What is Checkpoint?
A checkpoint is a control point in cell cycle. It is a internal surveillance mechanism present in eukaryotic cell. It ensures that cell division occurs in proper manner. It act like a quality control step where progression is checked before moving to next phase.
It is the process where cell checks cell size nutrient availability and completion of DNA replication. Integrity of DNA is also monitored. Correct attachment of chromosomes is verified during division.
If condition is favourable the cell cycle is moved forward by positive regulatory molecules. These are cyclins and cyclin dependent kinases (CDKs). If problem is detected such as DNA damage or replication error negative regulatory molecules act as roadblock and progression is stopped. This pause is provided for repair of DNA. If damage is not repaired it can trigger apoptosis (programmed cell death). It prevent replication of damaged cells and it helps in avoiding mutation and cancer.
These are three major checkpoints in cell cycle.
G1 checkpoint (restriction point) is present in G1 phase. Cell size reserves and genomic DNA damage is assessed. It determines whether cell will enter in division process or not.
G2 checkpoint is present in G2 phase. Entry into mitotic phase is blocked if DNA is damaged. It ensures that all chromosomes is fully replicated.
M checkpoint (spindle checkpoint) occurs during metaphase. It ensures that sister chromatids is correctly attached to spindle microtubules. It is checked before chromatids are separated.
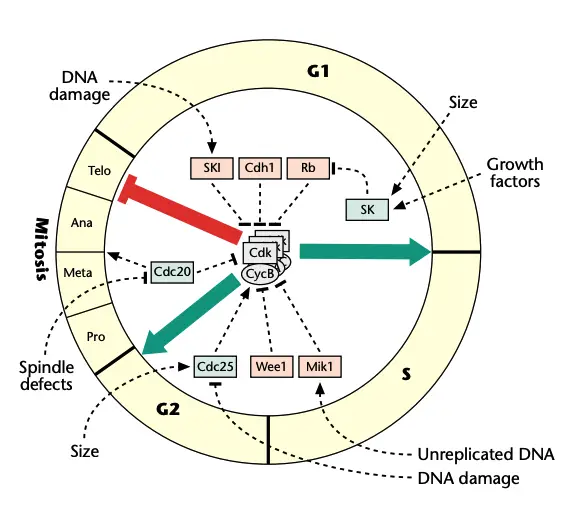
The Cell Cycle Engine
- The cell cycle engine is a control system which drives progression of cell cycle. It is driven by kinase complexes.
- Core components is CDK (cyclin dependent kinase) and cyclin. CDK is catalytic subunit and cyclin is regulatory subunit. The complex is formed and it phosphorylates specific target substrates.
- Primary function is to ensure that DNA is replicated only once in a cell cycle. It also ensure that chromosomes is segregated equally into daughter cells at correct time.
- Regulation by cyclin availability is important. Formation of active CDK cyclin complex depends on periodic synthesis of cyclins. Proteolysis (degradation) of cyclins also occurs throughout cell cycle.
- Regulation by phosphorylation occurs for activation. CDK cyclin complex must be phosphorylated by CDK activating complex (CAK) to become fully active. Inhibitory phosphorylation is done by wee1 and myt1 which keeps complex inactive. cdc25 family phosphatases removes inhibitory phosphates and complex becomes active.
- Regulation by inhibitor proteins is referred to as negative regulation. Cyclin dependent kinase inhibitors (CKIs) inhibits activity of engine. These are two main groups.
- INK4 family includes p16 p15 p18 and p19. These proteins bind with CDK4 and CDK6 and inhibition is done.
- Cip Kip family includes p21 p27 and p57. These have broader specificity. It can inhibit multiple cyclin CDK complexes such as cyclin D dependent kinases and cyclin E/A with CDK2.
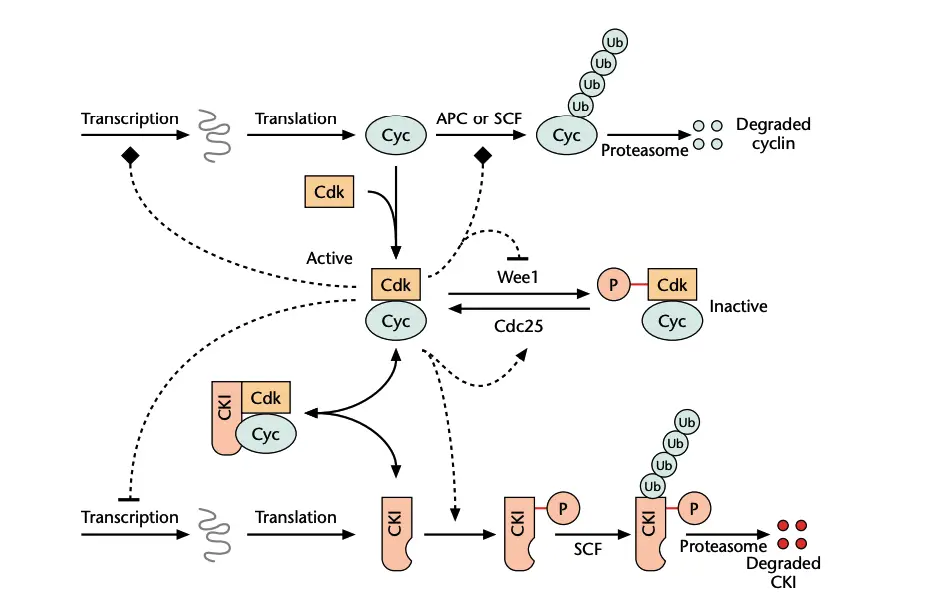
The Cell Cycle
Cell cycle is a series of events in which cell grows and divides to form two daughter cells. It is the process in which DNA is replicated and cell contents is distributed in proper manner. It includes interphase and M phase.
Interphase
Interphase is longest stage of cell cycle. In this stage cell grows and preparation is done for division. It is subdivided into three phases (G1 phase S phase and G2 phase).
- G1 Phase (Gap 1)- G1 phase is a growth phase. Cell is metabolically active and continuous growth occurs. Proteins and organelles is synthesized in this phase. It is prepared for DNA replication.
- S Phase (Synthesis)- S phase is referred to as synthesis phase. In this phase genomic DNA is replicated exactly and two identical copies is formed. Each chromosome forms sister chromatids. Centrosomes duplication also occurs in this phase.
- G2 Phase (Gap 2)- G2 phase is final growth period before division. Energy reserves is replenished. Specific proteins required for cell division is synthesized in this phase.
M Phase (Mitotic Phase)
M phase is the stage where cell divides. It consists of two main processes (mitosis and cytokinesis).
- Mitosis- Mitosis is the process in which duplicated chromosomes is separated and pulled to opposite poles. Two identical daughter nuclei is formed. It occurs in sequential manner. The stages are prophase prometaphase metaphase anaphase and telophase.
- Cytokinesis- Cytokinesis occurs after mitosis. Cytoplasm is divided and two distinct daughter cells is formed. It results in two independent daughter cells.
