DNA Analyzer is a precision scientific instrument that is used to automate the process of genetic analysis. It is also commonly referred to as DNA sequencer. It determines the exact order of four nucleotide bases such as adenine (A), guanine (G), cytosine (C), and thymine (T) present in a DNA sample.
Modern DNA analyzer combines advanced fluidics, high-resolution optics and computational algorithms for processing genetic samples quickly and in large volume. It is used to study the genetic sequence of DNA and gives accurate genetic information.
Many DNA analyzers also work as automated electrophoresis system. It is used for quality control of DNA or RNA fragments by measuring their size, quantity and overall integrity before further laboratory applications. DNA analyzer is important in disease diagnostics, personalized medicine, forensic science and agricultural improvement.
DNA Analyzer Principle
DNA Analyzer is based on the principle of automated capillary electrophoresis. In this process, DNA fragments are separated and detected according to their size. The DNA samples are first labelled with specific fluorescent dyes.
The labelled DNA samples are injected into narrow capillaries filled with polymer and then high-voltage electric field is applied. Due to negative charge, DNA molecules move through the capillary. Smaller DNA fragments move faster than larger fragments through the polymer matrix.
As the separated DNA fragments pass through the detection window, laser excites the fluorescent tags. The emitted light is detected by a sensor such as charge-coupled device (CCD) camera. Then the computer system converts these fluorescent signals into coloured peaks.
This visual graph is called electropherogram or chromatogram. It is used to determine the exact sequence of nucleotide bases or to measure the size of DNA fragments.
Parts of DNA Analyzer
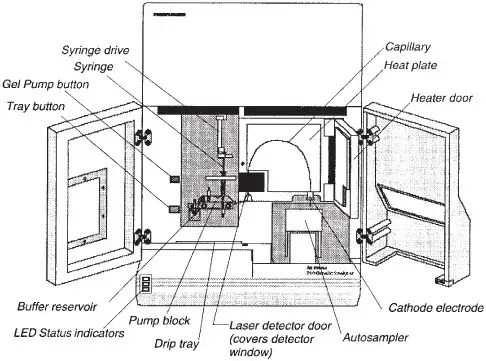
The following are the important parts of DNA analyzer-
- Capillary Electrophoresis Instrument
It is the main machine of DNA analyzer. It is used to automate the separation and reading of DNA samples. - Capillary Array
Capillary array is a preassembled set of capillaries. These capillaries may be 48 or 96 in number. DNA fragments migrate through these capillaries during electrophoresis. - Separation Matrix
Separation matrix is a special polymer that fills the capillaries. It helps in separation of DNA fragments according to their size. POP-7 polymer is commonly used in DNA analyzer. - Optical and Detection System
This system is used to detect the fluorescent labelled DNA fragments. It contains argon ion laser, capillary illumination system and CCD camera. The laser excites the fluorescent dye and CCD camera captures the emitted light. - Autosampler and Plate Stacker
Autosampler is a robotic part of the DNA analyzer. It holds sample plates and automatically loads them for continuous operation. It can hold 96-well or 384-well plates. - Fluidics System
Fluidics system manages the liquids required for electrophoresis. It contains polymer delivery pump, buffer reservoir, water reservoir and waste reservoir. It helps in movement of polymer and buffer during the experiment. - Temperature Controlled Oven
The oven houses the capillary array and maintains uniform temperature. It keeps the temperature suitable during sequencing run. The temperature may be maintained between 18-70°C. - Polymer Delivery Pump
Polymer delivery pump is used to deliver polymer into the capillary array. It is connected with polymer reservoir and anode buffer container by tubes. It helps in feeding polymer to the array. - Pump Block
Pump block is a pumping apparatus. It has syringe fittings, water seal, piston, pump chamber and water trap. It regulates the pump which delivers the polymer. - Lower Polymer Block
Lower polymer block is a polymer buffer valve present in the lower block. It regulates the flow of anode buffer from the buffer container. It is connected with the polymer delivery pump. - Polymer Pouch or Reservoir
Polymer pouch stores the polymer material required for the experiment. It is an important part because polymer is needed for separation of DNA fragments. - Cathode Buffer Reservoir
Cathode buffer reservoir contains 1x running buffer. This buffer is used to help electrophoresis and maintain the fluid level during experiment. - Waste Reservoir
Waste reservoir is used to collect leftover materials after the experiment. It may contain used buffer, water or polymer. - Water Reservoir
Water reservoir is used to store water or other liquid required in the procedure. It helps in maintaining the fluidics system. - Heat Plate
Heat plate helps to keep the capillary array at a constant temperature. It supports proper separation of DNA fragments. - Onboard Piercing Station
Onboard piercing station is used to pierce sealed sample plates. It helps in automatic processing of samples. - Internal Barcode Reader
Internal barcode reader scans the barcode of sample plates. It is used for automatic sample tracking. - Computer Workstation
Computer workstation is connected with the DNA analyzer. It is used to control the operation of the machine. It has a monitor and instrument control system. - Collection and Analysis Software
The software is used to collect raw optical data and analyze the result. It performs basecalling and interprets the genetic result. Data Collection Software, Sequencing Analysis Software and GeneMapper are examples. - Doors
DNA analyzer has two doors. One door is for the oven and another door is for the instrument. The oven door is present inside and the outer side has a glass panel. - Power Button
Power button is present outside the DNA analyzer. It is used to switch on or switch off the instrument. - LED Indicators
LED indicators are present near the power button. It shows whether the machine is on and working. - 96-well Plate
96-well plate is used for holding DNA samples. It is similar to the ELISA plate. Samples are placed in wells and then loaded into the analyzer.
