What is Oxidase Test?
Oxidase test is a biochemical test used in microbiology. It is used to identify bacteria producing cytochrome c oxidase enzyme. This enzyme is present in electron transport chain of aerobic respiration. It helps in transfer of electron to oxygen for formation of water or hydrogen peroxide.
In this test a colorless reagent is used. It is generally tetramethyl-p-phenylenediamine dihydrochloride. It acts as artificial electron donor. When bacteria contains cytochrome c oxidase the reagent is oxidized. It produces deep purple or dark blue colour.
The colour change occurs rapidly within 10 to 30 seconds. It indicates positive oxidase test. If no colour change is seen within this time it indicates negative result. This test is used for differentiating oxidase positive bacteria like Pseudomonas and Neisseria from oxidase negative bacteria like Escherichia coli and other Enterobacteriaceae.
Objectives of Oxidase Test
- To detect presence of cytochrome c oxidase enzyme (indophenol oxidase) in bacterial electron transport chain.
- To biochemically characterize the bacteria. It helps in identification of unknown bacterial isolate.
- To differentiate Enterobacteriaceae (generally oxidase negative) from other Gram negative bacilli like Pseudomonadaceae (typically oxidase positive).
- To help in presumptive identification of oxidase positive genera like Pseudomonas Neisseria Moraxella Campylobacter Aeromonas Vibrio and Pasteurella.
- To distinguish oxidase positive Neisseria species from other Gram negative diplococci. It also separates it from oxidase negative Gram positive cocci like Staphylococcus and Streptococcus.
- To rapidly screen and identify some fastidious pathogenic bacteria like Vibrio cholerae and Neisseria gonorrhoeae.
- To separate oxidase positive Plesiomonas shigelloides from morphologically similar oxidase negative Shigella species.
Principle of Oxidase Test
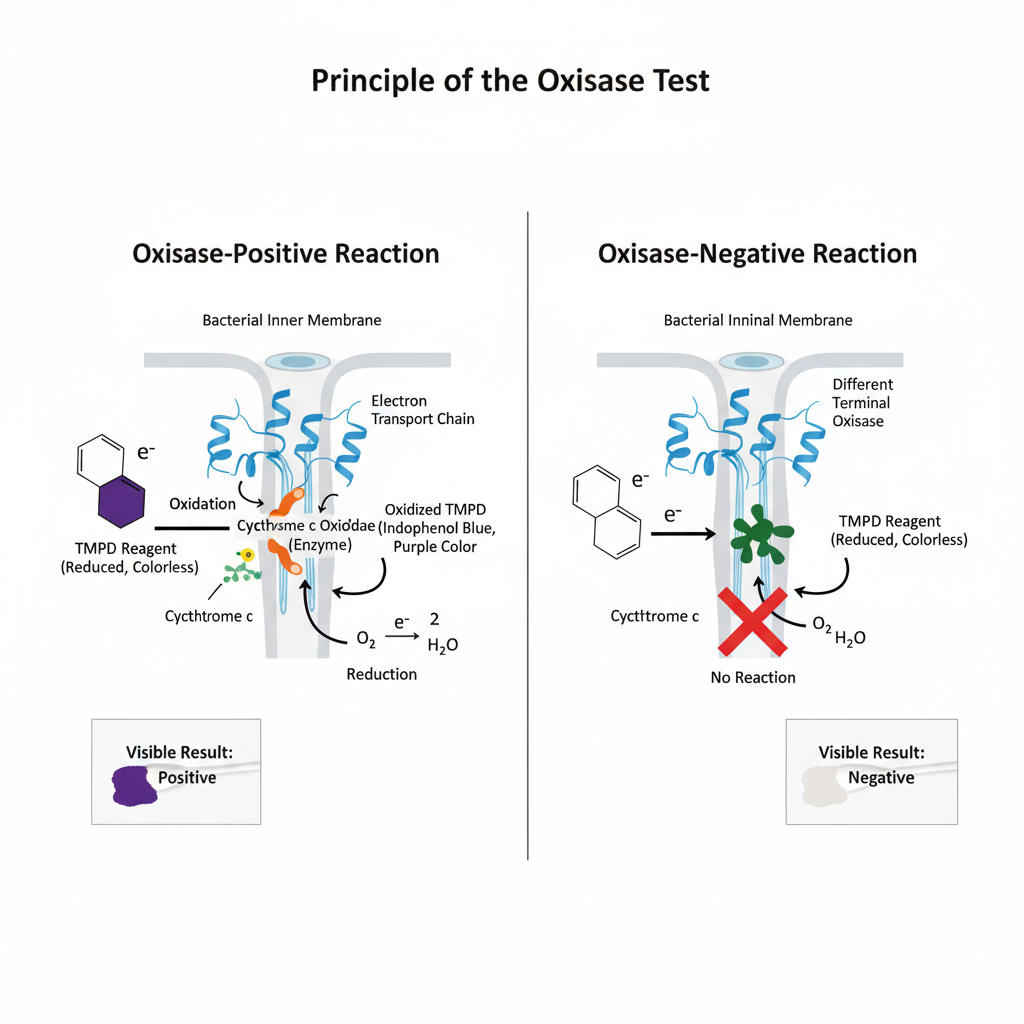
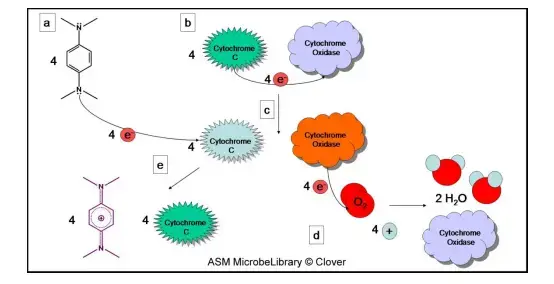
Principle of Oxidase test is based on detection of cytochrome c oxidase enzyme. It is an intracellular enzyme. It acts as final component of electron transport chain in aerobic respiration. It catalyzes transfer of electron from donor compound to molecular oxygen. Oxygen acts as final electron acceptor. Water or hydrogen peroxide is formed.
In this test an artificial electron donor is used. It is a colourless redox dye like tetramethyl-p-phenylenediamine dihydrochloride. When bacterial sample contains cytochrome c oxidase it oxidizes the dye rapidly. Electron is removed from the reagent. It converts colourless reagent into dark purple or blue coloured compound. This compound is known as indophenol blue (Wurster’s blue). It indicates positive oxidase test.