G0 Phase (Quiescent or Resting Phase)– G0 phase is an alternative state where cell exits from active cell cycle. In this stage cells are not preparing to divide. It may be temporary and cell can re-enter in G1 phase when external signals is present. Some cells remain permanently in G0 such as mature cardiac muscle cells and nerve cells.
What are the Checkpoints in the Cell Cycle?
Checkpoints in cell cycle is control points where cell cycle progression is checked. It ensures that events is completed properly before moving to next phase. These are major checkpoints.
G1 checkpoint (restriction point or START) occurs near end of G1 phase. It determines whether conditions are favourable for cell division. It is the point where cell commits to division process. It assesses cell size energy reserves growth factors and genomic DNA damage. If condition is unfavourable the cycle is stopped for repair. Cell may enter into G0 (resting phase). If DNA damage is not repaired apoptosis can occur.
G2 checkpoint (G2/M checkpoint) occurs at transition between G2 phase and mitotic phase. It prevents entry into mitosis if condition is not met. It also checks cell size and protein reserves. The major role is to ensure that chromosomes is fully replicated. It checks whether replicated DNA is free from damage. If problem is detected cell cycle is halted for completion of DNA replication or repair of damaged DNA before division.
M checkpoint (spindle or metaphase checkpoint) occurs near end of metaphase. It ensures that kinetochores of all sister chromatids is correctly attached to spindle microtubules from opposite poles. Separation of sister chromatids in anaphase is irreversible step so this checkpoint stops cycle until proper bipolar attachment is achieved. It prevents unequal distribution of chromosomes to daughter cells (aneuploidy).
G1 Checkpoints
- G1 checkpoint is a major checkpoint present near end of G1 phase. It is also referred to as restriction point in mammalian cells. In yeast it is called START point. It is also called as major checkpoint.
- Main purpose of G1 checkpoint is to act as quality control station. In this checkpoint cell commits irreversibly to cell division process. Once it is passed and cell enters S phase it is committed for DNA duplication. Extracellular growth signals is not required further for continuing proliferation.
- It evaluates internal and external conditions before progression. These are checked.
- Cell size is checked.
- Nutrient availability and adequate energy reserves is checked.
- External signals such as growth factors is checked.
- Genomic integrity is checked and DNA damage is verified.
- Key regulatory molecules drives progression. Positive regulators are cyclins (D and E) and cyclin dependent kinases (CDKs). Negative regulatory proteins act as roadblock if problem is detected. These are Rb p53 and p21. It stops cycle when DNA is damaged.
- If requirement is not met cell will not progress to S phase. These consequences may occur.
- Cell cycle can be halted temporarily for repair of damaged DNA by specific enzymes.
- Apoptosis (programmed cell suicide) can be triggered if DNA damage is beyond repair and duplication is prevented.
- Cell can enter into G0 phase (quiescent resting state) and it can wait for signals until conditions becomes favourable.
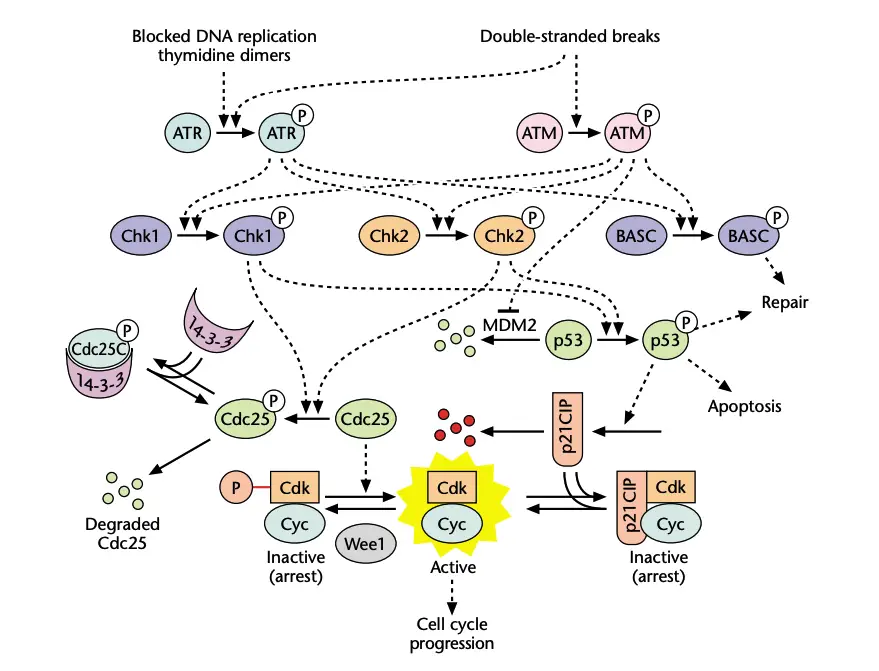
Functions of G1 Checkpoint
The G1 checkpoint is a regulatory mechanism that operates at the end of G1 phase. It is also referred to as restriction point or START point. It determines whether the cell will proceed to S phase or not. This checkpoint plays major role in maintaining genomic stability.
Some of the main functions are–
- Commitment to Cell Division –
It is the critical decision point where the cell becomes irreversibly committed to division. Once the cell passes this point it enters S phase and DNA replication is initiated. If the checkpoint is not satisfied the cell does not proceed further. - Monitoring Favorable Conditions –
The checkpoint evaluates internal and external environment of the cell. Adequate cell size nutrient availability and sufficient energy reserve is assessed. Presence of extracellular growth factors is also required. If these conditions is not favorable the progression is delayed. - Assessment of DNA Integrity –
It is the process by which genome is examined before DNA synthesis. DNA damage such as mutations or strand breaks is detected at this stage. If damage is present entry into S phase is blocked. This ensures that damaged DNA is not replicated. - Halting Cell Cycle Progression –
When unfavorable conditions or DNA damage is detected the cell cycle is arrested. Time is provided for DNA repair mechanisms to operate. This process occurs to prevent transmission of defective genetic material. - Directing Cells to Quiescent State (G0) –
If requirements for division is not met the cell exits active cycle and enters G0 phase. This is a resting state where cell remains metabolically active but does not divide. The cell can re-enter cycle when proper signals are received. - Triggering Programmed Cell Death –
If DNA damage is irreparable tumor suppressor proteins such as p53 is activated. p53 induces apoptosis (programmed cell death). This prevents replication of mutated or potentially cancerous DNA. Thus abnormal cells is eliminated from population. - Regulation of DNA Replication Machinery –
In response to genotoxic stress the p53 dependent pathway downregulates CDC7 kinase. CDC7 is essential for initiation of DNA replication. By suppressing CDC7 the G1 arrest is reinforced. This protects genome from instability.
Thus the G1 checkpoint functions as a major control point that determines cell fate and maintains integrity of genetic material before DNA replication.
G2 Checkpoint
- G2 checkpoint is also referred to as G2/M checkpoint. It is a surveillance mechanism which occurs at end of G2 phase. It occurs before cell enters into mitosis (M phase).
