role in protein synthesis during the process of translation. By pairing with the codon on the mRNA, the anticodon ensures that the correct amino acid is brought to the ribosome for incorporation into the growing polypeptide chain. The accurate alignment of the anticodon with the codon is essential for maintaining the fidelity and accuracy of protein production.
History of Anticodon
The history of the anticodon is closely tied to the discovery and understanding of the genetic code, which determines how the information encoded in DNA is translated into proteins. The genetic code was deciphered through a series of experiments conducted in the mid-20th century, and the role of the anticodon in this process gradually became clear.
In the early 1950s, scientists such as Francis Crick and Sydney Brenner proposed the concept of the “adaptor hypothesis.” They suggested that there must be intermediary molecules that could connect the genetic information stored in DNA with the synthesis of proteins. These intermediary molecules were later identified as transfer RNA (tRNA).
In 1958, scientists Severo Ochoa and Marianne Grunberg-Manago made a breakthrough discovery by isolating and characterizing tRNA. They found that tRNA molecules were small RNA molecules that carried specific amino acids to the ribosome, where protein synthesis occurred. However, the mechanism by which tRNA recognized the appropriate amino acids and codons remained unknown.
In the early 1960s, Marshall Nirenberg and Johann Matthaei conducted pioneering experiments to decipher the genetic code. They used synthetic RNA molecules with known repeating sequences to identify which amino acids were specified by each codon. Their experiments led to the discovery of the first codons, such as UUU (which codes for the amino acid phenylalanine) and AAA (which codes for lysine).
Around the same time, biochemist Har Gobind Khorana made significant contributions to understanding the genetic code. Khorana and his team synthesized RNA molecules with specific codons and used them to decipher the rest of the genetic code. Through their work, they identified the anticodons that corresponded to each codon.
The discovery of the anticodon helped scientists understand how tRNA molecules accurately recognize and bind to the appropriate codons on mRNA during protein synthesis. The complementarity between the anticodon and codon allows for the precise delivery of specific amino acids to the ribosome, ensuring the accurate assembly of the polypeptide chain.
Since these initial discoveries, further research has expanded our knowledge of the anticodon and its role in protein synthesis. Advances in molecular biology and genetics have provided insights into the structure and function of tRNA molecules, further deepening our understanding of the mechanisms underlying the translation process.
In summary, the history of the anticodon is intertwined with the deciphering of the genetic code and the elucidation of the molecular processes involved in protein synthesis. The discovery and characterization of the anticodon have been crucial in unraveling the mechanisms by which genetic information is translated into functional proteins.
Anticodon Principle
The Anticodon Principle, also known as the Wobble hypothesis, is a concept that explains how a limited number of tRNA molecules can recognize and bind to multiple codons, allowing for the efficient translation of the genetic code into proteins. This principle takes into account the imperfect pairing or weak hydrogen bonding between the third nucleotide of the codon (5′ to 3′) on the mRNA and the anticodon on the tRNA.
In the process of translation, the mRNA and tRNA are arranged antiparallel to each other. The first base of the codon on the mRNA binds with the third base of the anticodon on the tRNA to initiate the translation process. Each tRNA molecule carries a specific amino acid and binds to the complementary codon on the mRNA. In prokaryotes, this binding occurs on the 30s subunit, while in eukaryotes, it occurs on the 40s subunit.
The initial binding of the tRNA anticodon to the mRNA codon takes place at the A site of the ribosome. From there, the tRNA moves to the P site, where it is held together with the growing polypeptide chain of the protein. After the transfer of the amino acid, the tRNA moves to the E site (Exit) before being released from the ribosome.
The Anticodon Principle is based on the observation that the pairing between the codon and anticodon does not always follow strict Watson-Crick base pairing rules. Some tRNA molecules contain a nucleotide called Inosinate (I), which has the ability to form weaker hydrogen bonds with three different nucleotides: U (uracil), C (cytosine), and A (adenine). This weaker pairing allows for flexibility in the recognition and binding of multiple codons by a single tRNA molecule.
The inclusion of Inosinate in certain tRNA molecules enables a single tRNA to recognize multiple codons that differ in the third nucleotide position. Approximately 32 tRNA molecules can code for 61 different amino acids due to the imperfect pairing or “wobble” between the third nucleotide of the codon and the anticodon.
