Nuclear staining is the process in which the nucleus of a cell is given artificial colour so that it becomes visible under microscope. It is the main method used in histology and cytology because the nucleus is naturally transparent and it needs specific dyes for proper contrast. The nucleus contains DNA and chromatin materials which are acidic in nature, so these are stained by basic dyes. It is the basic principle followed during most nuclear staining procedures, and it helps in observing nuclear structure, shape and organisation.
There are different methods of nuclear staining which are based on chemical reactions or on fluorescent dyes. These methods allow the identification of DNA, RNA and chromatin in fixed or living cells.
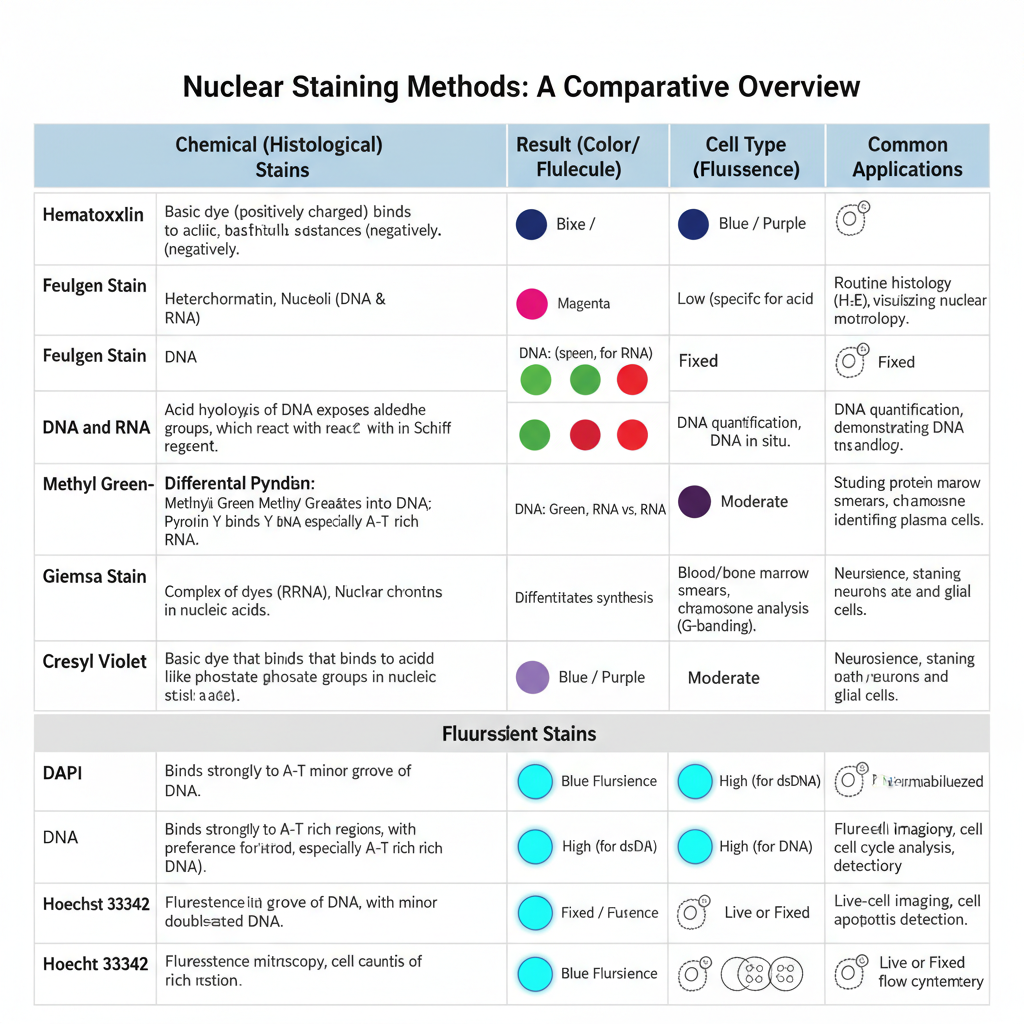
Chemical (Histological) Nuclear Staining Methods
In this method, staining occurs due to chemical interaction between dye and the nucleic acids. Feulgen stain is one important method in which DNA is hydrolysed by acid and forms aldehyde groups which bind with Schiff reagent to produce magenta colour. It is specific only for DNA. Methyl Green–Pyronin stain also distinguishes between DNA and RNA.
Methyl green binds with DNA and gives green colour while Pyronin Y binds with RNA and gives red colour. In tissues, Hematoxylin and Eosin stain is widely used where hematoxylin stains the nucleus blue, and it is the main stain to study nuclear details.
Giemsa stain is a polychromatic stain where nucleus is stained dark blue and is used for blood and bone marrow cells. Cresyl violet stain is used for Nissl substance and for nuclear chromatin which become blue to purple in neuronal cells.
Fluorescent Nuclear Staining Methods
In this method fluorochromes are used which bind with nucleic acids and produce fluorescence when exposed to UV light. DAPI (4′,6-diamidino-2-phenylindole) is a common dye which binds mainly with double-stranded DNA and gives blue fluorescence. It is used mostly in fixed cells because the dye needs permeabilisation of the membrane.
Hoechst 33342 also binds with DNA but it can enter live cells easily, so it is used for staining nuclei of living cells. These dyes are used in fluorescence microscopy and flow cytometry. The working concentration is low and staining occurs within a short time. Proper fixation like formalin or alcohol fixation is important to preserve morphology and to reduce background. Permeabilisation is also used in fixed tissues for entry of fluorescent dyes into the nucleus.
In all these methods, the aim is to make the nucleus clearly visible and to study its organisation for diagnostic and research purposes.
Nuclear Staining by DAPI
Principle
The principle of nuclear staining by DAPI is based on its strong affinity for the DNA present inside the nucleus. It is a blue-fluorescent dye that binds mainly with double-stranded DNA, and this binding takes place in the minor groove of the helix where AT-rich regions are present. It is the specific interaction that increases the fluorescence intensity of the dye, and when DAPI is attached to DNA the fluorescence becomes highly enhanced. This change occurs because water molecules are displaced from the dye and the groove region when binding takes place.
When DAPI binds with the nuclear DNA, it shows an excitation around 358 nm and gives a blue emission around 461 nm under UV light. The nucleus therefore becomes clearly visible as a bright blue structure. It may also bind with RNA but the fluorescence is weak and the emission shifts towards a longer wavelength which reduces the chance of cytoplasmic staining. Because of this property, it is mostly used as a nuclear counterstain in fluorescence microscopy.
Since DAPI cannot easily enter living cells, it is used mostly in fixed and permeabilised cells. Fixation opens the nuclear membrane and allows the dye to reach the DNA target. A low working concentration such as 300 nM is used to stain the nuclear material, and the result is a clear and high-contrast appearance of the nucleus and chromosomes.