If bacteria lacks cytochrome c oxidase enzyme it cannot oxidize the reagent. Even if aerobic respiration is present by different terminal oxidase the reagent remains colourless. It indicates negative oxidase test.
Requirements for Oxidase Test
- Fresh pure bacterial culture (18–24 hours old). Colonies should be taken from non fermentative solid media like nutrient agar or tryptic soy agar. Media with high glucose or dyes should be avoided due to interference.
- Oxidase reagent (fresh redox dye solution). Kovacs’ oxidase reagent is used commonly (1% tetramethyl-p-phenylenediamine dihydrochloride). Other reagent like Gordon and McLeod’s reagent or Gaby and Hadley reagent can also be used.
- Inoculating tools like sterile wooden applicator stick plastic loop or platinum loop. Iron or nichrome loop should be avoided because false positive result may occur.
- Reaction surface like sterile filter paper (Whatman No. 1) or oxidase disc/strip or sterile cotton swab (depending on method).
- Glassware or plasticware like clean petri plate glass slide or test tube (for liquid broth method).
- Timing device like stopwatch or timer. Result should be read within 10 to 30 seconds to avoid false positive due to air oxidation.
- Control organism (oxidase positive bacteria like Pseudomonas aeruginosa and oxidase negative bacteria like Escherichia coli). It is used to check reagent working.
- Miscellaneous laboratory items like distilled water (for moistening disc/strip) sterile forceps Bunsen burner and biohazard disposal or sharps container.
- Safety materials like PPE (lab coat and protective gloves).
Procedure of Oxidase Test
General precautions before test
- Fresh bacterial culture (18–24 hours) should be used. Fresh oxidase reagent should be taken.
- Nichrome loop or iron loop should not be used because false positive result is obtained. Sterile wooden stick plastic loop or platinum loop is used.
Filter paper method
- A sterile filter paper is placed on clean glass slide or inside petri dish.
- The filter paper is moistened by adding 1–2 drops of oxidase reagent.
- A well isolated colony is picked by sterile wooden stick or plastic loop. It is smeared on moistened area.
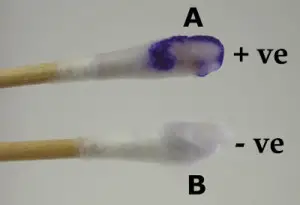
- Colour change is observed within 10 to 30 seconds. Deep purple or dark blue colour indicates positive test.