Operating Procedure of DNA Analyzer
The following are the operating procedure of DNA analyzer-
- The first step is DNA extraction. High quality and pure DNA is isolated from biological sample such as blood, tissue or cells. It is done to get reliable result in further process.
- The target DNA region is amplified by Polymerase Chain Reaction (PCR). This process is also used in cycle sequencing.
- In PCR reaction, template DNA, primers, DNA polymerase, dNTPs and fluorescent labelled ddNTPs are used for sequencing. For fragment analysis, fluorescent labelled primers are used.
- The amplified PCR products are purified after amplification. Leftover primers, free nucleotides, salts and unincorporated fluorescent dyes are removed in this step.
- If contaminants are not removed, they may enter into the DNA analyzer with the sample. This can decrease the quality of data.
- The purified DNA sample is resuspended in a special loading solution such as Hi-Di Formamide. It keeps the DNA sample ready for loading.
- In fragment analysis, internal size standard is mixed with the sample. It helps to measure the size of DNA fragments during analysis.
- The prepared samples are placed into 96-well or 384-well sample plate. The plate is sealed properly to prevent evaporation.
- The instrument is started by following correct sequence. First the computer workstation is turned on and login is done as local administrator.
- Then the DNA analyzer instrument is switched on. The green indicator light is checked before starting the software.
- Data Collection Software is opened after the instrument is ready. This software is used to control the run and collect the data.
- The prepared sample plate is placed into the integrated autosampler stacker of the DNA analyzer. The instrument automatically loads the samples for run.
- The samples are injected into the capillary array filled with separation polymer matrix. High voltage electric field is applied for separation of DNA fragments.
- DNA fragments are separated according to their size. Smaller fragments move faster and larger fragments move slowly through the polymer.
- As the separated fragments pass through detection window, laser excites the fluorescent tags. The emitted fluorescence is detected by a specialized camera.
- The raw fluorescent data is transferred to the computer. The data is then analyzed by specialized software.
- Software such as Sequencing Analysis Software, SeqScape or GeneMapper is used for interpretation. It produces electropherogram and performs basecalling or fragment sizing.
Applications of DNA Analyzer
The following are the applications of DNA analyzer-
- DNA analyzer is used in basic genomic research. It is used for de novo sequencing, targeted resequencing, microsatellite based fragment analysis and validation of molecular cloning experiments.
- It is used in precision medicine and clinical diagnosis. It helps to identify genetic variation and mutation for diagnosis of hereditary diseases and disease risk.
- DNA analyzer is used in personalized healthcare. It helps to prepare treatment plan according to genetic condition of individual patient.
- It is used in oncology for detection of cancer causing somatic mutations. It also helps in liquid biopsy for monitoring tumour progression and selecting targeted therapy.
- It is used in pharmacogenomics. It helps to study how genetic makeup of a person affects drug metabolism and response to medicines.
- DNA analyzer is used to optimize drug dose. It also helps to avoid harmful drug reactions in patients.
- It is used in forensic science and criminal justice. DNA profiling helps in human identification and matching suspects with crime scene samples.
- It is used in Short Tandem Repeat (STR) profiling. This method is useful for criminal investigation and also for exonerating wrongly convicted persons.