Procedure
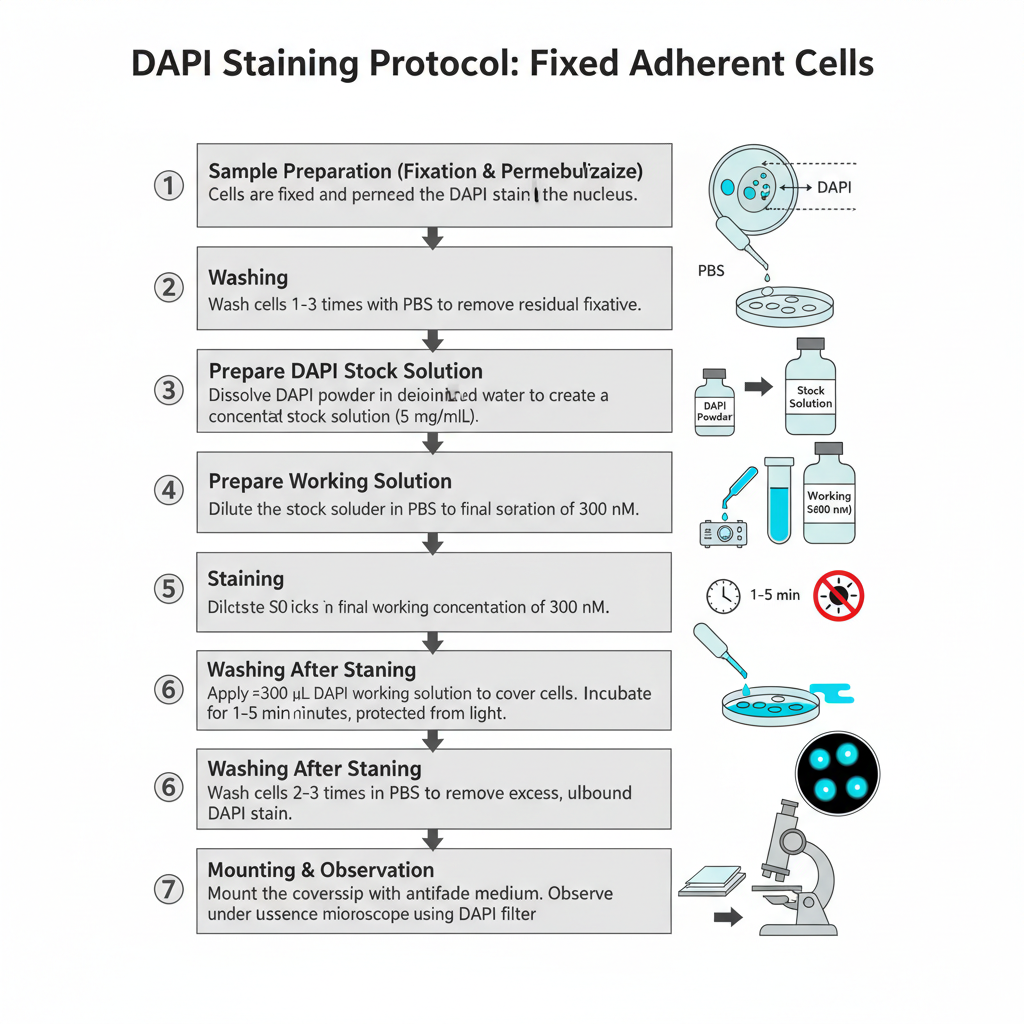
I. Procedure for Fixed Adherent Cells (Fluorescence Microscopy)
1. Sample Preparation (Fixation and Permeabilization)
Cells are fixed and permeabilized by using a suitable method. It is done earlier so that DAPI can enter inside the nucleus.
2. Washing
Cells is washed 1–3 times with PBS. This is done for removing residual fixative.
3. Preparation of DAPI Stock Solution
A concentrated stock (5 mg/mL) is made by dissolving the dye in deionized water or DMF. Sometimes complete dissolution is difficult so sonication is needed.
4. Preparation of Working Solution
The stock is diluted in PBS to make a final concentration of 300 nM. This is the commonly used concentration for staining fixed cells.
5. Staining
About 300 μL of DAPI working solution is applied over the coverslip so the cells are completely covered. It is incubated for 1–5 minutes and the preparation is kept protected from light.
6. Washing After Staining
The stain solution is removed. Cells is washed 2–3 times in PBS for removing excess unbound dye.
7. Mounting and Observation
The extra buffer is drained and the coverslip is mounted with an antifade mounting medium (for example SlowFade or ProLong type). It is then observed under a fluorescence microscope with DAPI filter.
II. Procedure for Cells in Suspension (Flow Cytometry)
1. Sample Preparation (Fixation)
A suspension containing approximately 2×10⁵ to 1×10⁶ cells is taken. Cells is pelleted by centrifugation. The pellet is gently resuspended and slowly transferred into cold absolute ethanol (−20°C) with vortexing. It is kept for 5–15 minutes for fixation.
2. Rehydration and Washing
The ethanol is removed after centrifugation. PBS (5 mL) is added and the cells is allowed to rehydrate for around 15 minutes.
3. Preparation of DAPI Working Solution
A working concentration of 3 μM DAPI is prepared in staining buffer. The staining buffer contains Tris (100 mM, pH 7.4), NaCl (150 mM), CaCl₂ (1 mM), MgCl₂ (0.5 mM), and 0.1% NP-40.
4. Staining of Cells
Cells is pelleted again and resuspended in 1 mL of the DAPI staining buffer. It is incubated for 15 minutes at room temperature.
5. Analysis
The stained cells is analysed by flow cytometry directly in the presence of dye.
Uses
- It is used as a nuclear counterstain in multicolor fluorescence.
- It is used for DNA content quantification in flow cytometry.
- It is used as a nucleic acid and chromosome counterstain in fluorescence microscopy.
- It is used in chromosome FISH staining.
- It is used for fungal nuclear staining in fixed and permeabilized cells.
Advantages
- It is highly efficient for staining fixed and permeabilized cells.
- It is used as a nuclear counterstain giving vivid contrast in multicolor fluorescence.
- It stains nuclei specifically with very little cytoplasmic labeling.
- It gives strong fluorescence enhancement when it binds with dsDNA.
- It is used in multicolor flow cytometry for labeling cells and estimating DNA content.
Limitations
- It is a known mutagen and must be handled carefully.
- It has emission close to natural autofluorescence which can reduce image clarity.
- It does not penetrate rigid barriers well and fungi often require aggressive cell wall digestion.
Nuclear Staining by Hoechst
Principle
The principle of nuclear staining by Hoechst 33342 is based on its ability to enter living cells and bind specifically with the DNA present in the nucleus. It is a blue-fluorescent dye which shows a strong affinity for double-stranded DNA, and the binding mainly occurs in the minor groove where AT-rich regions are present. It is the cell-permeant nature of Hoechst dye that allows the molecules to cross the plasma membrane of living cells, so the nucleus can be visualised without fixation. Because the dye binds directly with DNA, it has some toxic effects and therefore it is used in controlled concentration for live-cell studies.
When Hoechst binds with DNA, it shows excitation around 350 nm and gives blue emission around 461 nm under UV light. The nucleus becomes clearly visible with this fluorescence, and it can be used together with other fluorophores because the blue emission does not interfere with green or red channels. For live-cell staining, a higher concentration and longer incubation time is required, as the dye must pass through intact cellular barriers. A dilution such as 1:2000 is used, and the incubation is done for 5–15 minutes so that enough molecules can reach the nuclear DNA.
This dye is mostly used when nuclear events are to be observed in living cells because its permeability and fluorescence properties allow clear staining of DNA with minimal background.
Procedure
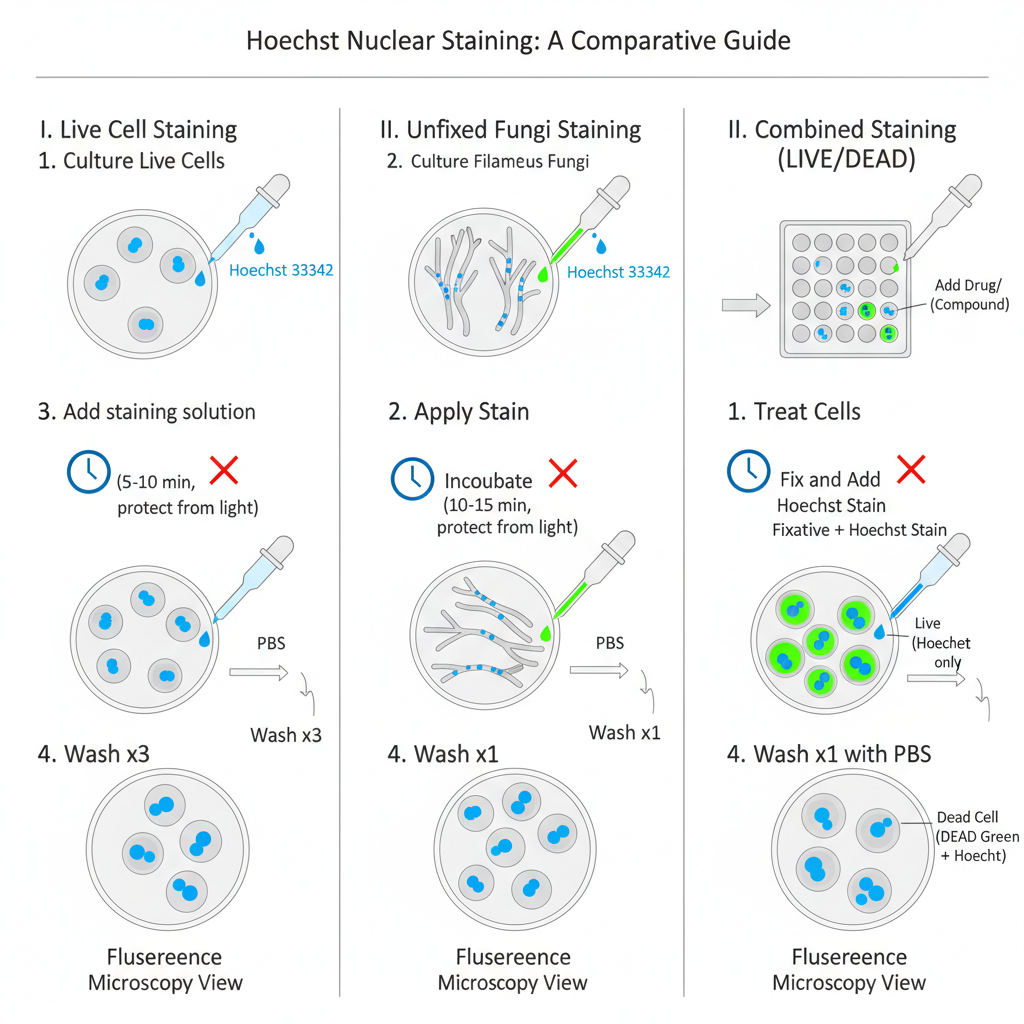
I. Hoechst 33342 Staining Procedure for Live Cells
1. Preparation of Hoechst Stock Solution
The dye is dissolved by taking 100 mg into 10 mL deionized water to make 10 mg/mL solution. Sometimes it has poor solubility so sonication is required for complete dissolution. The stock is stored at 2–6°C for several months or at −20°C for long storage.