Direct plate method
- 1–3 drops of oxidase reagent is added directly on well isolated colonies on agar plate. Whole plate should not be flooded.
- The colonies are observed on agar for colour change within 10 to 30 seconds. Dark purple or blue colour indicates positive test.
Swab method
- A sterile cotton swab is moistened by dipping in oxidase reagent.
- The moistened swab is touched gently on well isolated colony.
- The tip of swab is observed for colour change within 10 to 30 seconds. Deep purple or blue colour indicates positive test.
Impregnated disc or strip method
- Oxidase disc or test strip is placed on clean petri dish or glass slide. It is moistened with sterile distilled water if required.
- A bacterial colony is picked by sterile wooden stick or plastic loop. It is rubbed gently on disc or strip.
- The disc or strip is observed within 10 to 30 seconds. Deep purple colour indicates positive test.
Test tube method (Gaby-Hadley method)
- Liquid culture is prepared by inoculating bacteria in nutrient broth. It is incubated for 18 to 24 hours.
- 0.2 mL Reagent A (1% α-naphthol) and 0.3 mL Reagent B (1% p-amino dimethylaniline oxalate) is added into test tube.
- The tube is shaken vigorously. Mixing and oxygenation is done.
- Broth colour change is observed within 15 to 30 seconds. Blue or purple colour indicates positive oxidase test.
Result and Interpretation of Oxidase Test
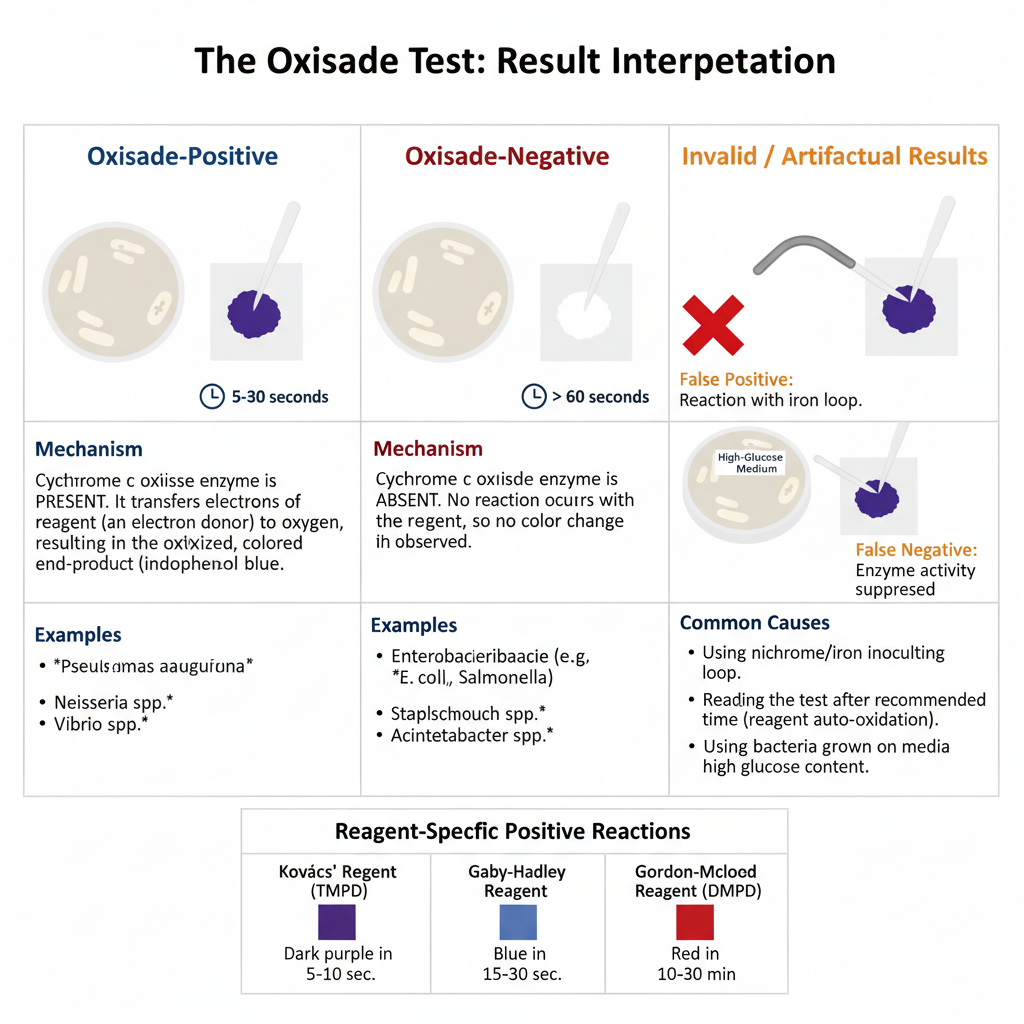
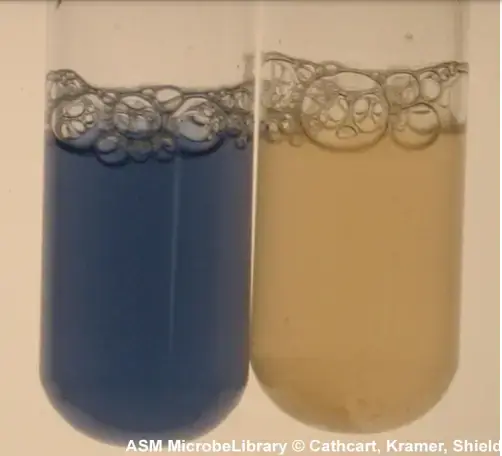
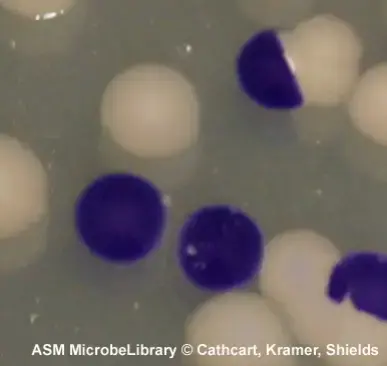
- Strong positive result– Deep purple or blue colour is formed within 10 to 30 seconds.
- Weak or delayed positive result– Purple colour is developed between 30 to 60 seconds. In Gaby-Hadley tube method colour may appear up to 2 to 3 minutes.
- Negative result– No colour change is seen within 60 seconds. Any colour developed after 60 seconds should be ignored because it is due to auto oxidation of reagent by oxygen.