- Primary assessment of this checkpoint is to ensure that DNA replication is completed. It monitors genome and it checks that replicated DNA is not damaged. Like G1 checkpoint it also checks cell size and protein reserves required for division.
- If DNA damage is detected or replication is incomplete cell cycle progression is halted. Time is provided for completing replication or for repair of damaged DNA before division.
- DNA damage response is initiated by sensor kinases. These are ATM and ATR. ATM detects double strand DNA breaks and ATR detects single stranded DNA lesions. These sensor kinases activates transducer checkpoint kinases. These are Chk1 and Chk2.
- Chk1 and Chk2 enforces arrest. It phosphorylates and inhibits Cdc25 phosphatases. Activation of CDK1/Cyclin B complex is prevented. CDK1/Cyclin B complex is master regulator of mitotic entry.
- p53 helps in maintaining G2 arrest. It induces target genes such as p21 GADD45 and 14-3-3 sigma. These further inhibits CDK1/Cyclin B complex or it sequester the complex in cytoplasm.
- If DNA is repaired and replication is completed checkpoint signalling diminishes and entry into mitosis is allowed. If DNA damage is severe and it is not repaired apoptosis (programmed cell death) can occur. Mitotic catastrophe can also occur and proliferation of mutated cells is prevented.
Functions of G2 Checkpoint
The G2 checkpoint is a surveillance mechanism that operates at the end of G2 phase. It is also referred to as G2/M checkpoint. It prevents the cell from entering mitosis when DNA is damaged or replication is incomplete. This process is important for maintaining genomic stability.
Some of the main functions are–
- Assessment of DNA Integrity –
It is the process by which newly replicated DNA is examined before mitosis. The checkpoint verifies the absence of DNA lesions such as single-stranded gaps and double-strand breaks. If damage is detected the progression into M phase is stopped. This allows repair of damaged DNA before chromosome segregation. - Ensuring Completion of DNA Replication –
The checkpoint ensures that DNA replication is fully completed. Incomplete replication is sensed and mitotic entry is blocked. This process occurs when replication forks are stalled or stress is present. Time is provided for finishing the replication process. - Monitoring Cell Size and Energy Status –
It is also responsible for checking adequate cell size and energy reserve. Sufficient proteins and metabolic energy must be accumulated for mitosis. If requirements are not fulfilled the cycle is delayed. This ensures proper division of cell contents. - Regulation of Cyclin B1/CDK1 Complex –
The progression into mitosis is driven by Cyclin B1/CDK1 complex (Cdc2). Under normal conditions CDK1 is kept inactive by inhibitory phosphorylation by Wee1 and Myt1 kinases. Activation occurs when CDC25 phosphatase removes these phosphates. When DNA damage is detected this activation is prevented. CDK1 remains in inactive phosphorylated state. - Activation of DNA Damage Sensors –
DNA damage sensors activate ATM and ATR kinases. ATR mainly responds to single-stranded DNA and replication stress whereas ATM responds to double-strand breaks. These kinases phosphorylate downstream checkpoint kinases Chk1 and Chk2. This signaling cascade leads to arrest of cell cycle. - Inhibition of CDC25 Phosphatases –
Activated Chk1 and Chk2 phosphorylate CDC25 proteins. Phosphorylated CDC25C binds with 14-3-3 proteins and is sequestered in cytoplasm. CDC25A is targeted for ubiquitin mediated degradation. As a result Cyclin B1/CDK1 complex is not activated and mitosis is blocked. - Maintenance of Inhibitory Kinases –
Chk1 activates Wee1 kinase by phosphorylation. This ensures that CDK1 remains inhibited. The inhibitory phosphorylation is maintained and entry into mitosis is prevented. - Role of p53 in Sustained Arrest –
The tumor suppressor protein p53 is stabilized by ATM ATR and Chk2. It acts as transcription factor. It induces p21 which is a CDK inhibitor and blocks Cyclin/CDK complexes. It upregulates 14-3-3 sigma which binds Cyclin B1/CDK1 complex and retains it in cytoplasm. It also induces GADD45 which dissociates Cyclin B1/CDK1 complex. Transcription of Cyclin B1 and CDK1 genes is repressed. Thus prolonged G2 arrest is maintained. - Prevention of Genomic Instability –
If the checkpoint fails cells enter mitosis with damaged DNA. This results in chromosome mis-segregation and accumulation of mutations. Severe instability can lead to cancer development. In extreme damage mitotic catastrophe and cell death may occur.
Thus the G2 checkpoint plays a critical regulatory role in preventing transmission of damaged genetic material to daughter cells.
Metaphase (Spindle) Checkpoint
- Metaphase checkpoint is also referred to as spindle checkpoint. It occurs near end of metaphase stage of mitosis. It acts as strict surveillance mechanism and premature separation of sister chromatids (anaphase) is prevented.
- It monitors attachment of kinetochores. It ensures that kinetochores of each pair of sister chromatids is firmly anchored to spindle microtubules. These microtubules arise from opposite poles and bipolar tension is generated. Separation of sister chromatids is irreversible step so cycle will not proceed until this condition is met.
- If kinetochore is unattached or improperly attached a “wait” signal is generated. It is a diffusible inhibitory signal and cell cycle progression is halted. Even a single unattached kinetochore is sufficient for maintaining arrest.
- Inhibitory complex is formed in this checkpoint. It is referred to as Mitotic Checkpoint Complex (MCC). It is a tetrameric complex. Proteins present are Mad2 BubR1 (or Mad3) Bub3 and Cdc20.
- Target of inhibition is APC/C (Anaphase Promoting Complex/Cyclosome). MCC binds and blocks APC/C. APC/C is E3 ubiquitin ligase and its activity is essential for sister chromatid separation and mitotic exit.
- When all chromosomes are attached properly to spindle apparatus and tension is present checkpoint signal is silenced. MCC disassembles and inhibition is removed.
- Disassembly of MCC frees Cdc20 and APC/C is activated. Active APC/C tags inhibitory proteins for degradation such as securin and cyclin B1. Cleavage of cohesin molecules is triggered and sister chromatids is pulled apart in anaphase.
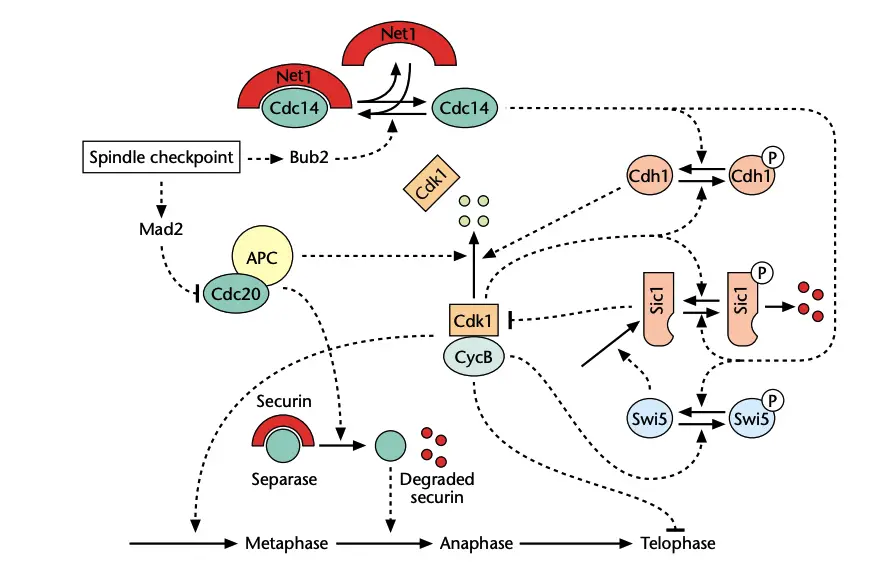
Functions of Metaphase (Spindle) Checkpoint
The metaphase checkpoint is a surveillance mechanism that operates during mitosis. It ensures that chromosomes are properly attached before separation. This process is referred to as spindle assembly checkpoint. It prevents unequal distribution of chromosomes in daughter cells.