2. Preparation of Culture
Cells is cultured in suitable medium and vessel for microscopy.
3. Preparation of Staining Solution
The stock is diluted 1:2000 in PBS to prepare the working solution.
4. Removal of Medium
The culture medium is removed from the cells carefully.
5. Staining
Staining solution is added so that the cells are fully covered. It is incubated for 5–10 minutes keeping the preparation protected from light.
6. Direct Imaging (Optional)
Cells can be imaged directly in staining solution if required.
7. Washing
The staining solution is removed and the cells is washed 3 times with PBS.
8. Imaging
The stained cells is imaged using a fluorescence microscope with DAPI filter set.
II. Hoechst 33342 Staining Procedure for Unfixed Filamentous Fungi
1. Culture Preparation
Filamentous fungi such as A. fumigatus is grown in appropriate culture medium until germination.
2. Dye Application
The supernatant is removed. Hoechst 33342 is added at 1:2000 dilution in PBS directly to the unfixed material.
3. Incubation
The sample is incubated for 10–15 minutes protected from light so that dye can cross the cell wall barrier.
4. Washing
The sample is washed once with PBS and resuspended in fresh PBS.
5. Imaging
Nuclei is observed and counted under a fluorescence microscope with UV filter. The dye has excitation at 350 nm and emission at 461 nm.
III. Hoechst Staining in Combination with Other Stains
Hoechst 33342 is used along with other dyes in multicolor staining because its blue emission do not interfere with green or red fluorophores.
Example Combined Staining Procedure (LIVE/DEAD Green Kit)
1. Culturing
Cells is cultured in a 96-well plate using proper medium.
2. Drug Incubation
The drug or compound is added and incubated for desired time.
3. DEAD Green Staining
About 50 μL of DEAD Green staining solution is added and the plate is incubated at 37°C for 30 minutes.
4. Fixation
The medium is removed. Fixation solution containing 16% paraformaldehyde and Hoechst 33342 (Component B) is added.
5. Fixation Incubation
The plate is kept at room temperature for around 15 minutes.
6. Washing and Analysis
Fixative is removed and the wells is washed once with PBS. The cells is analysed under high-content imaging system.
Uses
- It is used for live cell nuclear visualization as it is cell-permeant.
- It is used to distinguish condensed nuclei during apoptosis.
- It is used in cell cycle studies with BrdU.
- It is used in fungal research for studying nuclear dynamics.
- It is used for total cell demarcation in high-content screening assays.
- It is used in multi-color fluorescence imaging because spectral crosstalk is low.
Advantages
- It is cell-permeant and suitable for nuclear staining in live cells.
- It is suitable for multi-color fluorescence imaging with reduced spectral crosstalk.
- It is used to distinguish condensed nuclei in apoptotic cells.
- It can be used with viability stains for marking total cell populations.
Limitations
- It is a known mutagen.
- It is toxic because it binds directly to DNA.
- Its fluorescence signal is quenched by BrdU.
- Excess unbound dye can shift emission to green and produce haze.
- Live fungal cells need higher concentrations and longer incubation for penetration.
Nuclear Staining by Hematoxylin and Eosin (H&E) Stain
Principle
The principle of nuclear staining by Hematoxylin and Eosin stain is based on the acid–base interaction between the dye molecules and the cellular components. The nucleus contains DNA and RNA, and these nucleic acids are acidic in nature. Because of this property the nucleus behaves as a basophilic element, so it readily binds with basic dyes. Hematoxylin itself is not an active dye, but after oxidation it forms hematein which combines with a metallic mordant such as aluminum to form a dye–mordant complex. This complex carries a positive charge and binds strongly with the negatively charged chromatin materials present in the nucleus. It is this binding that stains the nucleus blue.
Eosin is used as the counterstain. It is an acidic dye with a negatively charged chromophore, and it binds with basic or eosinophilic components such as the cytoplasm, muscle fibres and collagen. These structures appear pink after staining. This combination of a blue nucleus and a pink cytoplasm creates clear contrast and allows the tissue organisation to be easily observed under the microscope.
The staining can be done either progressively where the tissue is stained to the required intensity, or regressively where the tissue is first overstained and then differentiated by acid alcohol to remove excess stain from non-nuclear parts. The principle therefore depends mainly on the charge-based attraction between the dye complexes and the tissue components, and it is the reason why H&E stain is widely used for general histological examination.
Procedure
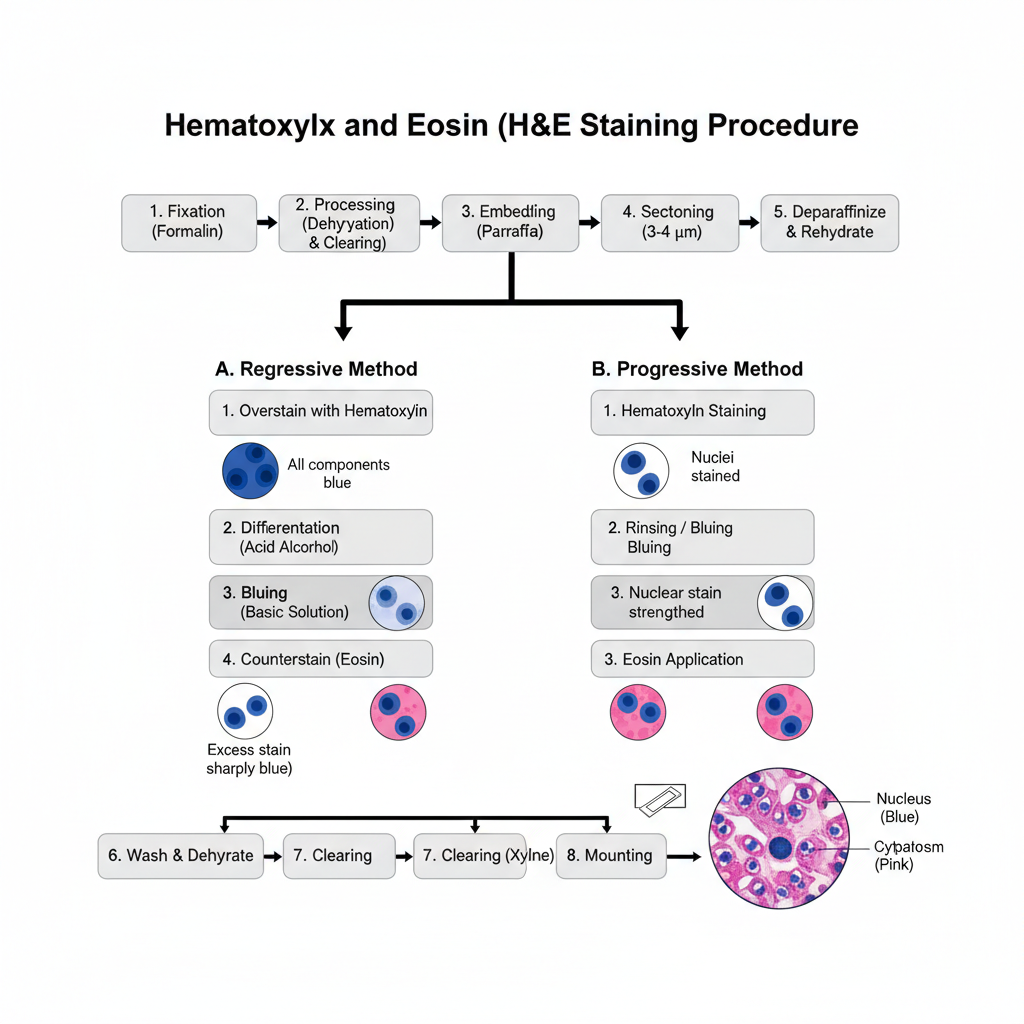
I. Specimen Preparation
Fixation is done before staining. Neutral buffered formalin is commonly used because it preserve proteins and nuclear structure. After fixation the tissue is processed.