In summary, the Anticodon Principle, or Wobble hypothesis, explains how the presence of Inosinate in some tRNA molecules allows for the recognition and binding of multiple codons with different third nucleotides. This principle expands the coding capacity of tRNA molecules, enabling efficient and accurate protein synthesis despite the redundancy in the genetic code.
Anticodon Functions
The anticodon has several important functions in the process of translation, where it plays a crucial role in the formation of protein polypeptides.
- Amino Acid Specificity: The anticodon is responsible for determining the specificity of the amino acid carried by the tRNA. Each tRNA molecule is associated with a specific amino acid, and the anticodon ensures that the correct amino acid is delivered to the growing polypeptide chain during translation.
- Initiation of Translation: The first anticodon in both prokaryotes and eukaryotes is UAC, which binds to the AUG codon on the mRNA. This initiator anticodon of the tRNA pairs with the complementary codon on the mRNA, allowing the addition of the first amino acid to initiate the protein synthesis process. In prokaryotes, the first amino acid is N-formyl-methionine, while in eukaryotes, it is unmodified methionine.
- Termination of Translation: Just as the anticodon is responsible for the initiation of translation, it also plays a role in the termination of the process. Certain anticodon sequences on tRNA, such as AUU, AUC, and ACU, are associated with termination. When these anticodons encounter their corresponding stop codons on the mRNA, it signals the termination of protein synthesis.
How Anticodons Work
Anticodons play a crucial role in the process of protein synthesis, ensuring the accurate transfer of genetic information from the mRNA to the growing protein chain. Here’s how anticodons work:
- Transcription: The genetic information present in the cell’s genome is transcribed into RNA molecules by RNA polymerase. During this process, base-pairing rules are followed, where each DNA nucleotide is paired with its complementary RNA nucleotide. This results in the synthesis of an mRNA molecule that carries the information necessary for protein production.
- mRNA and Ribosome: The mRNA molecule, also known as messenger RNA, travels to a ribosome, which serves as the site of protein synthesis. The ribosome consists of two subunits, the small subunit, and the large subunit.
- Codon-Anticodon Pairing: At the ribosome, the rules of base-pairing are employed once again to ensure the accurate transfer of information. Each three-nucleotide sequence on the mRNA, known as a codon, is matched with a corresponding three-nucleotide sequence on a transfer RNA (tRNA) molecule, known as an anticodon. The anticodon carries the complementary bases to the mRNA codon, ensuring precise alignment and decoding of the genetic information.
- tRNA and Amino Acid Attachment: Transfer RNA molecules, or tRNAs, act as carriers for specific amino acids. Each tRNA molecule has one anticodon that corresponds to one mRNA codon and carries a specific amino acid attached to it. When the correct tRNA molecule with its specific anticodon finds the complementary mRNA codon, its amino acid is added to the growing protein chain.
- Bonding of Amino Acids: Enzymes catalyze the bonding of amino acids together as the tRNA anticodons bind to the correct mRNA codons. This process, known as peptide bond formation, occurs sequentially, resulting in the elongation of the protein chain.
- Release and Recharging: Once the tRNA molecule has added its amino acid to the protein chain, it dissociates from the mRNA and ribosome, ready to pick up a new amino acid and bind to a different mRNA codon. The tRNA anticodon maintains the RNA version of the nucleotide sequence from the original gene, as it was transcribed using complementary nucleotides during transcription.
By correctly pairing with the mRNA codons, anticodons ensure the accurate translation of genetic information into a functional protein. The sequential addition of amino acids guided by the tRNA anticodons allows for the precise assembly of the protein chain, ultimately determining its structure and function.