Some of the main functions are–
- Surveillance of Kinetochore Attachment –
It is the process by which the kinetochores of sister chromatids are monitored for proper attachment with spindle microtubules. The checkpoint verifies that bipolar attachment (amphitelic orientation) is achieved. In this condition the kinetochores of each chromatid pair are attached to opposite poles. If attachment is improper the signal is generated and anaphase is delayed. - Mechanical Tension Sensing –
It is a mechanical and spatial sensing mechanism. Proper bipolar attachment generates tension across the centromere region. This tension is detected by the checkpoint proteins to confirm stability of microtubule binding. If tension is absent the process is stopped and correction is allowed. - Inhibition of Premature Anaphase –
The major function is to prevent entry into anaphase until all chromosomes are aligned at metaphase plate. Unattached kinetochore produces a diffusible “wait” signal. Even a single unattached kinetochore is sufficient to block the progression of cell cycle. This ensures that chromatids is not separated early. - Regulation of APC/C Activity –
To maintain arrest the Mitotic Checkpoint Complex (MCC) is formed. This complex binds to Anaphase-Promoting Complex/Cyclosome (APC/C) and inhibits its activity. As a result securin and cyclin B is not degraded. Separase enzyme remains inactive and cohesin rings are not cleaved. Sister chromatids remain attached until proper signal is received. - Providing Time for Error Correction –
By delaying cell cycle progression time is provided for correcting faulty attachments. Syntelic attachment (both chromatids attached to same pole) and monotelic attachment (only one kinetochore attached) can be corrected during this pause. This process occurs when improper microtubule orientation is detected. - Prevention of Genomic Instability –
It is the ultimate role of the checkpoint to prevent chromosome missegregation and aneuploidy (abnormal chromosome number). Accurate segregation is ensured only when all conditions are satisfied. This mechanism acts as barrier against genomic instability, developmental defects and tumor formation.
Thus the metaphase checkpoint plays an important regulatory role in maintaining chromosomal stability during mitosis.
Regulation of Checkpoint
Core Regulatory Molecules of the Cell Cycle
- Progression of cell cycle is driven by balance of regulatory molecules. These molecules assess internal and external conditions and cell cycle is allowed only when condition is suitable.
- Positive regulators (engine) are cyclins and cyclin dependent kinases (CDKs). These proteins propel cell cycle forward.
- Cyclin levels fluctuates cyclically throughout cell cycle. CDK levels remain relatively stable.
- For cell to advance to next phase CDKs must bind with specific cyclins and active complex is formed. This complex must be phosphorylated by CDK activating kinase (CAK) for full activity.
- Negative regulators (brakes) act to halt cell cycle when problematic condition is present such as DNA damage.
- Cyclin dependent kinase inhibitors (CKIs) directly inhibit cyclin/CDK activity. These are divided into two main groups.
- INK4 family includes p15 p16 p18 and p19. These specifically blocks CDK4 and CDK6.
- CIP/KIP family includes p21 p27 and p57. These broadly inhibits multiple cyclin/CDK complexes.
- Tumor suppressors are also negative regulators. Proteins such as p53 and retinoblastoma (Rb) act as major roadblock. It prevents replication of damaged or incomplete genomes.
G1 Checkpoint (Restriction Point) Regulation
- G1 checkpoint is located in late G1 phase. It determines whether cell commits irreversibly to DNA replication. It is based on cell size energy reserves and absence of DNA damage.
- Key regulatory molecules involved are Cyclin D CDK4/6 Cyclin E CDK2 Rb E2F transcription factors p53 and p21.
- Regulation mechanism (normal progression) occurs in presence of external growth factors. Cyclin D is synthesized and it binds with CDK4/6. Active Cyclin D–CDK4/6 complex is formed.
- Cyclin D–CDK4/6 complex mono phosphorylates Rb protein. Normally unphosphorylated Rb binds with E2F transcription factors and E2F is inhibited. Cycle is halted by this binding.
- Phosphorylation reduces Rb grip on E2F and E2F is released. E2F transcribes genes essential for entering S phase. Cyclin E is also produced by this transcription.
- Cyclin E pairs with CDK2 and Cyclin E–CDK2 complex is formed. It hyper phosphorylates Rb. Positive feedback loop is created and E2F is freed completely. Cell passes restriction point and entry into S phase is done.
- Arrest mechanism occurs when DNA damage is detected. Tumor suppressor p53 is stabilized and activated. Activated p53 acts as transcription factor and expression of CKI p21 is induced.
- p21 binds with Cyclin E/CDK2 complexes and inhibition is done. Rb remains in active suppressive state and cell is arrested in G1. Time is provided for DNA repair.
G2/M Checkpoint Regulation
- G2/M checkpoint occurs before cell enters into mitosis. It guarantees that DNA replication is completed and no DNA damage remains.
- Key regulatory molecules are Cyclin B CDK1 (Cdc2) Cdc25 phosphatases (A B C) Wee1 Myt1 ATM ATR Chk1 and Chk2.
- Regulation mechanism (normal progression) occurs for initiation of mitosis. Cyclin B–CDK1 complex is master regulator and it is also referred to as Mitosis Promoting Factor. This complex must be fully activated for mitotic entry.
- During G2 phase Cyclin B–CDK1 complex is kept inactive. Inhibitory phosphorylation is added by Wee1 and Myt1 kinases and complex remains off.
- When cell is ready to divide Cdc25 phosphatases remove inhibitory phosphates. Cyclin B–CDK1 complex is switched on and entry into mitosis is driven.
- Arrest mechanism occurs during DNA damage. Sensor proteins such as MRN complex or RPA detects DNA double strand breaks or single strand breaks. ATM and ATR transducer kinases is activated.
- ATM and ATR phosphorylates and activates checkpoint effector kinases Chk1 and Chk2. Activated Chk1/Chk2 phosphorylates Cdc25 phosphatases.
- Cdc25A is tagged for proteasomal degradation. Cdc25C is sequestered in cytoplasm by 14-3-3 proteins.