- DNA analyzer is used in victim identification and kinship testing. It helps in mass disaster victim identification, missing person cases and checking family relationship.
- It is used in agriculture and food science. Genome sequencing helps in marker assisted breeding for disease resistant and drought tolerant crops.
- It is used to improve livestock traits. It also helps in food safety by detecting genetically modified organisms.
- DNA analyzer is used in infectious disease surveillance. It helps to identify viral and bacterial pathogens rapidly.
- It is used to track mutation and spread of diseases such as COVID-19. It also helps in monitoring antimicrobial resistance.
- DNA analyzer is used in environmental genomics. It helps to study soil microbiomes, endangered species and global biodiversity from environmental samples.
- It is used in evolutionary biology and anthropology. It helps to trace genetic lineage, human migration pattern and evolutionary relationship among different species.
Advantages of DNA Analyzer
The following are the advantages of DNA analyzer-
- DNA analyzer gives high quality sequencing data. It also reduces the total cost per sample by using less amount of DNA, reagents and separation matrix.
- It is a fully automated system. After the setup, it can process the sample without much manual work.
- It gives result in short time. The workflow becomes faster and reliable result can be obtained in minutes or hours instead of days or weeks.
- DNA analyzer has high optical sensitivity. It can detect fluorescent signals uniformly and accurately from the DNA fragments.
- It can work with different concentration of input DNA. Due to this, the sample consumption is reduced.
- Modern DNA analyzer can give increased read length. It helps to read longer DNA sequences in less run time.
- It reduces the total number of traces needed for a sequencing project. This makes the analysis more efficient.
- DNA analyzer has high throughput capacity. It can process one sample to 96 samples at the same time.
- It is useful for both DNA sequencing and fragment analysis. Thus, it is a versatile instrument in molecular biology laboratory.
- It has software for automated data analysis. The software helps in basecalling, allele calling and interpretation of genetic result.
- Many DNA analyzers are compact in design. They occupy less laboratory bench space.
- Some DNA analyzers are portable and can be used in field condition. These systems reduce the need to send sample to central laboratory.
- DNA analyzer is reliable and easy to maintain. It reduces laboratory overhead expenses and service cost.
Limitations of DNA Analyzer
The following are the limitations of DNA analyzer-
- DNA analyzer may face difficulty in reading complex DNA sequences. Secondary structures such as hairpin, GC-rich regions and long repeated bases can produce mixed and unreadable signals.
- It needs proper quantity of DNA sample. Too much template DNA can give off-scale data or early termination of sequence.
- Very low amount of DNA sample can produce weak signal or no signal. So, DNA concentration should be optimized before running the instrument.
- Contaminants present in DNA sample can affect the sequencing reaction. Salts, proteins and phenol can inhibit the reaction and reduce the quality of result.
- Reagents and consumables used in DNA analyzer have short life span. Sequencing buffer, separation polymer and other materials should be used within their proper time limit.
- Capillary array has limited use. It is generally replaced after fixed number of runs or after its expiry period.
- DNA analyzer is sensitive to hardware and environmental condition. Air bubbles in lines, fluid leakage and room temperature changes can disturb the run.
- Unstable current may occur due to physical or fluidic problems. It can reduce peak resolution and affect data quality.
- Fluorescent dyes used in DNA analyzer are sensitive. They can degrade by light exposure, heat, oxidation or acidic pH.
- Dye degradation can cause missing peaks or irregular peaks in the result. This makes the data difficult to interpret.
- DNA analyzer produces large amount of raw data. Regular software maintenance and database cleanup are required.
- If the database becomes too full, the system may lock and new runs may not start properly.
Precautions of DNA Analyzer
The following are the precautions of DNA analyzer-
- Proper personal protective equipment should be worn while operating DNA analyzer. Gloves, eye protection and lab coat are used because some reagents may contain hazardous chemicals.
- Only clean distilled water, deionized water or laboratory grade water should be used for cleaning and filling reservoirs. Tap water should not be used because it can contaminate the system.
- Fluorescent labelled DNA and sequencing reactions should be protected from light, heat, oxygen and acidic condition. These conditions can damage the dye labels.
- Samples should be properly sealed before loading into the instrument. Tube septa or heat seals are used to prevent evaporation and exposure to air.
- Old polymer should not be mixed with fresh polymer. It can reduce the quality of data.