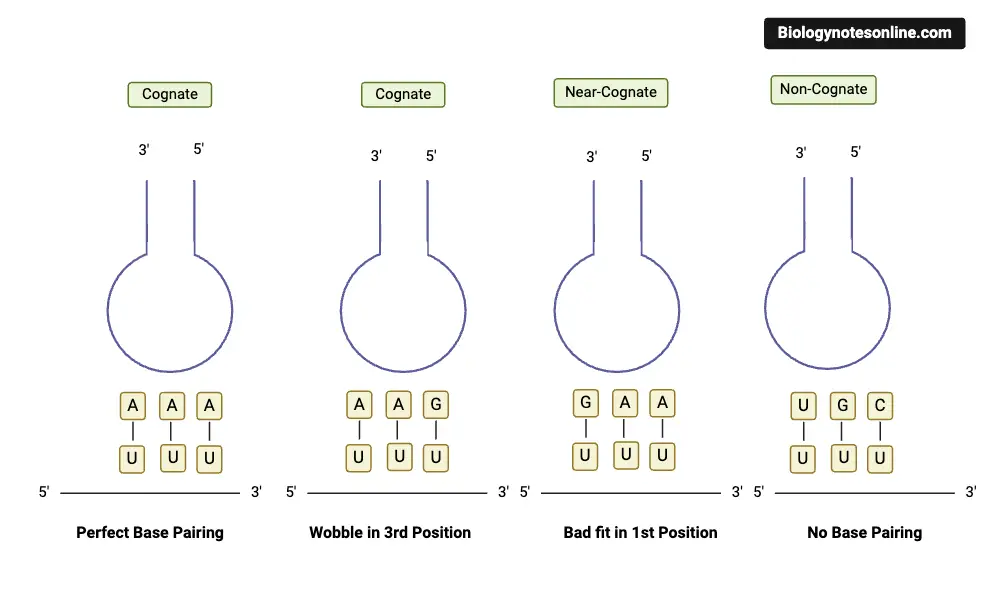
Types of interactions between anticodons of tRNA and codons of the mRNA
The four types of interactions between anticodons of tRNA and codons of the mRNA are as follows:
- Base pairing: The anticodon of tRNA and the codon of mRNA form complementary base pairs. Adenine (A) pairs with Uracil (U) in RNA, and Cytosine (C) pairs with Guanine (G). The base pairing between the anticodon and codon ensures the accurate reading of the genetic code during translation.
- Watson-Crick base pairing: The anticodon-codon interaction follows the Watson-Crick base pairing rules. Adenine (A) in the anticodon forms two hydrogen bonds with Uracil (U) in the codon, while Cytosine (C) in the anticodon forms three hydrogen bonds with Guanine (G) in the codon. These hydrogen bonds stabilize the interaction between the anticodon and codon.
- Wobble base pairing: In some cases, the third nucleotide of the codon and the first nucleotide of the anticodon can exhibit non-standard base pairing, known as wobble base pairing. This flexibility allows a single tRNA to recognize multiple codons with different nucleotides at the third position. For example, a tRNA with the anticodon 5′-GAA-3′ can recognize the codons 5′-UUU-3′ (Phenylalanine) and 5′-UUC-3′ (Phenylalanine) due to the wobble pairing between the third position of the codon and the first position of the anticodon.
- Specificity of anticodon-codon pairing: The interactions between anticodons and codons are highly specific, meaning each anticodon pairs with a specific codon. The specificity is determined by the genetic code, which is a set of rules that relate the sequence of nucleotides in mRNA codons to the amino acids they encode. The specificity ensures that the correct amino acids are incorporated into the growing polypeptide chain during translation.
What is the RNA Base Pairing Rule?
RNA base pairing rules dictate the specific interactions between nucleotides in RNA molecules, allowing for the accurate transfer and utilization of genetic information. Here are the key points about RNA base pairing rules:
- RNA Nucleotides: RNA is composed of four different nucleotides: Adenine (A), Cytosine (C), Guanine (G), and Uracil (U). These nucleotides are often represented by their first letters to simplify the representation of RNA sequences.
- Base Pairing Rules: The base pairing rules for RNA are as follows:
- Adenine (A) always pairs with Uracil (U).
- Cytosine (C) always pairs with Guanine (G).
- Guanine (G) always pairs with Cytosine (C).
- Uracil (U) always pairs with Adenine (A).
- Complementary Base Pairing: The base pairing rules establish complementary interactions between the nucleotides in an RNA molecule. These rules ensure that A always binds to U and C always binds to G, forming hydrogen bonds between the bases. This complementary base pairing is essential for maintaining the structural integrity of RNA and enables accurate replication, transcription, and translation processes.
In summary, RNA base pairing rules dictate the specific hydrogen bonding interactions between the four RNA nucleotides: A, C, G, and U. Adenine (A) pairs with Uracil (U), and Cytosine (C) pairs with Guanine (G). These rules ensure the precise alignment of nucleotides and facilitate the proper functioning of RNA in transferring and utilizing genetic information.