- Chk1 also phosphorylates and activates inhibitory Wee1 kinase. Cdc25 is not active for removing inhibitory phosphates and Wee1 applies inhibitory phosphorylation. Cyclin B–CDK1 complex remains locked inactive and cell cycle is halted safely in G2.
Metaphase (Spindle Assembly) Checkpoint Regulation
- Spindle assembly checkpoint occurs during M phase. It ensures that sister chromatids are not pulled apart until all kinetochores is attached properly with spindle microtubules from opposite poles. It is a surveillance mechanism and anaphase is prevented until condition is met.
- Key regulatory molecules are Mps1 kinase Knl1 Bub1 Bub3 BubR1 Mad1 Mad2 Cdc20 APC/C (Anaphase Promoting Complex/Cyclosome) securin and separase.
- Arrest mechanism occurs when kinetochore is unattached. Even a single unattached kinetochore can activate checkpoint and arrest is maintained. It acts as catalytic hub.
- Mps1 kinase localizes to unattached kinetochores. It phosphorylates MELT repeating motifs on Knl1 protein.
- These phosphorylated motifs recruit Bub3 and Bub1. These proteins attract and scaffold other checkpoint proteins such as Mad1 Mad2 BubR1 and Cdc20 in close proximity.
- This crowding catalyzes formation of Mitotic Checkpoint Complex (MCC). MCC is a tetrameric inhibitor made of Mad2 BubR1 Bub3 and Cdc20.
- MCC binds tightly with APC/C and it acts as roadblock. APC/C is inhibited and its E3 ubiquitin ligase activity is blocked.
- Regulation mechanism (normal progression or checkpoint silencing) occurs when APC/C inhibition is removed. As long as APC/C is inhibited by MCC securin and Cyclin B1 is spared from destruction and anaphase onset is prevented.
- When all kinetochores are securely attached and tension is present SAC signal is silenced. Mps1 is displaced from kinetochore. Phosphatases such as PP1 and PP2A remove activating phosphorylation from Knl1.
- MCC disassembles quickly and Cdc20 is freed. Cdc20 binds with APC/C and APC/C is activated.
- Active APC/C–Cdc20 complex polyubiquitinates securin and Cyclin B1 and rapid degradation is done. Destruction of securin unleashes separase. Separase cleaves cohesin rings holding sister chromatids together and separation of sister chromatids is triggered to opposite poles.
What are the roles of cyclins and CDKs?
- Cyclins and cyclin dependent kinases (CDKs) are primary positive regulators of cell cycle. It acts as engine and cell cycle progression is driven. Together it forms active complexes and timing of DNA replication and division is controlled.
- Role of CDKs
- CDKs are catalytic subunit of the complex.
- It is serine/threonine protein kinase and phosphorylation of target proteins is done.
- Phosphate group is attached and shape of substrate proteins is changed. Activation of proteins occurs and events required for next stage is carried out.
- Concentration of CDK proteins remain relatively stable throughout cell cycle.
- Role of Cyclins
- Cyclins are regulatory subunit and it does not have enzymatic activity alone.
- It provides substrate specificity and CDKs is directed to correct target proteins.
- Cyclins is strictly required for CDK activation.
- Cyclin levels fluctuates periodically in predictable pattern.
- It accumulates in response to internal and external growth signals. It is degraded rapidly by cytoplasmic enzymes after passing checkpoint.
- Mechanism of Action
- CDK remains inactive alone and regulation of cell cycle is done only when it binds tightly with specific cyclin.
- After CDK/cyclin complex is formed it must be phosphorylated by CDK activating kinase (CAK) for full function.
- Without proper concentration of fully activated complexes cell cycle is halted.
- Stage specific Cyclin–CDK combinations
- G1 phase and G1/S transition
- D type cyclins (D1 D2 D3) act as growth sensors. It accumulates in response to external growth factors.
- It binds with CDK4 and CDK6 and complex is formed. This complex phosphorylates retinoblastoma protein (Rb).
- E2F transcription factors is released and cell passes G1 restriction point.
- Cyclin E then binds with CDK2. Transition into S phase is facilitated and DNA replication is initiated.
- S phase and G2/M transition
- Cyclin A binds with CDK2 during S phase and progression is driven.
- Later Cyclin A partners with CDK1 and transition from G2 phase to mitosis is helped.
- M phase (mitosis)
- Cyclin B forms complex with CDK1 (cdc2 or mitosis promoting factor MPF).
- Activation of this complex is essential for initiation of mitosis.
- Chromosome condensation and assembly of mitotic spindle is enabled and mitotic events is carried out.
- G1 phase and G1/S transition
How do checkpoints help prevent cancer?
- Checkpoints help prevent cancer by acting as surveillance mechanism in cell cycle. It maintains genomic stability and proliferation of compromised cells is prevented. Cancer begins when genetic errors is not corrected and uncontrolled cell growth occurs.
- Checkpoints monitors DNA damage and incomplete chromosome replication. If problem is detected progression of cell cycle is halted. This arrest provides time for repair of damaged DNA before replication or division.
- If DNA damage is severe and it is not repaired checkpoint regulators can trigger apoptosis (programmed cell suicide). Permanent cellular senescence can also occur. It acts as fail safe and heavily mutated DNA is not duplicated and passed to daughter cells.
- Spindle checkpoint prevents unequal distribution of chromosomes during division. It protects against aneuploidy (chromosomal instability). Aneuploidy is a hallmark of many human solid tumors.
- Checkpoints act as brakes on uncontrolled division. Proteins enforcing checkpoints are mainly negative regulators such as p53 p21 and Rb. These are classified as tumor suppressor proteins.
- p53 is referred to as guardian of genome. It detects DNA damage and repair or cell death is controlled. If checkpoint proteins are mutated these roadblocks fails. Cells bypass checkpoints and mutations accumulates rapidly and uncontrolled replication occurs.
How do tumor suppressor genes like p53 protect the cell?
- Tumor suppressor genes codes for negative regulatory proteins. It act as roadblock and uncontrolled division of damaged cells is prevented. p53 is most important tumor suppressor gene and it is referred to as guardian of genome.
- Cell cycle arrest
- When cell is exposed to stress or DNA damage p53 acts at G1 and G2/M checkpoints. Cell cycle is halted temporarily.
- p53 acts as transcription factor and production of p21 is increased.
- p21 binds with cyclin dependent kinase (CDK) complexes and inhibition is done. Brake is applied and cell does not replicate damaged DNA or enter into mitosis.
- DNA repair
- When cell cycle is paused repair of genome is coordinated by p53.
- Repair enzymes is recruited to sites of DNA damage.
- Genes involved in DNA repair pathways is activated such as p53R2 and Mlh1.
- Apoptosis (programmed cell death)
- If DNA damage is severe and it cannot be repaired p53 triggers apoptosis. Mutated cell is removed permanently.
- p53 activates pro apoptotic genes such as Bax NOXA PUMA and Apaf-1. Anti apoptotic proteins is suppressed.
- p53 can translocate to mitochondria and membrane destabilization is induced. Cytochrome c is released and cell is destroyed.
