What is Rolling circle replication?
Rolling circle replication is a unidirectional and asymmetric process of nucleic acid replication. It is the process by which multiple copies of circular DNA or RNA molecules is synthesized rapidly. This mechanism is commonly seen in bacterial plasmids, bacteriophages and certain plant and animal viruses.
In this type of replication both strands are not synthesized simultaneously. The synthesis of leading strand and lagging strand are separated physically and temporally. This process occurs when a special initiator protein recognizes the origin of replication on a circular DNA molecule.
The replication begins with the action of Rep protein. It is the protein which identifies the double-strand origin (DSO) and introduces a nick in one strand of the circular DNA. The Rep protein binds covalently to the 5′ phosphate end of the nicked strand and leaves a free 3′ hydroxyl end. This 3′ OH group acts as a primer for DNA polymerase.
In this step DNA polymerase extends the 3′ end continuously by moving around the intact circular template. As the polymerase moves, the nicked strand is displaced gradually. The displaced strand is peeled off as a single-stranded DNA. The reaction continues until one complete round of the circle is achieved.
After completion of one round, the Rep protein cleaves the newly synthesized strand and ligates the ends. A closed circular single-stranded DNA (ssDNA) molecule is released. This is referred to as the displaced product.
The single-stranded DNA is then converted into double-stranded DNA. It is carried out by host enzymes which recognize a specific site known as single-strand origin (SSO). Lagging strand synthesis is initiated and complementary strand is formed. Finally a complete double-stranded circular DNA molecule is produced.
Rolling circle replication is also adapted in laboratory techniques. It is used in Rolling Circle Amplification (RCA), which is an isothermal amplification method. In this technique strand displacing DNA polymerase (such as phi29 DNA polymerase) is used to produce long DNA molecules containing repeated sequences of the template. This method is widely used in molecular diagnosis, biosensing and sequencing studies.
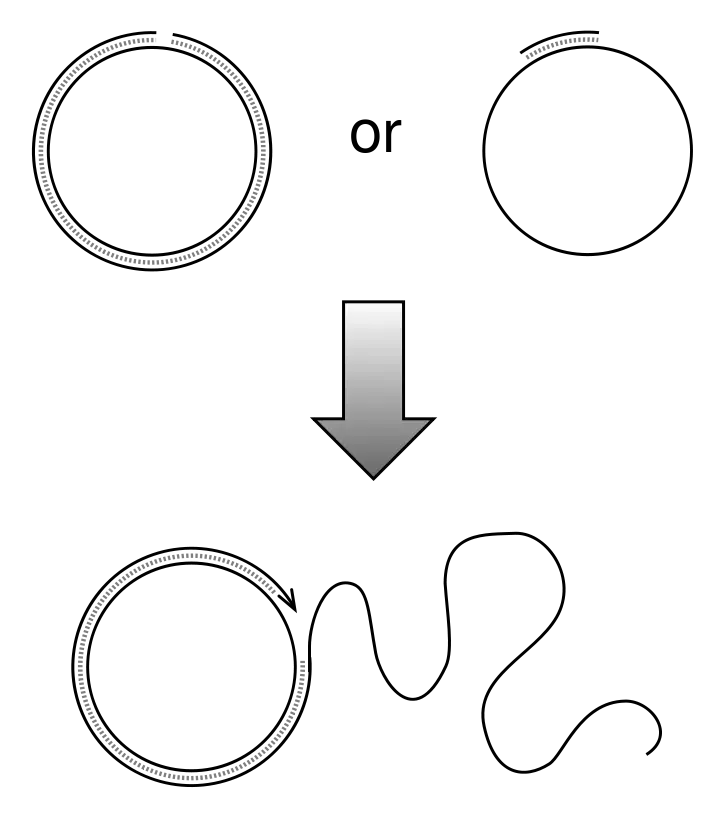

What Is Circular DNA?
- Circular DNA is a closed loop DNA molecule which does not possess free ends.
- It is structurally different from the linear DNA present in most eukaryotic organisms.
- It is the chromosomal DNA found in prokaryotic organisms.
- In bacteria and archaea the genetic material is present in circular form.
- Circular DNA is also present in plasmids and bacteriophages.
- It is found in some eukaryotic viruses and also in mitochondria of mammals and chloroplasts of plants.
- Since it does not have free ends, special types of replication is followed.
- These are– Theta (θ) replication, Rolling circle replication and Strand displacement (D-loop) replication.
- Theta replication is commonly seen in bacteria such as Escherichia coli.
- During this process a replication bubble is formed and the structure resembles the Greek letter theta (θ).
- Rolling circle replication is a unidirectional mode of replication.
- It produces multiple copies of circular template and forms a structure resembling the Greek letter sigma (σ).
- Strand displacement replication is also referred to as D-loop replication.
- This type of replication occurs in circular DNA of mammalian mitochondria.