Dehydration is done through graded alcohols. Clearing is done with xylene. Then infiltration and embedding in paraffin is carried out. The paraffin block is sectioned into 3–4 μm thick sections. These are deparaffinized in xylene and rehydrated to water.
II. Staining Methods
Two methods are used: regressive staining and progressive staining.
A. Regressive Method
1. Overstaining with hematoxylin
In this step the section is fully stained with hematoxylin. All components appear blue.
2. Differentiation
The section is placed in acid alcohol. Excess stain is removed from non-nuclear areas.
3. Bluing
The slide is placed in a basic solution. The nuclear colour becomes sharply blue.
4. Counterstaining
Eosin is applied. Cytoplasm and connective tissue appear pink.
B. Progressive Method
1. Hematoxylin staining
The section is stained until nuclei reach the required colour. Background remain almost clear.
2. Rinsing or bluing
The slide is rinsed in tap water or bluing reagent to strengthen the nuclear staining.
3. Eosin application
Eosin is applied as the final counterstain. It stains the cytoplasm pink.
III. Finishing
After staining the slide is washed. Dehydration is done again through graded alcohols. Clearing is done in xylene. The slide is mounted with a coverslip using mounting medium.
The nucleus appear blue. The cytoplasm appear pink. Other structures show different shades according to their affinity.
Uses
- It is used as the routine stain for microscopic diagnosis.
- It is used to show tissue structure and cellular architecture.
- It is used to diagnose disease and cancer and identify tissue type.
- It is used to observe intranuclear detail.
Advantages
- It is the routine stain for disease diagnosis and identification of cancer or tissue type.
- It shows tissue structure and cellular architecture clearly.
- It gives intranuclear detail.
- It provides differential staining of cytoplasm and connective tissue fibers in one application.
Limitations
- Poor fixation leads to weak staining and loss of nucleic acid preservation.
- Aluminum hematoxylins are removed when exposed to acidic solutions.
- Staining intensity changes with age of hematoxylin and solution reuse.
- Frozen sections can develop artifacts from excess saline.
- Old hematoxylin or chlorinated water may prevent chromatin staining.
Nuclear Staining by Feulgen Stain (DNA Specific)
Principle
The principle of nuclear staining by Feulgen stain is based on a specific chemical reaction that acts only on DNA. It is the acid hydrolysis of the deoxyribose sugar that produces aldehyde groups which can then bind with Schiff reagent. This reaction does not occur in RNA because the ribose sugar does not form the required aldehyde under the same hydrolysis condition. The method therefore becomes DNA specific, and it is used to visualise the nuclear DNA in fixed tissues.
During the procedure, the tissue is first treated with hydrochloric acid. It is the step that converts the deoxyribose of DNA into aldehyde groups, and the duration of hydrolysis is important because both under- and over-hydrolysis can weaken the reaction. When the aldehyde groups are formed, they react with Schiff reagent which is originally colourless. After binding takes place, the section is washed in running water, and the dye develops a pink to red colour. It is this reaction that shows the DNA as a magenta coloured structure inside the nucleus.
Usually a light green counterstain is applied to provide contrast, and the nucleus becomes clearly visible. The Feulgen stain therefore works on the principle of acid-induced aldehyde formation followed by Schiff reagent binding, making it one of the specific methods for DNA staining.
Procedure
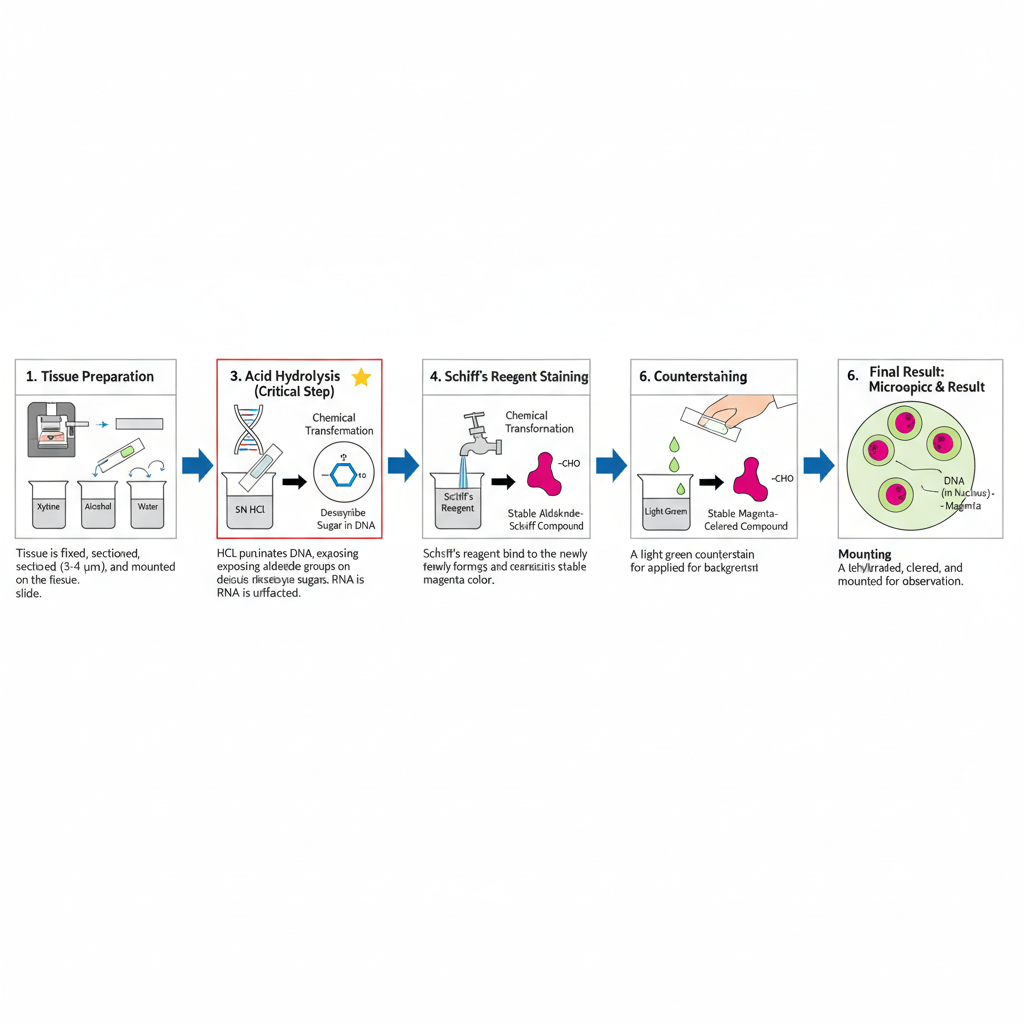
I. Specimen Preparation
1. Tissue preparation
The tissue specimen is fixed, processed and sectioned. It is generally cut into 3–4 μm thick sections. These are placed on clean slides.
2. Deparaffinization and rehydration
If the tissue is paraffin embedded, the paraffin is removed. It is passed through xylene and graded alcohol series. Finally the sections is brought back to water.
II. Feulgen Reaction Steps
1. Acid hydrolysis
In this step, the section is treated with hydrochloric acid. The hydrolysis converts the deoxyribose sugar into aldehyde groups. This is the most critical step because hydrolysis time control the reaction. Increasing the time increases the colour intensity up to an optimum point after which the reaction is weakened.