- Running buffer should be replaced regularly. Reservoir volume should be measured correctly with graduated cylinder and not only by seeing fill lines.
- Routine maintenance should be done regularly. Parts such as water trap should be flushed weekly to prevent polymer accumulation and blockage.
- Capillary array and other parts should be handled gently. If any part becomes stuck, excessive force or tools should not be used.
- Distilled water may be used to loosen the stuck part. If the problem remains, service should be called.
- Vibration should be avoided during the run. The instrument should not be touched and doors should not be opened after starting the run.
- Main door, oven door and detector cell door should be properly closed before starting the run. Open doors can stop the machine.
- Correct booting sequence should be followed. Computer should be turned on and logged in before switching on the instrument.
- Computer maintenance should be done carefully. In systems with built-in database, the database drive should not be defragmented because it can cause software problem.
References
- Thermo Fisher Scientific. (n.d.). 3730xl DNA Analyzer – FAQs.
- Applied Biosystems. (2006). DNA Sequencing by Capillary Electrophoresis Chemistry Guide.
- Colarossi, J. (2026, January 23). AI generates short DNA sequences that show promise for gene therapies. Broad Institute.
- ANDE. (n.d.). ANDE – Rapid DNA Technology.
- Agilent Technologies. (2024). Agilent TapeStation and Bioanalyzer Systems: Systems, consumables, and supplies.
- Applied Biosystems. (2006). Applied Biosystems 3730 and 3730xl DNA Analyzers. LabWrench.
- Thermo Fisher Scientific. (n.d.). Applied Biosystems 3730/3730xl DNA Analyzers Support.
- Agilent Technologies. (2019). Automated Nucleic Acid Sample Quality Control in NGS Workflows. Agilent TapeStation systems.
- Abcam. (n.d.). Bioanalyzer and TapeStation Analysis for ChIPSeq, CUT&RUN, and CUT&Tag.
- UBOS. (n.d.). Breakthroughs in DNA Sequencing Drive Down Costs and Expand Applications.
- College of American Pathologists. (n.d.). CAP 15189 Accreditation Program Frequently Asked Questions.
- College of American Pathologists. (n.d.). CAP 15189 Accreditation Program.
- Promega Corporation. (n.d.). Capillary Electrophoresis (CE) for Forensic DNA Analysis.
- PubMed Central. (n.d.). Case report: Liquid biopsy, the sooner the better?
- PubMed Central. (n.d.). Clinical applications of cell-free DNA-based liquid biopsy analysis.
- Graf, E. (2018). Comparison of DNA Assays Using the 4200 TapeStation System and 2100 Bioanalyzer System. Agilent Technologies.
- Comprehensive Analysis of DNA Sequencing Technologies: Mechanical Architectures, Biochemical Mechanisms, and the Global Genomic Infrastructure. (n.d.).
- Promega Corporation. (n.d.). DNA Purification | DNA Extraction Methods.
- Patil, V. (2026, March 16). DNA Sequencing Market Size & Trends 2026-2033. Coherent Market Insights.
- Mordor Intelligence. (2026). DNA Sequencing Market Size & Share Report.
- CD Genomics. (n.d.). DNA Sequencing in Practice: Crop Customization, Livestock Optimization and Microbiome-Based Agriculture.
- Mathiesen, A. M. (2024). DNA sequencing. Science | Research Starters. EBSCO.
- Mobley, I. (2025, July 16). DNA Sequencing: How to Choose the Right Technology. Front Line Genomics.
- Wikipedia. (2026, March 19). DNA sequencer.
- Abcam. (n.d.). DNA sequencing: Key methods and applications.
- Abcam. (n.d.). DNA sequencing: Decoding the genetic blueprint.
- Yale Research. (n.d.). DNA/RNA QC analysis service.
- Alketbi, S. K. (2024). Emerging Technologies in Forensic DNA Analysis. Perspectives in Legal and Forensic Sciences, 1(1), 10007.
- MyBioSource. (n.d.). Forensic Applications of PCR: DNA Profiling and Analysis.
- Thermo Fisher Scientific. (n.d.). Fragment Analysis Workflow.
- Intel Market Research. (n.d.). Gene Sequencing Market Outlook 2025-2032.
- Thermo Fisher Scientific. (n.d.). How a Flow Cytometer Works.
- Oxford Nanopore Technologies. (n.d.). How nanopore sequencing works.
- CD Genomics. (n.d.). How to Sequence a Gene: Step-by-Step Experiment Workflow.