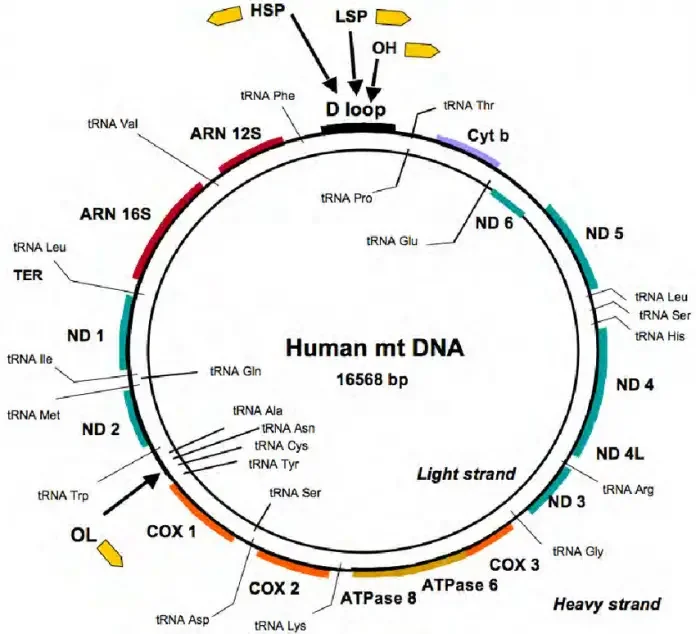
Mechanism of Rolling circle replication
Rolling circle replication is the process by which multiple copies of circular DNA is synthesized in a unidirectional manner. It is completed in different phases which are described below.
Initiation
In the first step, an initiator protein known as Rep protein recognizes a specific sequence on the circular double stranded DNA.
This specific region is referred to as Double-Strand Origin (DSO).
The Rep protein catalyzes a reaction and introduces a nick in one strand of the DNA helix.
The nick is produced in the phosphodiester bond of the plus (+) strand.
After making the cut, the Rep protein remains covalently attached to the 5′ phosphate end of the nicked strand.
This attachment is important for controlling the replication process.
Due to nick formation a free 3′ hydroxyl (3′-OH) end is generated.
This 3′-OH end acts as a primer for DNA synthesis and no RNA primer is required in this step.
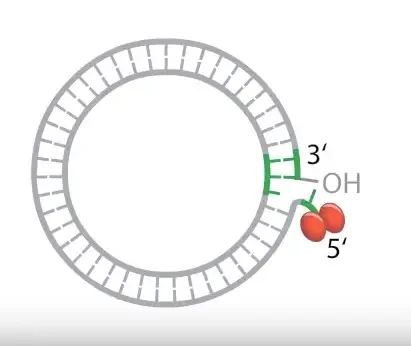
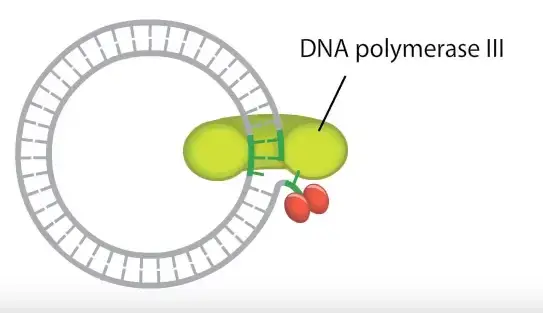
Elongation and Strand Displacement
In this step host DNA polymerase (mainly DNA polymerase III) binds to the free 3′-OH end.
It starts adding nucleotides by reading the intact inner strand as template.
As DNA polymerase moves forward, DNA helicase unwinds the parental double helix ahead of the replication fork.
This unwinding is necessary for continuous synthesis.
The movement of polymerase results in displacement of the previously nicked strand.
The displaced strand is peeled off and forms a long single stranded DNA (ssDNA) tail.
The exposed single stranded DNA is unstable and may undergo degradation.
Therefore it is coated rapidly by Single-Strand Binding (SSB) proteins which prevents folding and damage.
Termination and Release
DNA synthesis continues until the replication fork completes one full 360° round around the circular DNA.
The fork returns again to the DSO region.
At this stage the Rep protein introduces a second cleavage reaction.
This separates the newly synthesized strand from the original strand.
The Rep protein then joins the 5′ and 3′ ends of the displaced strand.
As a result a closed circular single stranded DNA molecule is released.
The original template strand remains paired with the newly synthesized complementary strand.
Thus a complete double stranded circular DNA molecule is formed.
Lagging Strand Synthesis (Conversion into Double-Stranded DNA)
The released single stranded circular DNA contains a specific sequence called Single-Strand Origin (SSO).
This region is recognized by host replication enzymes.
Host RNA polymerase or primase binds to the SSO and synthesizes a short RNA primer.
This primer provides a starting point for DNA synthesis.
DNA polymerase III extends the primer and synthesizes the complementary strand around the circular template.
The strand synthesis continues until a full circle is completed.
Finally DNA polymerase I removes the RNA primer and replaces it with DNA nucleotides.
DNA ligase seals the remaining nick and a mature double stranded circular DNA molecule is produced.