Different fixatives require different hydrolysis time. It is around 8 minutes for formalin fixed tissues. It is around 5 minutes for Zenker’s or Zenker–formol fixed tissues. Bouin’s fixative is unsuitable. The ribose sugar of RNA is not affected.
2. Schiff’s reagent staining
The aldehyde groups now react with Schiff’s reagent. Schiff’s reagent is prepared from basic fuchsin, HCl and sodium metabisulfite. Basic fuchsin combines with aldehydes to form a colourless compound at first.
3. Colour development
The slides are washed under running water. During washing, the colourless compound changes into a pink–red colour. This is the visible magenta colour of DNA.
III. Post-Staining Steps
1. Counterstaining
A light green counterstain is applied. It is used for background contrast.
2. Mounting
The sections are dehydrated, cleared and mounted with a coverslip. The slide is now ready for microscopic observation.
Uses
- It is used for selective DNA visualization.
- It is used for quantitative DNA analysis.
Advantages
- It is used for selective DNA demonstration.
- It gives DNA specificity because the acid hydrolysis modifies only deoxyribose.
- A light green counterstain can be used for better contrast.
Limitations
- The hydrolysis step is critical.
- Over-hydrolysis weakens and may ruin the staining.
- Some fixatives like Bouin’s are unsuitable.
Nuclear Staining by Methyl Green-Pyronin Y Stain (DNA and RNA Differentiation)
Principle
The principle of nuclear staining by Methyl Green–Pyronin Y stain depends on the differential binding of two basic dyes with the nucleic acids present inside the cell. Both dyes carry a positively charged chromophore which binds with the negatively charged groups of nucleic acids. Methyl green shows a strong affinity for highly polymerised DNA, so the DNA becomes green or blue-green in colour. Pyronin Y binds mainly with RNA, and because RNA is less polymerised compared to DNA, it stains red. This difference in polymerisation is considered the main reason for the selective staining of these two nucleic acids.
In this technique, the DNA–methyl green complex gives a clear nuclear colour, while the cytoplasmic RNA and nucleolus take up the pyronin stain. Fixation plays an important role, and Carnoy’s fixative usually gives the best staining because it preserves the nucleic acids in a condition suitable for dye binding. Formalin-fixed tissues can also be used with modification of the procedure.
After staining, the slides cannot be treated with water because water can remove the dyes and disturb the differentiation. Therefore, dehydration is done with acetone or tertiary butanol, and the slides are mounted in a synthetic resin. The stain thus works on the principle of selective binding of methyl green to DNA and pyronin Y to RNA, allowing clear differentiation between the two nucleic acids inside the cell.
Procedure
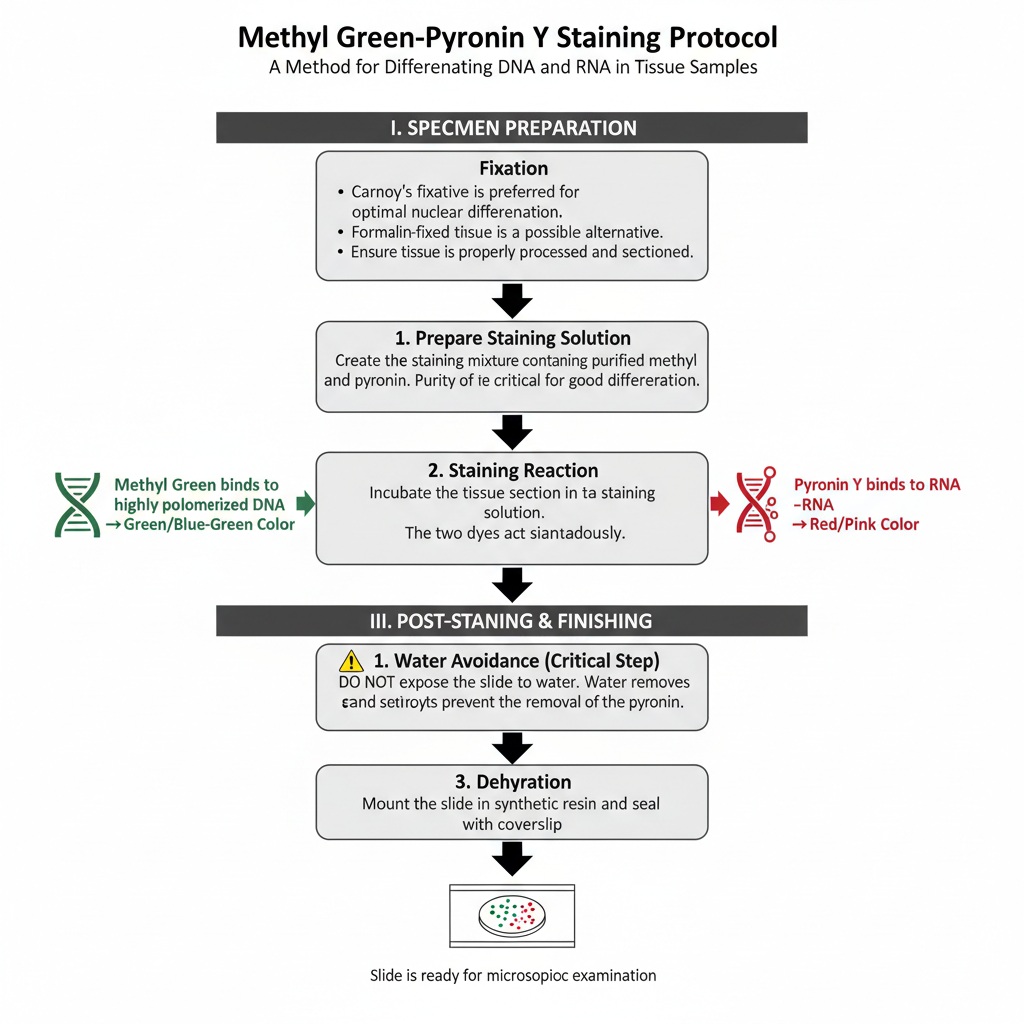
I. Specimen Preparation (Fixation)
Carnoy’s fixative is preferred for this staining. It gives the best nuclear differentiation. Formalin fixed tissues can also be used although the result is not always identical. The tissue must be processed and sectioned properly before staining.
II. Staining Procedure
1. Preparation of staining solution
In this step, the section is incubated in a staining mixture containing purified methyl green and pyronin Y. The purity of methyl green is important because impure dye causes poor differentiation.
2. Staining reaction
Methyl green binds with DNA forming a green or blue–green colour. Pyronin Y binds with RNA giving a red colour. The reaction is influenced by the degree of polymerization of nucleic acids. The two dyes act simultaneously.
III. Post-Staining and Finishing
1. Water avoidance
After staining the slide must not be exposed to water. Exposure to water remove the dye and destroy the differentiation.
2. Dehydration
Dehydration is carried out with acetone or tertiary butanol. These solvents prevent the removal of pyronin.
3. Mounting
The slide is mounted in a synthetic resin. It is then sealed with a coverslip and made ready for microscopic study.
Uses
- It is used to differentiate DNA and RNA.
- It is used to visualize nucleic acids together with the Feulgen stain.
Advantages
- It is used to demonstrate RNA specifically.
- It can differentiate DNA (green or blue-green) and RNA (red) in the same section.
- Dye concentration and pH can be adjusted for various tissues.
Limitations
- Slides must not contact water after staining.
- Optimal staining often requires Carnoy’s fixative.
- pH, dye purity, and concentration strongly affect staining success.
Nuclear Staining by Cresyl Violet Stain
Principle
The principle of nuclear staining by Cresyl violet stain depends on the affinity of this basic aniline dye for the acidic components of the cell. Cresyl violet carries a positive charge, so it binds strongly with basophilic structures. The nucleus contains chromatin materials which are acidic in nature because of the DNA content, and these components take up the dye readily. As a result, the nuclear chromatin becomes blue to purple after staining.
Besides the nucleus, Cresyl violet also stains the Nissl substance present in neurons. Nissl substance contains RNA and proteins associated with the rough endoplasmic reticulum, and due to its acidic nature it becomes basophilic. The stain highlights these structures clearly, and therefore it is used to study neuronal organisation. Loss of Nissl substance, known as chromatolysis, can also be detected using this stain and indicates neuronal damage.