- Positive interpretation– Positive result confirms presence of cytochrome c oxidase enzyme. Some common oxidase positive bacteria are Pseudomonas Neisseria Vibrio and Campylobacter.
- Negative interpretation– Negative result indicates absence of cytochrome c oxidase enzyme. It is used for identifying Enterobacteriaceae. Some oxidase negative organism are Escherichia coli Klebsiella and Salmonella.



List of Oxidase-Positive & Oxidase-Negative Organisms
Here is a list of organisms categorized by their typical results in the oxidase test:
Oxidase-Positive Organisms
- Pseudomonas species (e.g., Pseudomonas aeruginosa)
- Neisseria species (e.g., Neisseria gonorrhoeae, Neisseria meningitidis)
- Vibrio species (e.g., Vibrio cholerae, Vibrio parahaemolyticus)
- Campylobacter species (e.g., Campylobacter jejuni)
- Aeromonas species
- Alcaligenes species
- Moraxella species
- Pasteurella species (e.g., Pasteurella multocida)
- Brucella species
- Helicobacter pylori
- Legionella pneumophila
- Micrococcus species (e.g., Micrococcus luteus)
- Bordetella pertussis
- Plesiomonas shigelloides (the only member of the Enterobacteriaceae family that is oxidase-positive)
- Burkholderia cepacia and Burkholderia pseudomallei
Oxidase-Negative Organisms
- Most members of the Enterobacteriaceae family, including:
- Escherichia coli
- Klebsiella species (e.g., Klebsiella pneumoniae)
- Salmonella species
- Proteus species (e.g., Proteus mirabilis)
- Shigella species
- Staphylococcus species (e.g., Staphylococcus aureus, though a few rare species are positive)
- Streptococcus species
- Acinetobacter species
- Stenotrophomonas maltophilia (formerly Pseudomonas maltophilia)
- Listeria species
- Mycoplasma species
- Burkholderia gladioli
Note: Some organisms, such as certain species of Haemophilus and Pasteurella, can produce variable oxidase results.
Quality Control of Oxidase Test
Standard oxidase test
- Positive control– Pseudomonas aeruginosa (ATCC 27853). It gives rapid deep purple or blue colour.
- Negative control– Escherichia coli (ATCC 25922). It shows no colour change.