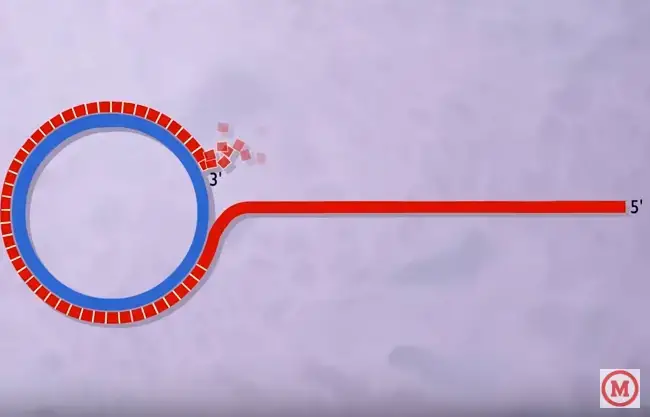
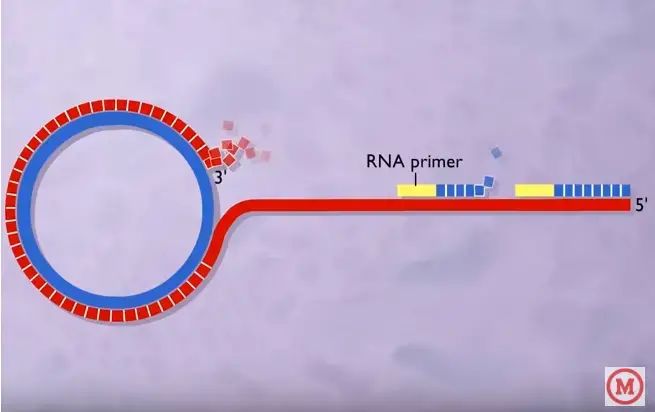
As a summary, a typical DNA rolling circle replication has five steps
- The dsDNA in circular form will be “nicked.”
- Using “unnicked” DNA as the leading strand (template), the 3′ end is lengthened; the 5′ end is displaced.
- A succession of Okazaki fragments are used to convert a lagging strand of DNA into a double-stranded molecule.
- Both “unnicked” and misplaced ssDNA replication.
- Displaced DNA circularises.
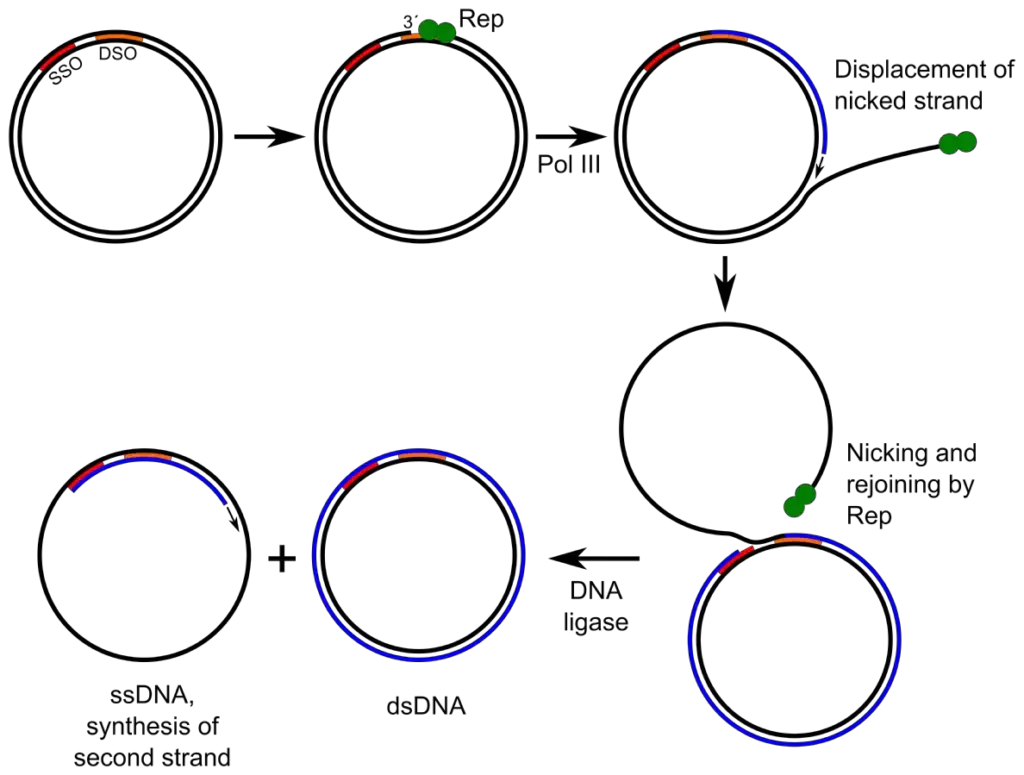
Replication of viral DNA using Rolling circle replication
- After entering into the host cell, the viral single stranded DNA genome is converted into double stranded DNA. This double stranded molecule is referred to as Replicative Form (RF) and it becomes supercoiled in structure.
- The formation of replicative form occurs by synthesis of complementary strand. Host enzymes synthesize the lagging strand using the viral single stranded DNA as template.
- A virus encoded initiator protein (Rep protein) then binds to a specific region on the double stranded DNA. This region is called Double-Strand Origin (DSO) and it often forms a stem-loop structure.