Where are anticodons located?
Anticodons are located on transfer RNA (tRNA) molecules. Each tRNA molecule has a specific anticodon sequence on one arm, known as the anticodon loop. The anticodon loop is positioned opposite to the amino acid attachment site on the tRNA molecule. The anticodon sequence consists of three nucleotides that are complementary to a specific codon on the messenger RNA (mRNA) during protein synthesis.
During translation, tRNA molecules bind to the mRNA at the ribosome, and the anticodon on the tRNA pairs with the complementary codon on the mRNA through base-pairing interactions. This complementary binding ensures the accurate alignment of the corresponding amino acid carried by the tRNA with the growing polypeptide chain. Thus, the anticodons on tRNA molecules play a crucial role in decoding the genetic information carried by mRNA and facilitating the precise incorporation of amino acids into the protein being synthesized.
Does mrna have codons or anticodons?
mRNA (messenger RNA) contains codons, not anticodons. Codons are specific sequences of three nucleotides on mRNA that encode for a particular amino acid or serve as start or stop signals during protein synthesis. Each codon on mRNA corresponds to a specific amino acid or a control signal in the genetic code.
On the other hand, tRNA (transfer RNA) molecules carry anticodons. Anticodons are sequences of three nucleotides on tRNA that are complementary to the codons on mRNA. The anticodon on tRNA base-pairs with the codon on mRNA during translation, ensuring the accurate pairing of amino acids with the mRNA codons.
So, while mRNA contains codons, tRNA carries anticodons that recognize and bind to the codons on mRNA to facilitate the correct incorporation of amino acids during protein synthesis.
What is a trna anticodon?
A tRNA anticodon refers to a specific sequence of three nucleotides on a transfer RNA (tRNA) molecule that is complementary to a corresponding codon on messenger RNA (mRNA) during protein synthesis. The tRNA anticodon plays a crucial role in ensuring accurate translation of the genetic code.
Each tRNA molecule carries a specific amino acid and has an anticodon region that recognizes and pairs with the complementary codon on mRNA. The base pairing between the tRNA anticodon and mRNA codon is facilitated by the rules of complementary base pairing in nucleic acids. This interaction allows the correct amino acid to be added to the growing polypeptide chain during protein synthesis. The tRNA anticodon acts as a molecular adaptor, ensuring that the correct amino acid is incorporated into the growing protein chain according to the genetic code.
What is the difference between a codon and an anticodon?
A codon and an anticodon are both sequences of three nucleotides, but they have different roles and locations within the process of protein synthesis.
- Codon:
- Location: Codons are found on the messenger RNA (mRNA) molecule.
- Function: Codons represent the genetic code and specify the sequence of amino acids in a protein. Each codon codes for a specific amino acid or serves as a start or stop signal.
- Complementary Pairing: Codons undergo complementary base pairing with anticodons during translation. For example, the codon “AUG” codes for the amino acid methionine and pairs with the anticodon “UAC” on the transfer RNA (tRNA) molecule.
- Anticodon:
- Location: Anticodons are found on the transfer RNA (tRNA) molecule.
- Function: Anticodons play a crucial role in recognizing and binding to the complementary codon on the mRNA molecule during translation. They ensure the accurate translation of the genetic code by pairing with the appropriate codon.
- Complementary Pairing: Anticodons undergo complementary base pairing with codons. The tRNA molecule carries a specific amino acid and binds to the corresponding codon on mRNA through its anticodon. This ensures that the correct amino acid is incorporated into the growing protein chain.
What is the purpose of an anticodon?
The purpose of an anticodon is to recognize and bind to the complementary codon on the messenger RNA (mRNA) during the process of protein synthesis.
During translation, the mRNA carries the genetic code in the form of codons, which are three-nucleotide sequences that specify the amino acids to be incorporated into the growing protein chain. The tRNA (transfer RNA) molecules act as adapters between the mRNA and the amino acids. Each tRNA molecule carries a specific amino acid and has an anticodon sequence that is complementary to a specific codon on the mRNA.
The anticodon of a tRNA molecule pairs with the codon on the mRNA through complementary base pairing. For example, if the codon on the mRNA is “AUG,” the corresponding anticodon on the tRNA would be “UAC.” This base pairing ensures that the correct amino acid is added to the growing protein chain according to the genetic code.