- Alternative cell death and stress responses
- p53 can induce autophagy (cellular self digestion) by activating genes like DRAM and Atg.
- p53 can trigger ferroptosis (iron dependent cell death) in response to high reactive oxygen species.
- p53 can also drive cellular senescence. It is permanent and irreversible growth arrest and damaged cells does not divide.
- Metabolic regulation and stemness inhibition
- Cancer cells use altered metabolism such as aerobic glycolysis (Warburg effect). p53 suppresses glycolysis and mitochondrial respiration is promoted.
- Lipid and cholesterol synthesis pathways is restricted.
- Stem like properties is inhibited by p53. Stem cell factors such as OCT4 and SOX2 is suppressed and differentiation is promoted.
- Danger of p53 mutation
- TP53 gene is commonly mutated in human cancers. It is found damaged missing or inactive in many tumors (more than 50%).
- When p53 function is lost detection of DNA errors is reduced. Repair signals and cell death response is not triggered.
- Cell passes checkpoints even when damage is present. Mutations accumulates and aggressive tumor growth can occur.
What is the difference between the G1 and G2 checkpoints?
- G1 and G2 checkpoints are quality control points in cell cycle. It occurs at different stages and different conditions is checked before division is allowed.
- Timing and location
- G1 checkpoint occurs near end of G1 phase. It occurs before entry into S phase (synthesis phase). It is also referred to as restriction point in mammalian cells or START in yeast.
- G2 checkpoint occurs at transition between G2 phase and M phase. It occurs after DNA is supposed to be fully replicated.
- Primary purpose and conditions monitored
- In G1 checkpoint irreversible commitment to cell division is made. Cell checks internal and external conditions for DNA replication. Cell size nutrient availability energy reserves extracellular growth factors and absence of genomic DNA damage is checked. If requirement is not met cycle can be halted for repair. Cell can enter into G0 (resting phase). Apoptosis can also occur if damage is not repaired.
- In G2 checkpoint entry into mitosis is blocked if condition is not met. It also checks cell size and protein reserves. Major role is to ensure that chromosomes is fully replicated accurately and replicated DNA is free from damage. If damage or stalled replication is detected cell cycle pauses for completion of replication or repair. If damage is irreparable apoptosis can occur and mitotic catastrophe can also occur.
- Molecular mechanisms and regulators
- G1 checkpoint regulators
- Forward progression is driven by cyclins D and E. It forms active complexes with CDK4 CDK6 and CDK2.
- Phosphorylation of Rb protein is defining mechanism. Active CDK complexes phosphorylates Rb and E2F transcription factors is released. Genes required for S phase is activated.
- If DNA damage is present p53 accumulates and p21 production is triggered. p21 is CDK inhibitor and cyclin–CDK complexes is blocked and cycle is halted.
- G2 checkpoint regulators
- Progression depends on activation of Cyclin B–CDK1 complex. It is master regulator of mitotic entry.
- Inhibitory phosphates must be removed by Cdc25 phosphatases for CDK1 activation.
- If DNA damage is sensed ATM and ATR kinases is recruited. It activates checkpoint kinases Chk1 and Chk2.
- Chk1 and Chk2 phosphorylates Cdc25 and Cdc25 is degraded or it is sequestered in cytoplasm by 14-3-3 proteins. Cdc25 cannot activate Cyclin B–CDK1 complex and cell remains locked in G2 until damage is resolved.
- G1 checkpoint regulators
How does the spindle assembly checkpoint prevent chromosomal errors?
- Spindle assembly checkpoint (SAC) prevents chromosomal errors by acting as surveillance mechanism during metaphase to anaphase transition. It ensures that cell will not separate sister chromatids until every chromosome is aligned and attached to mitotic spindle. It prevents aneuploidy (abnormal number of chromosomes).
- Generation of “wait” signal occurs when even a single kinetochore is unattached or improperly attached. A diffusible inhibitory signal is produced and cell cycle progression is halted. Unattached kinetochore acts as catalytic platform and Mitotic Checkpoint Complex (MCC) is formed. MCC is tetrameric complex made of Mad2 BubR1 Bub3 and Cdc20.
- MCC halts cell cycle engine by inhibiting APC/C (Anaphase Promoting Complex/Cyclosome). APC/C is E3 ubiquitin ligase and it normally tags securin and cyclin B1 for degradation. When APC/C is inhibited securin remains stable and separase is inhibited. Separase is protease enzyme which cleaves cohesin rings holding sister chromatids together. Cohesin remains intact so sister chromatids cannot be pulled apart early and unequal distribution of chromosomes is prevented.
- SAC also checks tension for correcting faulty attachments. It is not only microtubule presence but proper bipolar tension is required. Sister kinetochores must be attached to microtubules from opposite poles and tension is generated. If attachment is incorrect such as syntelic or merotelic proper tension is not formed and error is detected.
- Aurora B kinase acts as tension sensor. It is located at inner centromere. It detects low tension attachments and phosphorylates outer kinetochore proteins such as Ndc80 complex. This phosphorylation destabilizes faulty microtubule attachment and kinetochore releases microtubule. Attachment is tried again until proper bipolar tension bearing connection is established.
- When all chromosomes are bi oriented and proper tension is present SAC signal is silenced. MCC disassembles and APC/C is activated. APC/C degrades securin and separase is released. Separase cleaves cohesin and sister chromatids is pulled apart. Each daughter cell receives exact copy of genome and chromosomal errors is prevented.
How do Chk1 and Chk2 inhibit the CDK1 complex?
Chk1 and Chk2 inhibits CDK1 (cdc2) complex by targeting regulatory enzymes which controls phosphorylation state. It is done during DNA damage response and cell cycle is halted in G2.
Inhibition and sequestration of Cdc25 phosphatases is done. For CDK1/Cyclin B complex activation inhibitory phosphates (Thr14 and Tyr15) must be removed by Cdc25 family phosphatases. Chk1 and Chk2 phosphorylates Cdc25 and it becomes inactive.
Phosphorylation of Cdc25C creates binding site for 14-3-3 proteins. 14-3-3 binds with Cdc25C and it is sequestered in cytoplasm. Cdc25C is separated from nuclear CDK1 targets so activation is not done. Chk1 phosphorylates Cdc25B and it is sequestered away from centrosomes and CDK1 activation is reduced.
Degradation of Cdc25A is also done. Chk1 phosphorylates Cdc25A and it is tagged for ubiquitin mediated degradation by proteasome. Cdc25A is eliminated and it cannot activate CDK1.
Activation of Wee1 kinase is done by Chk1. Wee1 adds inhibitory phosphates to CDK1. When Wee1 activity is increased CDK1 remains suppressed and phosphorylated state is maintained. Myt1 is also stabilized and inhibitory phosphorylation is supported.
Chk1 and Chk2 activates p53 pathway. It phosphorylates and stabilizes tumor suppressor p53 and degradation is prevented. Stabilized p53 induces target genes such as p21 and 14-3-3 sigma. p21 inhibits cyclin/CDK complexes. 14-3-3 sigma binds with Cyclin B/CDK1 complex and it is sequestered in cytoplasm. Nuclear entry is blocked and mitotic entry is prevented.