Microdase (modified oxidase) test
- Positive control– Micrococcus luteus (ATCC 10240). It gives blue or purple colour (for Gram positive cocci testing).
- Negative control– Staphylococcus aureus (ATCC 25923). It shows no colour change.
Precautions during Oxidase Test
- Fresh oxidase reagent should be used. Reagent is unstable and it is photo sensitive. It undergoes auto oxidation in light and air. If reagent is already dark blue or purple it should be discarded.
- Metal inoculating loop should be avoided. Nichrome steel and iron loop gives false positive result due to oxidation of reagent. Sterile wooden applicator stick plastic loop or platinum loop is used.
- Fresh bacterial culture should be taken (18 to 24 hours old). Older culture gives weak reaction. False negative result can be obtained due to low metabolic activity.
- Glucose rich media should be avoided. Media like MacConkey agar or TSI agar is not used. Acid produced during fermentation inhibits cytochrome oxidase enzyme. False negative result is obtained.
- Blood agar should not be used for direct test. Hemoglobin shows pseudo peroxidase activity and it can give positive like reaction. Media containing dyes (like EMB) should also be avoided because colour change is interfered or it is masked.
- Time should be monitored strictly. Result is read within 10 to 30 seconds. It should not be read beyond 60 seconds for Kovacs’ reagent. Any colour developed after time limit is ignored (false positive due to reagent auto oxidation by atmospheric oxygen).
- Sub culturing should be done before test. Oxidase reagent is toxic and it kills microorganism.
- Well isolated pure colony should be used. Mixed culture gives inaccurate reading.
- Pigmented colonies are interpreted carefully. Distinct dark blue black or purple colour should be seen. Natural pigment should not be mistaken as positive test.
- Strict anaerobes should not be tested. Oxidase test is meant for organism utilizing oxygen.
Uses of Oxidase Test
- It is used to detect cytochrome c oxidase enzyme in bacteria.
- It is used to differentiate Enterobacteriaceae (oxidase negative like E. coli) from oxidase positive Gram negative bacilli like Pseudomonadaceae.
- It is used for presumptive identification of pathogenic bacteria like Neisseria Vibrio cholerae Campylobacter jejuni and Pasteurella species.
- It is used to differentiate oxidase positive Neisseria and Moraxella from other Gram negative diplococci like Acinetobacter (oxidase negative).
- It is used to distinguish oxidase positive Plesiomonas shigelloides from oxidase negative Shigella species in stool culture (morphology may be similar).
- It is used in Microdase test (modified oxidase test) for Gram positive catalase positive cocci. It differentiates Micrococcus (oxidase positive) from Staphylococcus (oxidase negative).
- It is used for species level differentiation in some non enteric Gram negative genera. Example Burkholderia cepacia (positive) is differentiated from Burkholderia gladioli (negative). It also helps in differentiating some species within Haemophilus and Bordetella.
Limitations of Oxidase Test
- Oxidase reagent is unstable. It is photo sensitive and it undergoes auto oxidation in presence of light and air. Old or degraded reagent gives unreliable result. False negative result can be obtained. Reagent should be prepared fresh (often daily).
- Inoculating tool creates interference. Iron steel or nichrome loop catalyzes oxidation of reagent. False positive result is obtained. Only sterile wooden stick plastic loop or platinum loop should be used.
- Culture age affects test. Fresh culture (18 to 24 hours) should be used. Older colonies shows low metabolic activity. Respiratory enzyme synthesis decreases and weak or false negative reaction is produced.
- Media interference is present. Glucose rich media like MacConkey agar or TSI agar produces acidic byproduct. It inhibits cytochrome oxidase enzyme and false negative test is obtained. Media containing dyes also affects result and colour change may be masked.
- Blood agar gives problem. Colonies taken directly from blood agar may give false positive result because hemoglobin shows pseudo peroxidase activity.
- Timing requirement is strict. Result should be observed within 10 to 30 seconds. Colour developed after time limit is generally due to auto oxidation of reagent not due to bacterial enzyme activity.
- Oxidase reagent is toxic. It kills the microorganism. In direct plate method colony becomes non viable quickly. Sub culturing should be done before adding reagent or immediately after.
- It is only presumptive test. Confirmatory identification is not done by oxidase test alone. Closely related oxidase positive bacteria cannot be differentiated. It is interpreted with Gram stain and other biochemical test.