- The Rep protein introduces a nick in the positive (+) strand at a specific site within the origin. This cleavage occurs in the phosphodiester bond of the DNA backbone.
- After nicking, the Rep protein remains covalently attached to the 5′ phosphate end of the cut strand. A free 3′ hydroxyl (3′-OH) end is produced which acts as a primer for DNA synthesis.
- Host DNA polymerase binds to the free 3′-OH end and begins synthesis of new DNA. The intact negative (–) strand serves as the template for elongation.
- As DNA polymerase moves around the circular genome, continuous synthesis takes place. The original nicked positive strand is gradually displaced as a single stranded DNA.
- The elongation continues until the replication machinery completes one full round around the circular DNA. The replication fork returns to the origin region.
- The Rep protein makes a second cleavage reaction at the origin site. It then ligates the ends of the displaced strand forming a closed circular single stranded DNA molecule.
- After release, two forms of DNA are present. The original template remains as double stranded circular DNA and the newly displaced strand exists as single stranded circular viral DNA.
- The newly formed single stranded viral DNA may undergo encapsidation. It is packaged into viral capsids forming new infectious virions.
- In some cases the released single stranded DNA is again converted into double stranded replicative form. This allows further rounds of rolling circle replication to occur within the host cell.
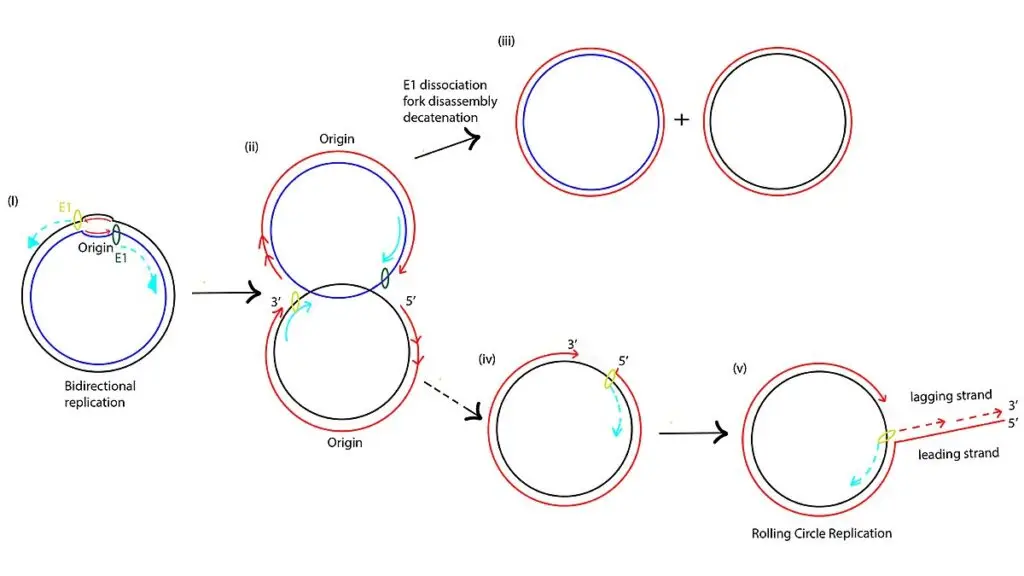
Replication of viral RNA using Rolling circle replication
Rolling circle replication in RNA viruses and viroids is the process by which circular RNA molecule is copied continuously to produce multiple RNA units.
This process follows two different pathways– Asymmetric pathway and Symmetric pathway.
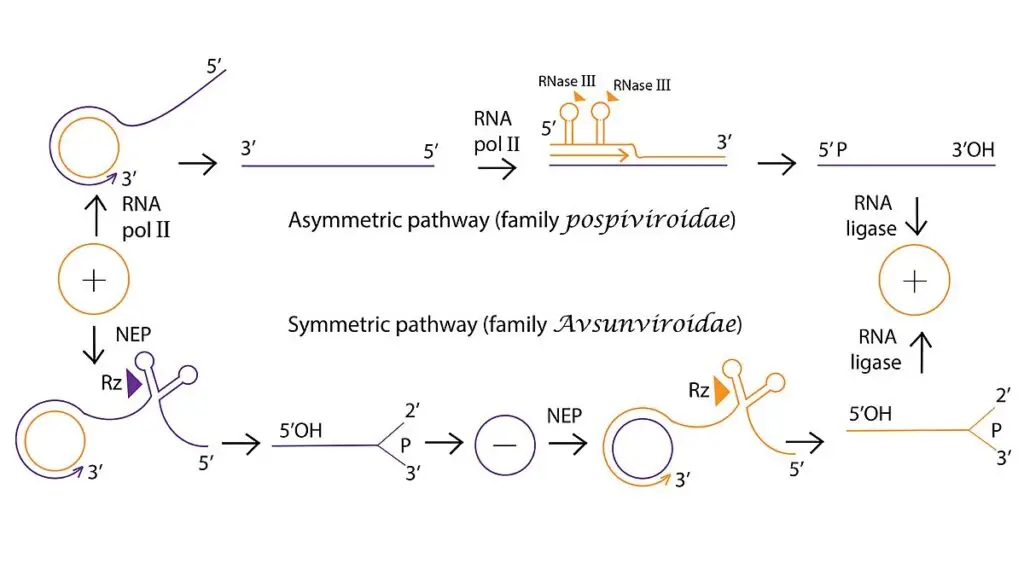
Asymmetric Replication Pathway
This pathway is seen in certain viroids such as members of Pospiviroidae family.