After staining, differentiation is done with alcohol to remove the excess colour from the background tissue. Only the nuclear chromatin and Nissl substance retain the blue to purple shade. Formalin fixation is generally preferred, and the choice of basic dye can be adjusted depending on the fixation method. The staining works mainly on the principle of charge-based attraction between the basic dye and the acidic components of the cell.
Procedure
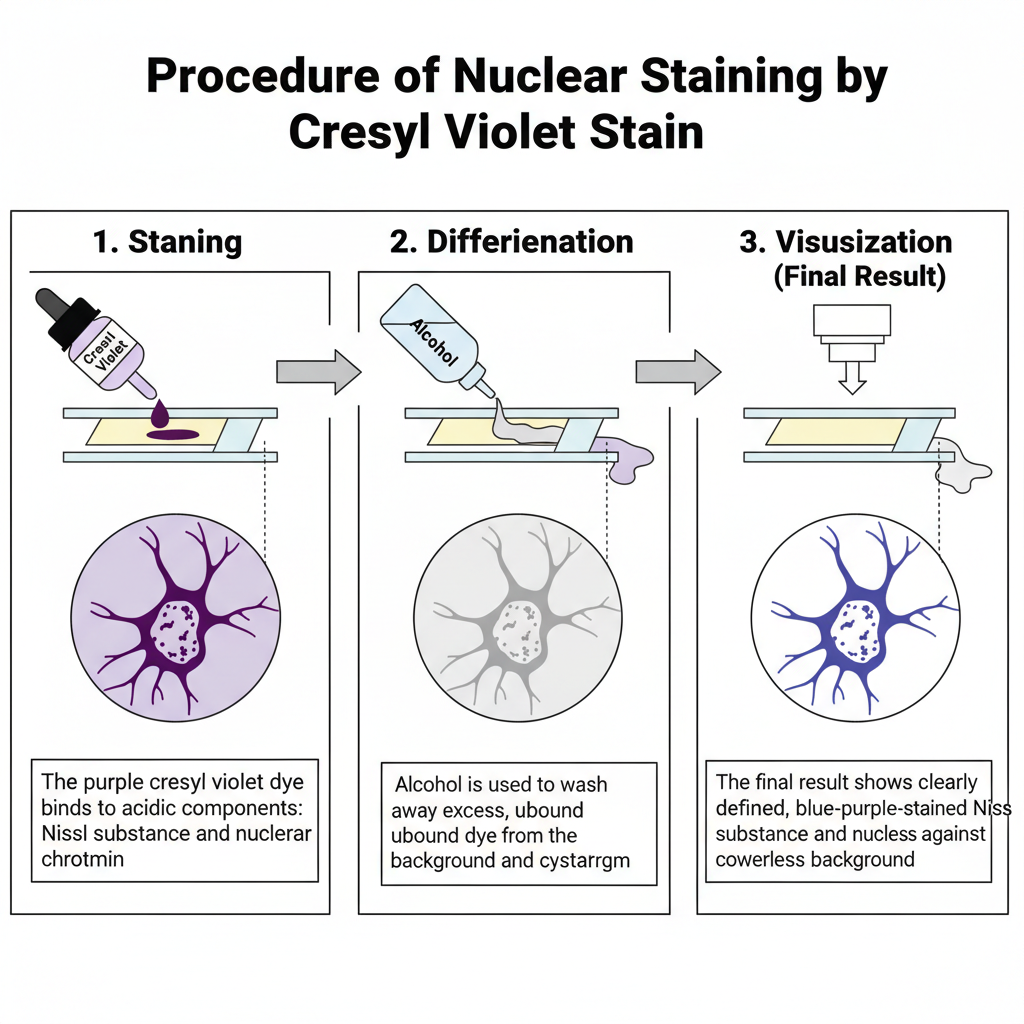
I. Specimen Preparation and Fixation
Formalin is generally the preferred fixative. Other fixatives can also be used but the staining behaviour may change depending on the basic dye. For example thionin is suitable for alcohol-fixed tissues but does not stain formalin fixed tissue well. After fixation, the tissue is processed and sectioned.
II. Core Staining Procedure
1. Staining
In this step, the tissue section is stained with cresyl violet dye. The dye binds with Nissl substance and nuclear chromatin. These are highly acidic components of the neuron.
2. Differentiation
The stained section is differentiated in alcohol. This step removes excess dye. Differentiation is continued until the background becomes colourless.
3. Visualization
The Nissl substance appears blue to purple. The nuclear chromatin also take similar colour. The background remain colourless after proper differentiation.
Uses
- It is used for neuron identification.
- It is used to assess neuronal damage by showing loss of Nissl substance.
- It is used for staining nuclear chromatin and Nissl substance blue to purple.
Advantages
- It is useful for identifying neurons.
- It is effective for assessing neuronal damage by loss of Nissl substance.
- It stains nuclear chromatin and Nissl substance blue to purple.
- Differentiation time can be adjusted to stain nuclei or only Nissl substance.
Limitations
- Choice of basic dye must match fixation type because some dyes do not stain well on certain fixatives.
- The differentiation step in alcohol must be controlled to avoid excess background.
Nuclear Staining by Nuclear Fast Red
Principle
The principle of nuclear staining by Nuclear Fast Red depends on its property as a basic dye which binds with the acidic components of the nucleus. The nucleus contains DNA and RNA, and these materials are acidic in nature, so the nucleus behaves as a basophilic structure. Because Nuclear Fast Red has a positively charged chromophore, it attaches strongly to these basophilic elements and stains the nuclei pink to red. It is mainly used as a counterstain in different special staining methods where the background tissue requires a contrasting colour.
In silver impregnation techniques where reticular fibres become black, Nuclear Fast Red is applied to colour the nuclei and surrounding tissue elements pink to red. It is also used in the colloidal iron method for acid mucins where the mucins are stained dark blue, and the nuclear counterstain provides a clear contrast. In lipid stains such as Sudan Black B, it stains the nuclei red and helps in distinguishing the cellular structures. For effective staining, the dye must be mixed properly before use, and overstaining must be avoided because excess colour can mask the delicate pigments formed by other reactions.
The stain therefore works mainly on the acid–base affinity between the basic dye and the acidic nuclear contents, giving a clear and contrasting nuclear appearance in special histological procedures.
Procedure
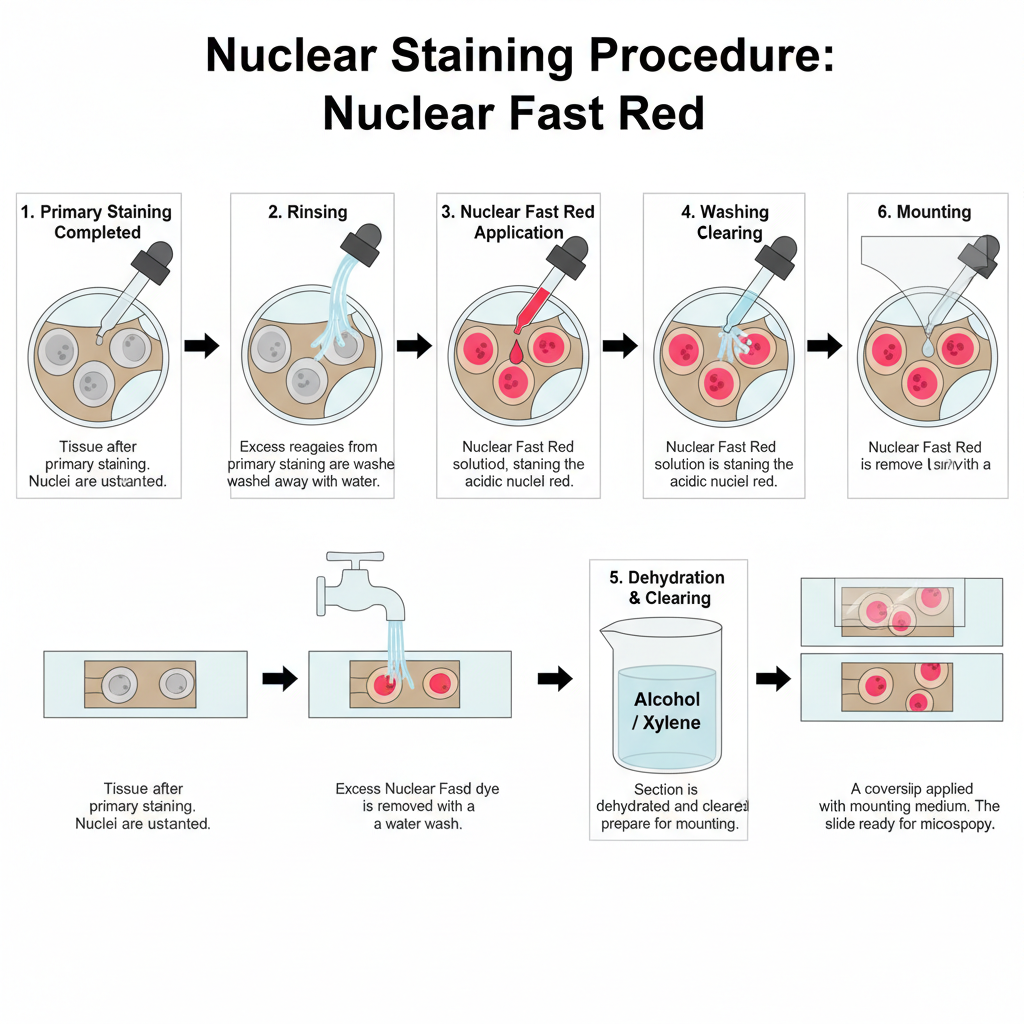
I. Preparation Before Counterstaining
The tissue must be fixed and processed. Paraffin embedded sections are used. The sections is deparaffinized and taken to water. The primary staining steps are completed first. These are different depending on the technique like reticulin staining, iron staining or colloidal iron staining.