By binding to the complementary codon, the anticodon ensures the accurate translation of the genetic code, allowing the correct amino acid to be added at each step of protein synthesis. The proper pairing between the codon and anticodon is essential for the fidelity and specificity of protein synthesis.
How do codon and anticodon work together?
Codons and anticodons work together to ensure accurate translation of the genetic code during protein synthesis. Here’s how they work together:
- Codons: Codons are three-nucleotide sequences on the messenger RNA (mRNA) that specify a particular amino acid or a start/stop signal. Each codon represents a specific amino acid or a functional instruction.
- Anticodons: Anticodons are three-nucleotide sequences on the transfer RNA (tRNA) molecules. Each tRNA molecule carries a specific amino acid and has an anticodon that is complementary to a specific codon on the mRNA.
- Recognition and Binding: During translation, the ribosome moves along the mRNA, and tRNA molecules bind to the codons on the mRNA in a complementary manner. The anticodon of the tRNA recognizes and binds to the corresponding codon on the mRNA through base pairing.
- Complementary Base Pairing: The pairing between the codon and anticodon follows the rules of complementary base pairing in nucleic acids. Adenine (A) pairs with uracil (U), and cytosine (C) pairs with guanine (G). For example, if the codon on the mRNA is “AUG,” the corresponding anticodon on the tRNA would be “UAC.”
- Amino Acid Delivery: Each tRNA molecule carries a specific amino acid that is attached to its 3′ end. The anticodon ensures that the correct amino acid is brought to the ribosome, aligning with the codon on the mRNA. The ribosome catalyzes the formation of peptide bonds between the amino acids, ultimately leading to the synthesis of a protein chain.
By binding to the complementary codon on the mRNA, the anticodon ensures the accurate translation of the genetic code, allowing the correct amino acid to be added to the growing protein chain according to the genetic instructions. The codons and anticodons work together to maintain the fidelity and specificity of protein synthesis.
How does an anticodon participate in protein synthesis?
During protein synthesis, an anticodon participates in the process of translation by facilitating the accurate pairing between the codons on messenger RNA (mRNA) and the corresponding amino acids carried by transfer RNA (tRNA) molecules. Here’s how the anticodon participates:
- tRNA Binding: The first step is the binding of specific tRNA molecules to the ribosome, which is the cellular machinery responsible for protein synthesis. Each tRNA molecule carries a specific amino acid that is attached to its 3′ end.
- Complementary Base Pairing: The anticodon, which is a three-nucleotide sequence on the tRNA molecule, recognizes and binds to the complementary codon on the mRNA through base pairing. Adenine (A) pairs with uracil (U), and cytosine (C) pairs with guanine (G).
- Correct Alignment: The binding of the anticodon to the codon ensures the correct alignment of the tRNA molecule and its associated amino acid with the growing protein chain. The complementary base pairing between the codon and anticodon helps in selecting the appropriate tRNA molecule that carries the specific amino acid required for that position in the protein chain.
- Amino Acid Transfer: Once the tRNA molecule with the anticodon has paired with the codon on the mRNA, the ribosome facilitates the transfer of the amino acid carried by the tRNA to the growing protein chain. The ribosome catalyzes the formation of a peptide bond between the amino acids, resulting in the elongation of the protein chain.
- Translocation: After the amino acid transfer, the ribosome moves along the mRNA to expose the next codon, and the process repeats. The empty tRNA is released, and a new tRNA with the appropriate anticodon and amino acid binds to the ribosome, ensuring the accurate addition of amino acids to the protein chain.
How to convert mRNA codon to tRNA anticodon?
To convert an mRNA codon to a tRNA anticodon, you need to follow the rules of complementary base pairing. The mRNA codon and tRNA anticodon form a complementary pair, with adenine (A) pairing with uracil (U) and cytosine (C) pairing with guanine (G). Here’s how you can convert an mRNA codon to a tRNA anticodon:
- Identify the mRNA codon: The mRNA codon is a three-nucleotide sequence on the mRNA molecule. Each codon represents a specific amino acid.
- Determine the complementary bases: Based on the mRNA codon, determine the complementary bases for each nucleotide. For example, if the mRNA codon is AUG, the complementary bases would be UAC.