In this process only the plus strand is circularized at the end.
- Transcription of minus strand. The circular plus-strand RNA is used as template. Host RNA polymerase transcribes it continuously and forms a long oligomeric minus-strand RNA.
- Transcription of plus strand. The long minus-strand RNA now acts as template. It is again transcribed by host RNA polymerase forming a long oligomeric plus-strand RNA.
- Cleavage of oligomeric RNA. The long plus-strand RNA contains multiple genome copies in series. Host RNase enzyme cleaves this long RNA into linear monomeric plus-strand units.
- Ligation and circularization. The linear monomeric plus-strand RNAs are joined at their ends. Host RNA ligase catalyzes the ligation and mature circular plus-strand RNA molecules are formed.
Symmetric Replication Pathway
This pathway is observed in Avsunviroidae family.
In this process both plus and minus strands are circularized during replication.
- Transcription of minus strand. The circular plus-strand RNA is transcribed into a long linear oligomeric minus-strand RNA. Host RNA polymerase carries out this reaction.
- Cleavage of minus strand. The oligomeric minus-strand RNA is cleaved into linear monomers. This cleavage is mediated by a self-cleaving hammerhead ribozyme structure present within the RNA.
- Ligation of minus strand. The linear minus-strand monomers are ligated at their ends. Circular monomeric minus-strand RNAs are produced.
- Transcription of plus strand. These circular minus-strand RNAs now serve as templates. Long oligomeric plus-strand RNAs are synthesized.
- Cleavage of plus strand. The oligomeric plus-strands undergo self-cleavage by hammerhead ribozymes. Linear monomeric plus-strand units are formed.
- Ligation of plus strand. The linear plus-strand monomers are ligated. Mature circular plus-strand RNA molecules are produced completing the replication cycle.
What is Rolling circle amplification?
- Rolling Circle Amplification is a nucleic acid amplification technique. It is derived from rolling circle replication mechanism and the reaction is carried out at constant temperature (isothermal condition).
- It is used to amplify DNA or RNA rapidly without thermal cycling. The amplification takes place generally between 30°C to 37°C.
- The main components required for RCA are– circular DNA template, short primer (DNA or RNA), deoxynucleotide triphosphates (dNTPs), suitable buffer and a strand displacing DNA polymerase (commonly phi29 DNA polymerase).
- In this process the primer anneals to the circular template. DNA polymerase binds to the 3′-OH end of the primer and initiates DNA synthesis.
- The polymerase moves continuously around the circular DNA. As it moves forward, the previously synthesized strand is displaced and elongation continues.
- The product formed is a long single stranded DNA molecule. It contains multiple tandem repeats complementary to the circular template and this long repeated strand is referred to as concatemer.
- Since the reaction occurs at constant temperature, no thermocycler is required. It can be performed using simple heating block or water bath.
- High fidelity is observed in RCA. As amplification occurs directly from circular template, early replication errors are not exponentially amplified.
- The reaction is highly stable and resistant to inhibitors. It can function efficiently even in crude biological samples.
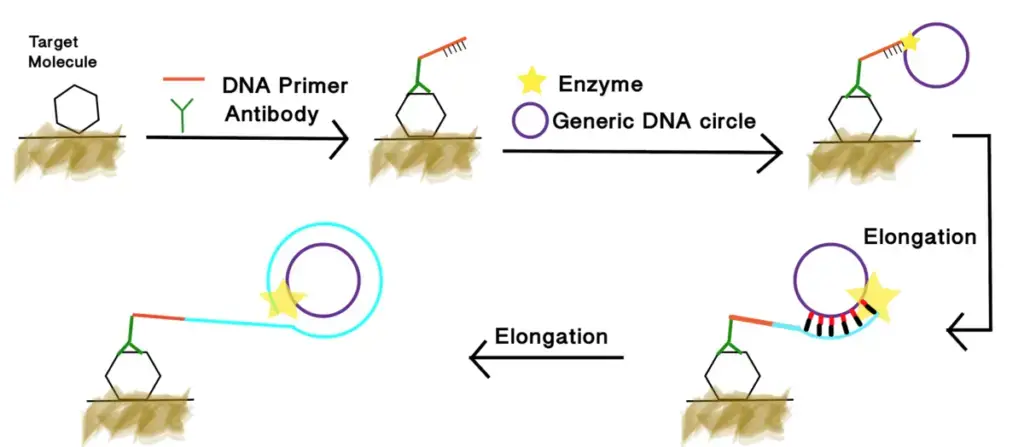
- RCA is modified into different formats such as– Linear RCA, Hyperbranched RCA, Padlock probe RCA and Immuno-RCA.