- Pure culture is required. Well isolated colony should be used. Mixed culture gives misleading result.
- Pigmented organism creates confusion. Natural pigment (like Pseudomonas aeruginosa) may interfere in interpretation of colour change.
References
- AAT Bioquest. (2024, May 9). What are the limitations of oxidase tests?
- abalidu91. (n.d.). Oxidase test procedure and results. Scribd.
- Abbott, I. J., & Peleg, A. Y. (2015). Stenotrophomonas, Achromobacter, and nonmelioid Burkholderia species: Antimicrobial resistance and therapeutic strategies. Seminars in Respiratory and Critical Care Medicine, 36(1), 99–110. https://doi.org/10.1055/s-0034-1396929
- American Society for Microbiology. (2019, August 12). Oxidase test.
- Centers for Disease Control and Prevention. (n.d.). How to perform an oxidase test [Video transcript].
- Centers for Disease Control and Prevention. (n.d.). How to perform the oxidase test [Video]. YouTube.
- Chauhan, A. (2025, May 19). Oxidase test procedure for bacterial identification. Flabs.
- Chavan, D. D., Khatoon, H., Anokhe, A., & Kalia, V. (2022). Oxidase test: A biochemical method in bacterial identification. AgriCos e-Newsletter, 3(1), 31–33.
- Clinical and biochemical foundations of the oxidase test in diagnostic microbiology. (n.d.).
- Dahal, P. (2023, April 4). Oxidase test- Principle, procedure, types, results, uses. Microbe Notes.
- Dalynn Biologicals. (2014, October). Oxidase reagent [Technical Data].
- DrChika. (2022, December 28). Oxidase test. Microbiology Class.
- GMP Plastic. (2025, April 2). Oxidase test – A test for detecting cytochrome c oxidase in bacteria.
- Hardy Diagnostics. (2020). Neisseria test – tablets for the detection of Neisseria gonorrhoeae and other species.
- Harrens Lab. (n.d.). What is oxidase test microbiology?
- HiMedia Laboratories. (2022, March). R028 Gaby-Hadley reagent B [Technical Data].
- Key Scientific Products. (n.d.). K520 oxidase test strips discussion.
- Michigan State University. (2021). Oxidase test. Virtual Interactive Bacteriology Laboratory.
- Modified oxidase and benzidine tests for separation of staphylococci from micrococci – PMC. (n.d.).
- Neisseria spp. (n.d.).
- Oxidase test. (2025, July 28). In Wikipedia.
- Periaiah, P., Antony, T., & Samuel, S. (2024). Identification of Burkholderia cepacia complex: Comparing conventional, automated, and molecular methods in a tertiary care center. Cureus, 16(10), e70847. https://doi.org/10.7759/cureus.70847
- Picmonic. (n.d.). Neisseria meningitidis – Gram negative cocci.
- Plesiomonas shigelloides. (2025, May 23). In Wikipedia.
- Plesiomonas shigelloides: An unusual cause of septic abortion – PMC. (n.d.).
- Rapid, modified oxidase test for oxidase-variable bacterial isolates – PMC – NIH. (n.d.).
- Reassessment of Stenotrophomonas maltophilia phenotype – PMC – NIH. (n.d.).
- Reynolds, J. (2024, February 6). 27: Oxidase test. Biology LibreTexts.
- Roy, B., Das, T., & Bhattacharyya, S. (2023). Overview on old and new biochemical test for bacterial identification. Journal of Surgical Case Reports and Images, 6(5). https://doi.org/10.31579/2690-1897/163
- Shields, P., & Cathcart, L. (2010, November 11). Oxidase test protocol. American Society for Microbiology.
- Sigma-Aldrich. (2018). 70439 Oxidase test. MilliporeSigma.
- Tetramethylphenylenediamine. (2025, March 22). In Wikipedia.
- The brief case: Wound infection with Plesiomonas shigelloides following a freshwater injury – PMC. (n.d.).
- van Pelt, C., Verduin, C. M., Goessens, W. H. F., Vos, M. C., Tümmler, B., Segonds, C., Reubsaet, F., Verbrugh, H., & van Belkum, A. (1999). Identification of Burkholderia spp. in the clinical microbiology laboratory: Comparison of conventional and molecular methods. Journal of Clinical Microbiology, 37(7), 2158–2164. https://doi.org/10.1128/jcm.37.7.2158-2164.1999
- VetBact. (2023, March 29). Plesiomonas shigelloides.