- Convert uracil (U) to thymine (T): In DNA, thymine is used instead of uracil. If you are working with DNA, replace each occurrence of uracil (U) in the mRNA codon with thymine (T) to obtain the DNA codon. For example, UAC would become TAC.
- Write the tRNA anticodon: Write the tRNA anticodon using the complementary bases obtained in step 2 or step 3. This is the anticodon that pairs with the mRNA codon. For example, if the mRNA codon is AUG, the tRNA anticodon would be UAC.
How to determine anticodon sequence?
To determine the anticodon for a specific mRNA codon, you need to consider the rules of complementary base pairing. The anticodon is a sequence of three nucleotides on a transfer RNA (tRNA) molecule that pairs with the complementary codon on the mRNA during protein synthesis. Here’s how you can determine the anticodon:
- Identify the mRNA codon: The mRNA codon is a three-nucleotide sequence on the mRNA molecule that specifies a particular amino acid.
- Determine the complementary bases: Based on the mRNA codon, determine the complementary bases for each nucleotide. Remember that in RNA, adenine (A) pairs with uracil (U), and cytosine (C) pairs with guanine (G). For example, if the mRNA codon is AUG, the complementary bases would be UAC.
- Write the anticodon: Write the tRNA anticodon using the complementary bases obtained in step 2. This is the anticodon that pairs with the mRNA codon. For example, if the mRNA codon is AUG, the anticodon would be UAC.
The anticodon on the tRNA molecule ensures accurate pairing with the mRNA codon during translation, allowing the correct amino acid to be added to the growing polypeptide chain. It’s important to note that the anticodon is complementary to the mRNA codon, not identical. The anticodon and codon form a base-pairing interaction that is essential for proper protein synthesis.
How to determine tRNA anticodon?
To determine the tRNA anticodon sequence, you need to follow the rules of base pairing with the mRNA codon. The anticodon sequence of tRNA is complementary to the mRNA codon and binds to it during protein synthesis. Here’s how you can determine the tRNA anticodon sequence:
- Identify the mRNA codon: The mRNA codon is a three-nucleotide sequence on the mRNA molecule that corresponds to a specific amino acid or a stop signal.
- Determine the complementary bases: Based on the mRNA codon, determine the complementary bases for each nucleotide. Remember that in RNA, adenine (A) pairs with uracil (U), and cytosine (C) pairs with guanine (G).
- Write the anticodon sequence: Write the anticodon sequence using the complementary bases obtained in step 2. The anticodon sequence is the three-nucleotide sequence on the tRNA molecule that will pair with the mRNA codon during protein synthesis. The order of the bases in the anticodon is opposite to that of the mRNA codon.
For example, let’s say the mRNA codon is AUG. The complementary bases would be UAC. To determine the tRNA anticodon sequence:
mRNA codon: AUG Complementary bases: UAC Anticodon sequence: 3′-AUG-5′
So, the tRNA anticodon sequence for the mRNA codon AUG is 3′-AUG-5′. This tRNA molecule with the anticodon AUG will pair with the mRNA codon AUG during translation, ensuring the correct amino acid is added to the growing polypeptide chain.
Remember that the tRNA anticodon sequence is specific to each tRNA molecule and is designed to recognize a particular mRNA codon. Multiple tRNA molecules exist, each with a specific anticodon sequence that corresponds to a particular amino acid.
Is anticodon DNA or RNA?
The anticodon is part of the transfer RNA (tRNA) molecule, which is made of RNA, not DNA. tRNA molecules are involved in protein synthesis and function as adapters between the mRNA codon and the corresponding amino acid.
During protein synthesis, tRNA molecules carry specific amino acids to the ribosome, where the mRNA codon is read. The anticodon, a sequence of three nucleotides on the tRNA molecule, is complementary to the mRNA codon. It allows the tRNA to recognize and bind to the mRNA, ensuring that the correct amino acid is added to the growing polypeptide chain.
Unlike DNA, which uses thymine (T) as one of its bases, RNA uses uracil (U) instead. So the anticodon sequence found in tRNA is composed of three RNA nucleotides, which can be complementary to the corresponding mRNA codon.
What are anticodons used for?