II. Application of Nuclear Fast Red
1. Primary staining completed
The main stain is already applied to the tissue. For example silver reaction for reticulin, Prussian blue for iron or Sudan Black for lipids.
2. Rinsing
In this step, excess reagents from primary staining are washed off. It removes leftover chemicals such as sodium thiosulfate in silver impregnation.
3. Nuclear fast red staining
The section is treated with nuclear fast red. The dye binds with the acidic nuclear material. The nuclei turn pink to red.
4. Washing
The slide is washed in water to remove extra dye. The cytoplasm remain lightly coloured by the primary stain only.
5. Mounting
The section is dehydrated, cleared and mounted. It is sealed with coverslip for microscopic view.
Uses
- It is used as a counterstain for various histological special stains.
- It is used in reticulin stains to color nuclei pink to red.
- It is used in iron stains (Perl’s method) to stain nuclei red.
- It is used in colloidal iron method as a nuclear counterstain.
- It is used in lipid stains (Sudan Black B) as a red nuclear counterstain.
Advantages
- It is an excellent counterstain to improve contrast.
- It stains nuclei pink to red in silver-based stains so the main structures stand out.
- It is the typical counterstain used in detecting ferric iron deposits.
Limitations
- It is mainly a counterstain.
- Overstaining may hide pigments like iron and reduce visibility.
Nuclear Staining by Giemsa Stain
Principle
The principle of nuclear staining by Giemsa stain is based on its nature as a polychromatic Romanowsky-type stain that contains both basic and acidic dye components. The mixture is prepared from methylene blue as the basic dye together with acidic dyes such as eosin and azure compounds. These dyes interact differently with acidic and basic elements inside the cell, which produces multiple colours in the stained tissue. It is this combination of neutral dyes that allows Giemsa to give a characteristic staining pattern.
The nucleus behaves as a basophilic structure because nucleic acids are acidic in nature. The basic dye portion of the Giemsa mixture binds strongly with these acidic nuclear materials, and the nucleus becomes dark blue. The acidic dye components then stain the cytoplasmic elements pink or purple depending on their chemical properties. This differential staining gives a clear contrast and helps in identifying various cell types, especially in hematopoietic tissues.
Differentiation with a weak acid solution is an important step because it enhances the colour range and improves the clarity of the stained cells. The stain is widely used for identifying blood cells and diagnosing disorders such as leukemia and lymphoma, where both nuclear and cytoplasmic details are important. Fresh working solution is usually prepared because the stain is unstable, and proper pH of the buffer is necessary to obtain consistent nuclear staining.
Procedure
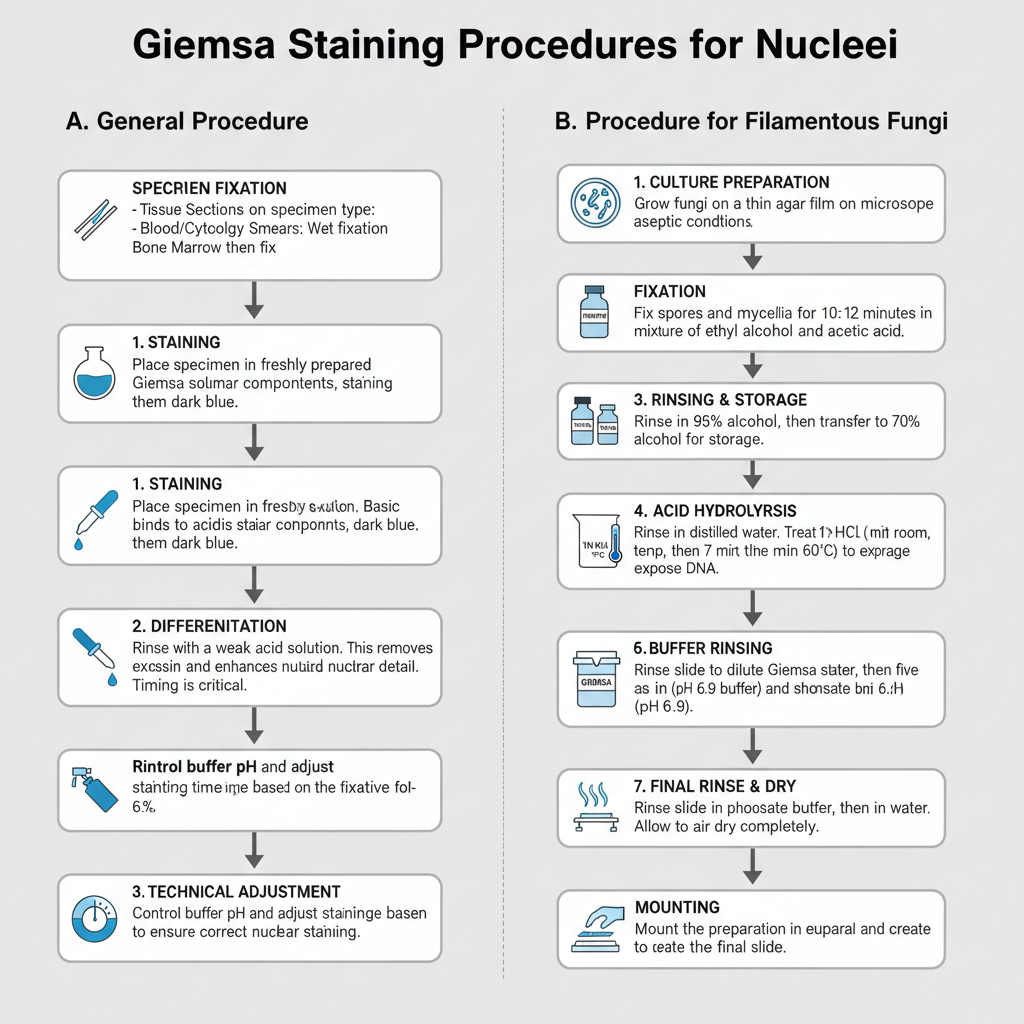
I. General Steps and Fixation
Fixation depends on specimen type. Tissue sections are fixed before staining. Blood films and cytology smears require immediate wet fixation to preserve chromatin. Bone marrow smears are usually air dried and then fixed. The Giemsa working solution is prepared fresh every time because it is not stable.
1. Staining
In this step, the section is placed in Giemsa solution. The nucleus becomes dark blue as basic dye binds to nuclear components.
2. Differentiation
The stained section is differentiated with weak acid solution. It removes excess colour and improves nuclear detail. The timing of acid differentiation is important to balance shades.
3. Technical adjustment
The pH of buffers must be controlled. Different fixatives may require changes in pH or staining time to obtain correct nuclear staining.