What is the role of p53 in G2 arrest?
p53 plays role in G2 arrest when DNA damage is detected. It stops cell cycle progression into mitosis at G2/M checkpoint.
p53 is activated by upstream checkpoint kinases such as ATM/ATR and Chk1/Chk2. It acts as transcription factor and inhibition of CDC2/Cyclin B complex (CDK1/Cyclin B) is done. This complex is master kinase required for mitotic entry.
Induction of 14-3-3 sigma is done by p53. 14-3-3 sigma binds with CDC2/Cyclin B complex and it is sequestered in cytoplasm. Nuclear targets is not reached and premature mitosis is prevented.
Induction of GADD45 is done by p53. GADD45 disrupts CDC2/Cyclin B complex. Dissociation of two subunits is caused and complex becomes inactive.
Induction of p21 and Reprimo occurs by p53. p21 is CDK inhibitor and CDC2 kinase activity is inhibited. Reprimo also inhibits CDC2 kinase activity and G2 arrest is maintained.
Transcriptional repression is also done by p53. p53 represses transcription and downregulation of genes coding for CDC2 and Cyclin B is mediated. CDK1/Cyclin B level is reduced and entry into mitosis is blocked.
How do ATM and ATR kinases detect DNA damage?
- ATM and ATR kinases are central transducers in DNA damage response. It detects different type of DNA lesion and signalling is initiated by different sensor complexes.
- Detection by ATM kinase
- ATM mainly monitors DNA double strand breaks (DSBs) such as damage caused by ionizing radiation.
- ATM does not sense break directly. MRN complex acts as initial detector. MRN complex is made of Mre11 Rad50 and Nbs1.
- Mre11 binds to DNA at break site. Rad50 bridges broken double stranded ends and lesion is stabilized.
- MRN complex recruits ATM to site of break through interaction with Nbs1 subunit.
- In normal cells ATM is maintained inactive as homodimer. When ATM is recruited conformational change occurs and autophosphorylation is triggered (Ser1981).
- Autophosphorylation forces dimer to dissociate into active ATM monomers. These monomers phosphorylates downstream targets such as Chk2 and cell cycle arrest is supported.
- Detection by ATR kinase
- ATR is activated mainly by single stranded DNA (ssDNA). It arises from stalled replication forks UV induced damage or resection of double strand breaks.
- When ssDNA is exposed it is coated by Replication Protein A (RPA) and stabilization is done.
- RPA coated ssDNA acts as landing pad and ATR is recruited with ATRIP (ATR interacting partner).
- For full activation additional factors is required at damage site. Rad17 complex is recruited and 9-1-1 sliding clamp is loaded (Rad9 Rad1 Hus1).
- TopBP1 activator protein is also required and kinase activation is enhanced.
- An alternative pathway is also present where ATR can be activated by mismatch repair proteins recognizing DNA lesions.
What happens if a cell enters the G0 phase?
G0 phase is referred to as quiescent or resting phase. In this phase cell exits from active cell cycle. Cell is not preparing for division and cell cycle progression is stopped.
Entry into G0 usually occurs from G1 phase. It occurs when extracellular growth factors is not available. It can also occur when G1 checkpoint requirement is not met and restriction point is not passed.
In G0 state cell remains metabolically active. Growth is stopped and rate of protein synthesis is reduced. Division related preparation is not continued.
G0 can be temporary in many cells. Cell remains in this arrested state until condition becomes favourable. When mitogenic signals (growth factors) is received cell can re-enter into G1 phase and cell cycle is resumed.
G0 can be permanent in some specialised cells. Mature cardiac muscle cells and cortical neurons remain locked in G0 for lifetime. Cell division is not done in these cells.
Functions of Checkpoints
- Checkpoints acts as internal surveillance mechanism in cell cycle. Proper and accurate progression of cell division is ensured. Overall genomic stability is maintained.
- Cell cycle arrest is done by checkpoints. Progression is halted and time is provided for completing critical processes. Repair of problematic condition such as damaged DNA is allowed before moving to next phase.
- If DNA damage is severe and it is not repaired apoptosis (programmed cell death) can be induced. Permanent arrest (senescence) can also be induced. It acts as fail safe and replication of mutated potentially cancerous cells is prevented.
- G1 checkpoint (restriction point) functions
- Cell size is assessed.
- Nutrient availability and energy reserves is checked.
- Extracellular growth signals is checked.
- Genomic integrity is verified and DNA damage is checked.
- It determines whether cell will commit irreversibly to DNA replication or it will enter into G0 (resting state).
- S phase checkpoint functions
- DNA replication process is monitored.
- Replication fork stability is evaluated.
- DNA synthesis is slowed down or halted when damage or errors is detected.
- DNA repair is facilitated during this arrest.
- G2 checkpoint (G2/M checkpoint) functions
- Entry into mitotic phase (M phase) is prevented when condition is not met.
- It verifies that entire genome is fully replicated accurately.
- Newly replicated DNA is checked and damage should not be present.
- M checkpoint (spindle assembly checkpoint) functions
- Premature separation of sister chromatids in anaphase is prevented.
- It ensures that all kinetochores is firmly and correctly attached to spindle microtubules from opposite poles.
- Chromosome missegregation is prevented and aneuploidy is avoided.
Refereneces
- Blagosklonny, M. V., & Pardee, A. B. (2000-2013). The restriction point of the cell cycle. In Madame Curie Bioscience Database. Landes Bioscience.
- Breastcancer.org. (n.d.). What are CDK4/6 inhibitors?
- Bucher, N., & Britten, C. D. (2008). G2 checkpoint abrogation and checkpoint kinase-1 targeting in the treatment of cancer. British Journal of Cancer, 98(3), 523–528. https://doi.org/10.1038/sj.bjc.6604208
- Calonge, T. M., & O’Connell, M. J. (2008). Turning off the G2 DNA damage checkpoint. DNA Repair, 7(2), 136–140. https://doi.org/10.1016/j.dnarep.2007.07.017
- Carter, S., & Lumen Learning. (n.d.). Introduction to cell cycle checkpoints. In Biology for majors I. Lumen Learning.
- Cell cycle checkpoint. (2025, September 30). In Wikipedia. https://en.wikipedia.org/wiki/Cell_cycle_checkpoint
- Cooper, G. M. (2000). The eukaryotic cell cycle. In The Cell: A Molecular Approach (2nd ed.). Sinauer Associates.