- CD Genomics. (n.d.). How to Troubleshoot Sequencing Preparation Errors (NGS Guide).
- The University of Melbourne. (n.d.). INTRODUCTION TO FLOW CYTOMETRY.
- myLabCompliance.io. (2025). ISO 15189 vs CLIA Standards: Understanding the Differences.
- Thermo Fisher Scientific. (n.d.). IntegenX by Thermo Fisher Scientific RapidHIT Instrument and Consumables.
- IPIRA-Berkeley. (n.d.). IntegenX | Intellectual Property & Industry Research Alliances.
- PubMed Central. (n.d.). Integration of liquid biopsy and pharmacogenomics for precision therapy of EGFR mutant and resistant lung cancers.
- PubMed Central. (n.d.). International Organization for Standardization (ISO) 15189.
- Geneious. (n.d.). Introduction to DNA Sequencing.
- Thermo Fisher Scientific. (n.d.). Liquid Biopsy Presentations and Case Studies.
- Illumina. (n.d.). MiSeq System specifications | Key specifications and performance parameters.
- Illumina. (2018). MiSeq System Guide.
- Illumina. (2024). MiSeq System specification sheet.
- Illumina. (n.d.). MiSeq System | Rapid and cost-effective sequencing.
- Illumina. (n.d.). MiSeq i100 Series Specifications | Run time, output, and more.
- Integrated DNA Technologies. (n.d.). NGS Workflow Steps.
- Illumina. (n.d.). NGS Workflow Steps | Illumina sequencing workflow.
- Illumina. (n.d.). NGS vs Sanger Sequencing.
- PatSnap Insights Team. (2026, April 20). Nanopore DNA sequencing technology landscape 2026. PatSnap.
- Wikipedia. (n.d.). Nanopore sequencing.
- Feng, Y., Zhang, Y., Ying, C., Wang, D., & Du, C. (2015). Nanopore-based fourth-generation DNA sequencing technology. Genomics, Proteomics & Bioinformatics, 13(1), 4-16.
- Roche Diagnostics. (n.d.). Next-generation sequencing (NGS) platform.
- CD Genomics. (n.d.). Overview of Nanopore Sequencing Technology.
- CD Genomics. (n.d.). Overview of PacBio SMRT sequencing: principles, workflow, and applications.
- LabX. (n.d.). PCR and Thermal Cycler Technology: Fundamentals, Applications, and Emerging Trends in DNA Amplification.
- DNA Technologies Core. (n.d.). Pacbio Sequencing.
- Wikipedia. (2025, November 9). Rapid DNA.
- FBI. (n.d.). Rapid DNA — LE.
- Roche Diagnostics. (n.d.). Roche SBX Webinar: Introduction to sequencing by expansion.
- Mannion, J., & Kokoris, M. (2025, May 20). Sequencing by expansion: A novel approach to next generation sequencing. LabLeaders – Roche Diagnostics.
- MGH DNA Core. (n.d.). Sanger DNA Sequencing: Troubleshooting.
- Thermo Fisher Scientific. (n.d.). Sanger Sequencing Support—Troubleshooting.
- CD Genomics. (n.d.). Sanger Sequencing: Introduction, Principle, and Protocol.
- Wikipedia. (2026, March 29). Sanger sequencing.
- Abcam. (n.d.). Sanger sequencing: Process and applications.
- PacBio. (2026, March 30). Sequencing 101: long-read sequencing.
- bioRxiv. (n.d.). Sequencing by Expansion (SBX) – a novel, high-throughput single-molecule sequencing technology.
- Roche Diagnostics. (n.d.). Sequencing by expansion (SBX) technology.
- Wikipedia. (2026, March 11). Single-molecule real-time sequencing.
- MyBioSource. (n.d.). The Advancements and Applications of Genome Sequencing.
- Alliant University. (n.d.). The Role of DNA Analysis in Forensic Science.
- Griffith University. (n.d.). Troubleshooting.
- Ultima Genomics. (n.d.). UG 100™ Sequencing Platform | High-Throughput, Cost-Effective Genomics.
- University of Minnesota Genomics Center. (n.d.). Ultima Genomics UG 100 Sequencing.
- Kiernan, E., & Mathews, K. (2026, April 12). Ultima Genomics Whole Genome Germline Overview. Broad Institute WARP.
- Agilent Technologies. (n.d.). What is Flow Cytometry?