- It has wide applications in molecular diagnosis and research. It is used in detection of pathogens, next generation sequencing platforms and DNA nanotechnology studies.
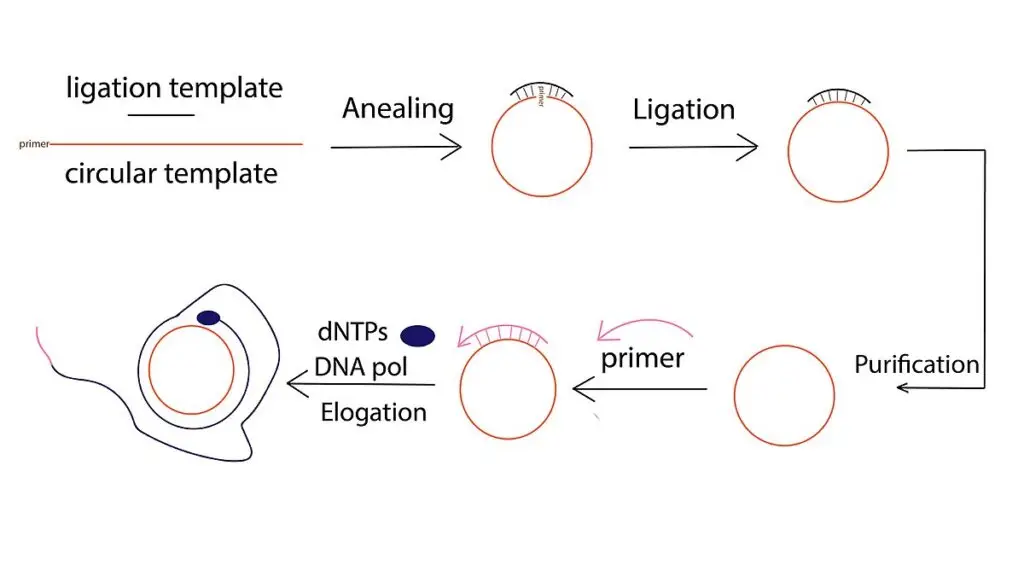
Components Required for Rolling circle amplification
Components Required for Rolling Circle Amplification (RCA)
- Circular DNA template
It is the closed circular DNA molecule used as template in RCA.
It may be a plasmid DNA or a circularized oligonucleotide and it does not have free ends. - Primer
It is a short DNA or RNA sequence required to start the synthesis.
The primer anneals to the complementary region of circular template and provides a free 3′-OH end. - DNA polymerase
It is the enzyme that synthesizes new DNA strand from the template.
A highly processive polymerase with strong strand displacement activity is required (phi29 DNA polymerase is commonly used, Bst and Vent (exo–) polymerases are also used). - Deoxynucleotide triphosphates (dNTPs)
These are the building blocks required for DNA synthesis.
dATP, dTTP, dCTP and dGTP are incorporated sequentially by the polymerase during elongation. - Reaction buffer
It is required to maintain suitable condition for enzyme activity.
The buffer maintains optimum pH and provides essential cofactors (mainly Mg²⁺ ions) for the polymerase to function.
Steps of DNA RCA (Rolling circle amplification) reaction
Step 1- Circular template preparation (ligation)
In DNA RCA, a circular DNA template is required for starting the reaction. It is commonly prepared by cyclizing a linear oligonucleotide (C-probe or padlock probe) into a closed circle. This circularization is done by enzymatic ligation using a ligase enzyme (e.g. T4 DNA ligase) with a splint DNA, or by template-free ligation using CircLigase.
Step 2- Primer annealing
A short DNA or RNA primer is added in the reaction mixture and it anneals to the complementary region of the circular template. The primer provides a free 3’-OH end for polymerase to start synthesis. In some reactions, multiple primers are used to anneal on same circular template to initiate multiple amplification events (Multiprimed RCA).
Step 3- Elongation and strand displacement
A strand-displacing DNA polymerase (phi29 DNA polymerase, Bst polymerase, Vent exo- polymerase) binds to the annealed primer and starts adding dNTPs to the 3’-OH end. The polymerase moves continuously on the circular template and synthesizes a new complementary strand. When one round is completed, the polymerase does not stop, it displaces the previously synthesized strand and continues synthesis again around the circle.
Step 4- Concatemer generation
Because the polymerase is moving again and again on the circular template in a continuous manner a long single stranded DNA product is formed. This product is referred to as concatemer. It contains many tandem repeats which are complementary to the original circular template and the repeats are formed in large number (tens to hundreds or more).