II. Procedure for Filamentous Fungi
This method is used for studying nuclei in filamentous fungi. It requires chemical fixation and acid hydrolysis to expose DNA due to the tough fungal cell wall.
1. Culture preparation
Fungi is grown on thin agar film on slides. The growth is maintained under aseptic conditions.
2. Fixation
Spores and mycelia are fixed for 10–12 minutes in a 3:1 mixture of ethyl alcohol and acetic acid. A 3:1:1 mixture with lactic acid can also be used.
3. Rinsing and storage
The preparation is rinsed in 95% alcohol. It is transferred to 70% alcohol where it can be stored for some days.
4. Water rinse
The slide is rinsed in distilled water.
5. Acid hydrolysis
In this step, the preparation is treated with 1N HCl at room temperature for 5 minutes. Then it is kept in 1N HCl at 60°C for 7 minutes. This exposes the DNA.
6. Buffer rinsing
The slide is rinsed in water and then in phosphate buffer (pH 6.9). Five changes are done for each.
7. Staining
The preparation is transferred into dilute Giemsa stain. A solution of 1 mL Giemsa in 15 mL phosphate buffer pH 6.9 is used. Staining is continued for 2 hours.
8. Final rinsing and drying
The slide is rinsed in phosphate buffer and then in water. It is air dried.
9. Mounting
The preparation is mounted in euparal. A coverslip is placed to complete the slide.
Uses
- It is used to identify hematopoietic cell lineages.
- It is used to diagnose blood disorders and tumors.
- It is used to detect microorganisms like H. pylori, Rickettsia, Toxoplasma gondii, and Plasmodium vivax.
- It is used for cytology screening of cervical cells.
- It is used for fungal nuclear staining in filamentous fungi.
Advantages
- It is a polychromatic stain.
- It identifies cell lineages and developmental stages in hematopoietic tissues.
- It helps in diagnosing tumors and blood disorders.
- It shows cellular morphology and nuclear-cytoplasmic detail clearly.
- Acid differentiation enhances the color contrast.
- It can detect microorganisms such as H. pylori, Rickettsia, and Toxoplasma gondii.
Limitations
- Working solution is unstable and must be prepared fresh.
- pH of solutions and buffers is critical.
- Different fixation methods may need pH adjustments to get proper staining.
Result of Nuclear Staining
Nuclear Staining by DAPI
- It gives clear visualization of nuclei and chromosomes as blue fluorescence.
- It shows strong emission when bound with dsDNA which helps in multicolor fluorescence imaging.
- It is used as a reliable nuclear counterstain because the stained nuclei appear bright against other fluorophores.
Nuclear Staining by Hoechst
- It stains nuclei in live cells showing blue fluorescence.
- It is suitable for multi-color imaging because its emission fits well with green or red fluorophores.
- It helps to clearly identify the nuclear region during live-cell observations.
Nuclear Staining by H&E
- It clearly shows nuclei in blue and cytoplasm in pink.
- It gives differential staining of many tissue elements with a single protocol.
- It provides clear contrast for routine histological diagnosis.
Nuclear Staining by Feulgen Stain
- It selectively demonstrates DNA by producing a magenta colored reaction.
- It gives clear visualization of DNA distribution within the nucleus.
- A light green counterstain can be added to enhance the contrast.
Nuclear Staining by Methyl Green-Pyronin Y
- It differentiates DNA and RNA in the same tissue section.
- It stains DNA green or blue-green and RNA red for clear interpretation.
- It provides a simple way to study nucleic acid localization.
Nuclear Staining by Cresyl Violet
- It shows Nissl substance and nuclear chromatin in blue to purple.
- It is useful to identify neuronal cells.
- Differentiation produces a clean colorless background.
Nuclear Staining by Nuclear Fast Red
- It provides pink to red nuclear staining as a counterstain.
- It offers good contrast when used with silver-based or iron stains.
Nuclear Staining by Giemsa Stain
- It shows nuclei dark blue and cytoplasm pink with good clarity.
- It gives polychromatic staining allowing visualization of multiple cell details.
- It helps to distinguish leukocytes and other blood elements due to its color range.
- AAT Bioquest. (2025). Cellular fluorescence microscopy troubleshooting & best practices.
- Cell Signaling Technology. (2019). Flow cytometry, Triton X-100 permeabilization protocol.
- Champouret, N., & Kamoun, S. (2004). An improved protocol for the protoplast method of stable DNA transformation of Phytophthora infestans. Retrieved from http://www.KamounLab.net
- DakoCytomation. (n.d.). Dako guide to special stains.
- Danion, F., van Rhijn, N., Dufour, A. C., Legendre, R., Sismeiro, O., Varet, H., Olivo-Marin, J.-C., Mouyna, I., Chamilos, G., Bromley, M., Beauvais, A., & Latgé, J.-P. (2021). Aspergillus fumigatus, one uninucleate species with disparate offspring. Journal of Fungi, 7(1), 30. https://doi.org/10.3390/jof7010030
- de Bekker, C., Wiebenga, A., Aguilar, G., & Wösten, H. A. B. (2009). An enzyme cocktail for efficient protoplast formation in Aspergillus niger. Journal of Microbiological Methods, 76(3), 305–306. https://doi.org/10.1016/j.mimet.2008.11.001
- Editorial Team. (2024, May 18). Principle for nuclear staining of filamentous fungi. Biology Notes Online.
- Fernando, L. D., Dickwella Widanage, M. C., Shekar, S. C., Mentink-Vigier, F., Wang, P., Wi, S., & Wang, T. (2022). Solid-state NMR analysis of unlabeled fungal cell walls from Aspergillus and Candida species. Journal of Structural Biology X, 6, 100070. https://doi.org/10.1016/j.yjsbx.2022.100070
- Fischer, R. (1999). Nuclear movement in filamentous fungi. FEMS Microbiology Reviews, 23(1), 39–68. https://doi.org/10.1111/j.1574-6976.1999.tb00391.x
- Gealt, M. A., Sheir-Neiss, G., & Morris, N. R. (1976). The isolation of nuclei from the filamentous fungus Aspergillus nidulans. Journal of General Microbiology, 94(1), 204–210. https://doi.org/10.1099/00221287-94-1-204
- Jung, M. K., Ovechkina, Y., Prigozhina, N., Oakley, C. E., & Oakley, B. R. (2000). The use of beta-D-glucanase as a substitute for Novozyme 234 in immunofluorescence and protoplasting. Fungal Genetics Reports, 47(1), Article 8. https://doi.org/10.4148/1941-4765.1204
- KEYENCE. (n.d.). Top 10 microscopy troubleshooting tips for clearer fluorescence images.
- OracleBio Limited. (2025). Combating autofluorescence in multiplex IF image analysis.
- Otali, D. (2007). The combined effect of formalin fixation and individual steps in tissue processing on immunorecognition [Master’s thesis, University of Alabama at Birmingham]. UAB Digital Commons. https://digitalcommons.library.uab.edu/etd-collection/3618
- Parker, N., Schneegurt, M., Tu, A.-H. T., Lister, P., & Forster, B. M. (2016). 2.4 Staining microscopic specimens. In Microbiology. OpenStax. Retrieved from https://openstax.org/books/microbiology/pages/2-4-staining-microscopic-specimens
Proteintech Group, Inc. (2025). How to reduce autofluorescence. - Rahman, M. A., Sultana, N., Ayman, U., Bhakta, S., Afrose, M., Afrin, M., & Haque, Z. (2022). Alcoholic fixation over formalin fixation: A new, safer option for morphologic and molecular analysis of tissues. Saudi Journal of Biological Sciences, 29(1), 175–182. https://doi.org/10.1016/j.sjbs.2021.08.075
- Thermo Fisher Scientific. (2025). DAPI counterstaining protocols.
- Thermo Fisher Scientific. (2025). DAPI protocol for fluorescence imaging.
- Thermo Fisher Scientific. (2025). Fixation strategies and formulations used in IHC staining.
- Thermo Fisher Scientific. (2025). HCS LIVE/DEAD green kit using Hoechst 33342 protocol.
- Thermo Fisher Scientific. (2025). Hoechst 33342 protocol for imaging.
- Van Genechten, W., & Van Dijck, P. (2021). Fluorescent toys ‘n’ tools lighting the way in fungal research. FEMS Microbiology Reviews, 45(5). https://doi.org/10.1093/femsre/fuab013