Anticodons are used in protein synthesis to ensure the accurate pairing of codons on mRNA with the corresponding amino acids carried by transfer RNA (tRNA) molecules. The primary function of anticodons is to recognize and bind to the complementary codon on the mRNA during translation.
During protein synthesis, the mRNA molecule contains a sequence of codons that encode the specific order of amino acids in a protein. Each codon consists of three nucleotides, and each codon corresponds to a specific amino acid. The tRNA molecules have anticodons that are complementary to the codons on the mRNA. This complementarity allows the tRNA to recognize and bind to the appropriate codon, ensuring the correct placement of the corresponding amino acid in the growing polypeptide chain.
The anticodon on the tRNA forms base pairs with the codon on the mRNA through complementary base pairing. For example, if the codon on the mRNA is “AUG,” the corresponding tRNA anticodon is “UAC,” which binds to the codon through base pairing (A-U and G-C). This binding of the anticodon to the codon brings the correct amino acid carried by the tRNA to the ribosome, allowing it to be incorporated into the growing protein chain.
In summary, the anticodons of tRNA molecules play a crucial role in ensuring the fidelity and accuracy of protein synthesis by facilitating the matching of codons on the mRNA with the appropriate amino acids carried by tRNA molecules.
What bases would be found in the complementary tRNA anticodon?
The bases found in the complementary tRNA anticodon would depend on the bases in the mRNA codon it is pairing with. The complementary base pairing rules in RNA are:
- Adenine (A) pairs with Uracil (U)
- Uracil (U) pairs with Adenine (A)
- Guanine (G) pairs with Cytosine (C)
- Cytosine (C) pairs with Guanine (G)
So, the bases in the tRNA anticodon would be complementary to the bases in the mRNA codon. For example, if the mRNA codon is “AUG,” then the complementary tRNA anticodon would be “UAC” since A pairs with U, U pairs with A, and G pairs with C.
It’s important to note that tRNA molecules have specific anticodons that are complementary to specific codons, allowing them to recognize and bind to the corresponding codon during protein synthesis. The complementary base pairing between the anticodon and codon ensures the accurate placement of the correct amino acid in the growing protein chain.
What is the anticodon for CCA?
The anticodon for the mRNA codon CCA would be GGU. In RNA, adenine (A) pairs with uracil (U), cytosine (C) pairs with guanine (G), and guanine (G) pairs with cytosine (C). Therefore, to find the anticodon, we replace each base in the codon with its complementary base.
In this case, the codon CCA has the bases C, C, and A. Replacing C with G and A with U, we get the anticodon GGU.
What is the anticodon for leucine?
The anticodon for leucine can vary depending on the specific codon for leucine. Leucine is encoded by multiple codons, including UUA, UUG, CUU, CUC, CUA, and CUG.
The corresponding anticodon for each of these codons would be:
- UUA (Leucine codon): The anticodon would be AAU.
- UUG (Leucine codon): The anticodon would be CAU.
- CUU (Leucine codon): The anticodon would be GAA.
- CUC (Leucine codon): The anticodon would be GAG.
- CUA (Leucine codon): The anticodon would be UAG.
- CUG (Leucine codon): The anticodon would be CAG.
Remember that the anticodon is the complementary sequence to the codon, so the bases in the anticodon are paired with the corresponding bases in the codon using the rules of base pairing (A-U and G-C).
What is the anticodon for valine?
The codons for valine are GUC, GUA, GUG, and GUU. The anticodons for these codons would be CAG, CUA, CAC, and CAA, respectively.
To summarize:
- Codon GUC (Valine) → Anticodon CAG
- Codon GUA (Valine) → Anticodon CUA
- Codon GUG (Valine) → Anticodon CAC
- Codon GUU (Valine) → Anticodon CAA
The anticodon is the complementary sequence to the codon, so the bases in the anticodon are paired with the corresponding bases in the codon using the rules of base pairing (A-U and G-C).
What is the complementary tRNA anticodon for the mRNA codon?
To find the complementary tRNA anticodon for an mRNA codon, you need to follow the base-pairing rules. The complementary bases are as follows:
- Adenine (A) pairs with Uracil (U) in RNA.
- Cytosine (C) pairs with Guanine (G).
Therefore, to determine the complementary tRNA anticodon for an mRNA codon, you would replace each base in the mRNA codon as follows:
- Adenine (A) in the mRNA codon pairs with Uracil (U) in the tRNA anticodon.