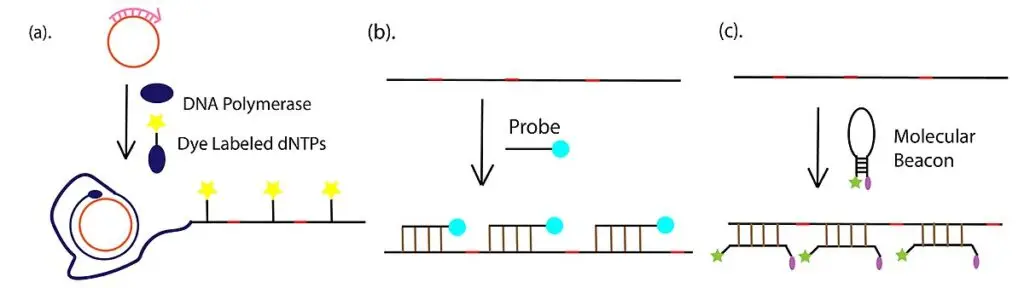
Step 5- Detection and visualization
The final RCA product is detected by different methods based on the experiment. Fluorescent detection is commonly used by incorporating labeled dNTPs or by hybridization of fluorophore-labeled probes/molecular beacons with the concatemer. The amplified product can also be analysed by gel electrophoresis for confirming formation of long DNA strands.
Importance of Rolling Circle Amplification (RCA)
- RCA is an isothermal amplification technique. It amplifies nucleic acids at constant temperature and it eliminates the requirement of thermocycler as in PCR.
- It is a simple and cost effective method. The reaction can be carried out using water bath or heating block and complex thermal cycling set up is not required.
- RCA is used for ultra-sensitive molecular diagnosis. It can detect very low amount of target DNA or RNA and it is used in diagnosis of infectious pathogens (SARS-CoV-2, HIV, Ebola, Tuberculosis, Malaria) and also cancer biomarkers.
- High fidelity amplification is achieved in RCA. By using highly processive enzyme like phi29 DNA polymerase very long DNA strand is synthesized (more than 70 kb) with low error rate.
- It avoids the mutant jackpot effect seen in PCR. Since copies are generated directly from circular template early errors are not amplified exponentially.
- RCA is important for cell-free DNA cloning. It amplifies circular DNA in vitro without using bacterial host and it is useful for genes which are toxic to host cells like E. coli.
- It has major importance in next generation sequencing. RCA is used for generating DNA nanoballs (DNBs) and it helps in producing more accurate sequencing data as clonal error propagation is reduced.
- RCA helps in single cell and spatial analysis. The amplified product remains tethered near target site and it allows localized signal amplification for detecting DNA, RNA and proteins within single cells.
- RCA is used in DNA nanotechnology and DNA origami. It is used to synthesize long single stranded DNA scaffolds which are folded into programmable 2D and 3D nanostructures.
- These DNA nanostructures have different applications. It is used in drug delivery studies, bio-computing and high density DNA data storage.
Importance of Rolling Circle Replication
- Rolling circle replication is an important mechanism for rapid multiplication of genetic material. It is mainly used when circular DNA genome is present and multiple copies are required in short time.
- It is the major replication mechanism used by many bacteriophages. Small single stranded DNA bacteriophages use this process for fast genome replication inside the host cell.
- It is also important in replication of several plant viruses and animal viruses. Viruses such as geminiviruses, nanoviruses and human papillomavirus uses rolling circle type replication for genome propagation.
- Rolling circle replication is linked with conjugational transfer in bacteria. In this process plasmid DNA is transferred from one bacterial cell to another and this is helping in horizontal gene transfer.
- It plays a major role in spread of antibiotic resistance genes. Many resistance genes are carried on plasmids and these plasmids can replicate and transfer by rolling circle mechanism.
- Rolling circle replication is involved in genome plasticity. Certain mobile genetic elements and transposons (e.g. IS91 type, helitrons) show rolling circle transposition and this modifies the host genome.
- The same principle is used in laboratory as Rolling circle amplification (RCA). It is an isothermal amplification method where circular template is amplified without thermal cycling.
- RCA is important for ultra-sensitive diagnosis. It is used to detect low amount of DNA or RNA targets and it is useful in diagnosis of infectious diseases and cancer biomarkers.
- It is used in next generation sequencing methods. DNA nanoballs are produced by RCA and since copies are made directly from template, clonal error propagation is reduced.
- RCA is also important in cell-free DNA cloning and DNA nanotechnology. Circular DNA molecules can be amplified in vitro and long single stranded DNA scaffolds can be produced for DNA origami studies.
References
- Bestard-Cuche, N., Munro, D. A. D., Beniazza, M., Priller, J., Williams, A., & Corsinotti, A. (2024). Illumina SBS sequencing and DNBSEQ perform similarly for single-cell transcriptomics. Genes, 15(11), 1436. https://doi.org/10.3390/genes15111436
- BYJU’S. (n.d.). Theta mode of replication. Retrieved from https://byjus.com/neet/theta-mode-of-replication/
- Cheung, A. K. (2006). Rolling-circle replication of an animal circovirus genome in a theta-replicating bacterial plasmid in Escherichia coli. Journal of Virology, 80(17), 8686–8694. https://doi.org/10.1128/JVI.00655-06
- Complete Genomics. (n.d.). Performance and compatibility evaluation of Celemics NGS panels with DNBSEQ-G400 sequencing platform. Complete Genomics.