- di Rorà, A. G. L., Martinelli, G., & Simonetti, G. (2019). The balance between mitotic death and mitotic slippage in acute leukemia: a new therapeutic window? Journal of Hematology & Oncology, 12, 123. https://doi.org/10.1186/s13045-019-0808-4
- Ding, L., Cao, J., Lin, W., Chen, H., Xiong, X., Ao, H., Yu, M., Lin, J., & Cui, Q. (2020). The roles of cyclin-dependent kinases in cell-cycle progression and therapeutic strategies in human breast cancer. International Journal of Molecular Sciences, 21(6), 1960. https://doi.org/10.3390/ijms21061960
- Engeland, K. (2022). Cell cycle regulation: p53-p21-RB signaling. Cell Death & Differentiation, 29(5), 946–960. https://doi.org/10.1038/s41418-022-00988-z
- Foster, D. A., Yellen, P., Xu, L., & Saqcena, M. (2010). Regulation of G1 cell cycle progression: Distinguishing the restriction point from a nutrient-sensing cell growth checkpoint(s). Genes & Cancer, 1(11), 1124–1131. https://doi.org/10.1177/1947601910392989
- Gao, T., Sun, Y., Leng, P., Liu, D., Guo, Q., & Li, J. (2025). CDK4/6 inhibitors in breast cancer therapy: mechanisms of drug resistance and strategies for treatment. Frontiers in Pharmacology, 16, 1549520. https://doi.org/10.3389/fphar.2025.1549520
- Hardy, M. (n.d.). 17.2 Regulator molecules of the cell cycle – College biology I. OPEN SLCC.
- Keezer, S. (2026, March 4). Hallmarks of cancer: Genome instability and mutation. CST Blog. Cell Signaling Technology.
- Kousholt, A. N., Menzel, T., & Sørensen, C. S. (2012). Pathways for genome integrity in G2 phase of the cell cycle. Biomolecules, 2(4), 579–607. https://doi.org/10.3390/biom2040579
- Lampson, M. A., & Cheeseman, I. M. (2011). Sensing centromere tension: Aurora B and the regulation of kinetochore function. Trends in Cell Biology, 21(3), 133–140.
- Lara-Gonzalez, P., Pines, J., & Desai, A. (2021). Spindle assembly checkpoint activation and silencing at kinetochores. Seminars in Cell & Developmental Biology, 117, 86–98. https://doi.org/10.1016/j.semcdb.2021.06.009
- Lischetti, T., & Nilsson, J. (2015). Regulation of mitotic progression by the spindle assembly checkpoint. Molecular & Cellular Oncology, 2(1), e970484. https://doi.org/10.4161/23723548.2014.970484
- Liu, Y., Park, S., & Li, Y. (2025). Breaking cancer’s momentum: CDK4/6 inhibitors and the promise of combination therapy. Cancers, 17(12), 1941. https://doi.org/10.3390/cancers17121941
- McVey, S. L., Cosby, J. K., & Nannas, N. J. (2021). Aurora B tension sensing mechanisms in the kinetochore ensure accurate chromosome segregation. International Journal of Molecular Sciences, 22(16), 8818. https://doi.org/10.3390/ijms22168818
- Med-Ace. (n.d.). Cell cycle & regulation, mitosis, cyclins, RB, p53 & tumor suppressors (USMLE essentials) [Video]. YouTube.
- Molinari, M. (2001). Cell cycle checkpoints and their inactivation in human cancer. Cell Proliferation, 33(5), 261–274. https://doi.org/10.1046/j.1365-2184.2000.00191.x
- Nakanishi, M., Shimada, M., & Niida, H. (2006). Genetic instability in cancer cells by impaired cell cycle checkpoints. Cancer Science, 97(10), 984–989. https://doi.org/10.1111/j.1349-7006.2006.00289.x
- Patil, M., Pabla, N., & Dong, Z. (2013). Checkpoint kinase 1 in DNA damage response and cell cycle regulation. Cellular and Molecular Life Sciences, 70(21), 4009–4021. https://doi.org/10.1007/s00018-013-1307-3
- Price, K. (2018, August 24). The stop and go of the cell cycle: Research reveals an important checkpoint. Stanford Medicine News Center.
- Pun, R., & North, B. J. (2025). Role of spindle assembly checkpoint proteins in gametogenesis and embryogenesis. Frontiers in Cell and Developmental Biology, 12, 1491394. https://doi.org/10.3389/fcell.2024.1491394
- QIAGEN. (n.d.). Cell cycle & cell division. GeneGlobe.
- QIAGEN. (n.d.). Cell cycle: G2/M DNA damage checkpoint regulation. GeneGlobe.
- Restriction point. (2026, March 11). In Wikipedia. https://en.wikipedia.org/wiki/Restriction_point
- Spindle checkpoint. (2025, October 3). In Wikipedia. https://en.wikipedia.org/wiki/Spindle_checkpoint
- Stier, A. B., Bonaiuti, P., Juhász, J., Gross, F., & Ciliberto, A. (2025). lncreased risk of slippage upon disengagement of the mitotic checkpoint. PLoS Computational Biology, 21(3), e1012879. https://doi.org/10.1371/journal.pcbi.1012879
- Stracker, T. H., Usui, T., & Petrini, J. H. J. (2009). Taking the time to make important decisions: the checkpoint effector kinases Chk1 and Chk2 and the DNA damage response. DNA Repair, 8(9), 1047–1054. https://doi.org/10.1016/j.dnarep.2009.04.012
- Susan G. Komen. (2025). CDK4/6 inhibitors. https://www.komen.org/breast-cancer/treatment/type/cdk4-6-inhibitors/
- Taveras, C., Liu, C., & Mao, Y. (2019). A tension-independent mechanism reduces Aurora B-mediated phosphorylation upon microtubule capture by CENP-E at the kinetochore. Cell Cycle, 18(12), 1349–1363. https://doi.org/10.1080/15384101.2019.1617615
- Tudzarova, S., Mulholland, P., Dey, A., Stoeber, K., Okorokov, A. L., & Williams, G. H. (2016). p53 controls CDC7 levels to reinforce G1 cell cycle arrest upon genotoxic stress. Cell Cycle, 15(21), 2958–2972. https://doi.org/10.1080/15384101.2016.1231281
- University of Southern Queensland. (n.d.). Cell cycle checkpoints. In Book 1: Biosciences for health professionals.
- Yao, Y., & Dai, W. (2014). Genomic instability and cancer. Journal of Carcinogenesis & Mutagenesis, 5, 1000165. https://doi.org/10.4172/2157-2518.1000165
- Yoo, T. Y., Choi, J.-M., Conway, W., Yu, C.-H., Pappu, R. V., & Needleman, D. J. (2018). Measuring NDC80 binding reveals the molecular basis of tension-dependent kinetochore-microtubule attachments. eLife, 7, e36392. https://doi.org/10.7554/eLife.36392
- Zhang, J., Xu, D., Zhou, Y., Zhu, Z., & Yang, X. (2021). Mechanisms and implications of CDK4/6 inhibitors for the treatment of NSCLC. Frontiers in Oncology, 11, 676041. https://doi.org/10.3389/fonc.2021.676041
- Zhou, Y., Nakajima, R., Shirasawa, M., Fikriyanti, M., Zhao, L., Iwanaga, R., Bradford, A. P., Kurayoshi, K., Araki, K., & Ohtani, K. (2023). Expanding roles of the E2F-RB-p53 pathway in tumor suppression. Biology, 12(12), 1511. https://doi.org/10.3390/biology12121511