- Cytosine (C) in the mRNA codon pairs with Guanine (G) in the tRNA anticodon.
- Guanine (G) in the mRNA codon pairs with Cytosine (C) in the tRNA anticodon.
- Uracil (U) in the mRNA codon pairs with Adenine (A) in the tRNA anticodon.
For example, if the mRNA codon is “AUG,” the complementary tRNA anticodon would be “UAC” because:
- A (Adenine) in the mRNA codon pairs with U (Uracil) in the tRNA anticodon.
- U (Uracil) in the mRNA codon pairs with A (Adenine) in the tRNA anticodon.
- G (Guanine) in the mRNA codon pairs with C (Cytosine) in the tRNA anticodon.
Keep in mind that the tRNA anticodon is the reverse complement of the mRNA codon.
Examples of Anticodons
Anticodons are specific sequences of nucleotides that correspond to codons on the mRNA and carry specific amino acids during protein synthesis. Here are examples of anticodons and their corresponding amino acids:
- Phenylalanine: AAA, AAG
- Leucine: AAU, AAC, GAA, GAG, GAU, GAC
- Isoleucine: UAA, UAG, UAU
- Methionine: UAC
- Valine: CAA, CAG, CAU, CAC
- Serine: AGA, AGG, AGU, AGC, UCA, UCG
- Proline: GGA, GGG, GGU, GGC
- Threonine: UGA, UGG, UGU, UGC
- Alanine: CGA, CGG, CGU, CGC
- Tyrosine: AUA, AUG
- Histidine: GUA, GUG
- Glutamine: GUU, GUC
- Asparagine: UUA, UUG
- Lysine: UUU, UUC
- Aspartic acid: CUA, CUG
- Glutamic acid: CUU, CUC
- Cysteine: ACA, ACG
- Tryptophan: AGU, ACC
- Arginine: GCA, GCG, GCU, GCC, UCU, UCC
- Glycine: CCA, CCG, CCU, CCC
These examples demonstrate the diverse range of anticodons that exist in the genetic code, each corresponding to a specific amino acid. The accurate pairing between anticodons and codons ensures the proper incorporation of the corresponding amino acids into the growing polypeptide chain during protein synthesis.
Difference between codon and anticodon – Codon vs Anticodon Table
| Codon | Anticodon | |
|---|---|---|
| Definition | A three-nucleotide sequence on mRNA that codes for a specific amino acid or serves as a start/stop signal in protein synthesis. | A three-nucleotide sequence on tRNA that is complementary to the codon on mRNA during protein synthesis. |
| Location | Found on mRNA | Found on tRNA |
| Function | Determines the sequence of amino acids in a protein during translation. | Helps to ensure the accurate pairing of amino acids with the mRNA codons during translation. |
| Binding | Binds with complementary anticodons on tRNA during translation. | Binds with complementary codons on mRNA during translation. |
| Base Pairing | Follows base pairing rules: A-U, C-G, G-C, U-A. | Follows base pairing rules: A-U, C-G, G-C, U-A (complementary to codons). |
| Number of Codons/Anticodons | 64 possible codons (4^3) encode for 20 amino acids and stop signals. | 64 possible anticodons (4^3) correspond to the 64 codons on mRNA. |
| Initiation and Termination | Certain codons (e.g., AUG) serve as start codons to initiate protein synthesis, and others (e.g., UAA, UAG, UGA) act as stop codons to terminate translation. | Anticodons play a role in the initiation and termination of translation by binding to the corresponding start and stop codons on mRNA. |
References
- Liljas, A. (2001). Anticodons. Encyclopedia of Genetics, 78–79. doi:10.1006/rwgn.2001.0059
- https://biologydictionary.net/anticodon/
- https://www.dictionary.com/compare-words/codon-vs-anticodon
- https://www.merriam-webster.com/dictionary/anticodon
- https://study.com/academy/lesson/codon-recognition-how-trna-and-anticodons-interpret-the-genetic-code.html
- https://www.pearson.com/channels/biology/asset/fa16d43f/the-anticodon-of-a-particular-trna-molecule-is-a-complementary-to-the-correspond
- https://rnajournal.cshlp.org/content/early/2021/07/27/rna.078863.121
- https://www.waynesword.net/codons.htm