- EmaSushan. (n.d.). Various model of DNA replication. SlideShare. Retrieved from https://www.slideshare.net
- Espinosa, M., del Solar, G., Rojo, F., & Alonso, J. C. (1995). Plasmid rolling circle replication and its control. FEMS Microbiology Letters, 130(2-3), 111–120. https://doi.org/10.1016/0378-1097(95)00194-8
- Faurez, F., Dory, D., Grasland, B., & Jestin, A. (2009). Replication of porcine circoviruses. Virology Journal, 6, 60. https://doi.org/10.1186/1743-422X-6-60
- Garafutdinov, R. R., Sakhabutdinova, A. R., Gilvanov, A. R., & Chemeris, A. V. (2021). Rolling circle amplification as a universal method for the analysis of a wide range of biological targets. Russian Journal of Bioorganic Chemistry, 47(6), 1172–1189. https://doi.org/10.1134/S1068162021060078
- Garcillán-Barcia, M. P., Pluta, R., Lorenzo-Díaz, F., Bravo, A., & Espinosa, M. (2021). The facts and family secrets of plasmids that replicate via the rolling-circle mechanism. Microbiology and Molecular Biology Reviews, 86(1), e00222-20. https://doi.org/10.1128/MMBR.00222-20
- Garg, M. (n.d.). A generalised model of DNA replication (with variations) | Microbiology. Biology Discussion. Retrieved from https://www.biologydiscussion.com
- He, Z., Shi, K., Li, J., & Chao, J. (2023). Self-assembly of DNA origami for nanofabrication, biosensing, drug delivery, and computational storage. iScience, 26(5), 106638. https://doi.org/10.1016/j.isci.2023.106638
- Hu, T., Chen, J., Lin, X., He, W., Liang, H., Wang, M., Li, W., Wu, Z., Han, M., Jin, X., Kristiansen, K., Xiao, L., & Zou, Y. (2024). Comparison of the DNBSEQ platform and Illumina HiSeq 2000 for bacterial genome assembly. Scientific Reports, 14, 1292. https://doi.org/10.1038/s41598-024-51725-0
- Hutchison, C. A., III, Smith, H. O., Pfannkoch, C., & Venter, J. C. (2005). Cell-free cloning using φ29 DNA polymerase. Proceedings of the National Academy of Sciences of the United States of America, 102(48), 17332–17336. https://doi.org/10.1073/pnas.0508809102
- McInerney, P., Adams, P., & Hadi, M. Z. (2014). Error rate comparison during polymerase chain reaction by DNA polymerase. Molecular Biology International, 2014, 287430. https://doi.org/10.1155/2014/287430
- NPTEL. (n.d.). Module 7: Microbial molecular biology and genetics, lecture 2: DNA replication.
- Ouyang, X., Li, J., Liu, H., Zhao, B., Yan, J., Ma, Y., Xiao, S., Song, S., Huang, Q., Chao, J., & Fan, C. (2013). Rolling circle amplification-based DNA origami nanostructures for intracellular delivery of immunostimulatory drugs. Small, 9(18), 3082–3087. https://doi.org/10.1002/smll.201300458
- Park, J. (2025). Rolling circle amplification as a molecular tool for spatially resolved signal amplification in single molecule counting assay. Biosensors, 15(9), 628. https://doi.org/10.3390/bios15090628
- Póliska, S., Fareh, C., Lengyel, A., Göczi, L., Tőzsér, J., & Szatmari, I. (2024). Comparative transcriptomic analysis of Illumina and MGI next-generation sequencing platforms using RUNX3- and ZBTB46-instructed embryonic stem cells. Frontiers in Genetics, 14, 1275383. https://doi.org/10.3389/fgene.2023.1275383
- Rajendran, A., Endo, M., & Sugiyama, H. (2012). DNA origami: Synthesis and self-assembly. Current Protocols in Nucleic Acid Chemistry, 48, 12.9.1–12.9.18.
- Reem, A., Almezgagi, M., Al-Shaebi, F., Al-Shehari, W. A., & Kumal, J. P. P. (2020). Application of rolling circle amplification (RCA) to detect the pathogens of infectious diseases. Infectious Diseases Diagnosis & Treatment, 4, 158. https://doi.org/10.29011/2577-1515.100158
- The biomechanics and biotechnological evolution of rolling circle replication. (n.d.).
- Thermo Fisher Scientific. (n.d.). Rolling circle amplification (RCA) and whole genome amplification (WGA). Retrieved from https://www.thermofisher.com
- Wawrzyniak, P., Płucienniczak, G., & Bartosik, D. (2017). The different faces of rolling-circle replication and its multifunctional initiator proteins. Frontiers in Microbiology, 8, 2353. https://doi.org/10.3389/fmicb.2017.02353
- Wikipedia contributors. (2025, September 30). Rolling circle replication. In Wikipedia, The Free Encyclopedia. Retrieved from https://en.wikipedia.org/w/index.php?title=Rolling_circle_replication
- Zhang, Z., Zhang, H., Wang, F., Zhang, G., Zhou, T., Wang, X., Liu, S., & Liu, T. (2018). DNA block macromolecules based on rolling circle amplification act as scaffolds to build large-scale origami nanostructures. Macromolecular Rapid Communications, 39(15), e1800263. https://doi.org/10.1002/marc.201800263